Design and Optimization of Universal DNA Arrays Ion








![Algorithms for c-h Code Problem • [Ben-Dor et al. 00] approximation algorithm based on Algorithms for c-h Code Problem • [Ben-Dor et al. 00] approximation algorithm based on](https://slidetodoc.com/presentation_image_h2/6f013cf02dda9ebbb7d9d30d49c50611/image-11.jpg)
![Integer Program Formulation [MPT 05] • Maximum integer flow problem w/ set capacity constraints Integer Program Formulation [MPT 05] • Maximum integer flow problem w/ set capacity constraints](https://slidetodoc.com/presentation_image_h2/6f013cf02dda9ebbb7d9d30d49c50611/image-14.jpg)
![Periodic Tags [MT 05] • Key observation: c-token uniqueness constraint in c-h code formulation Periodic Tags [MT 05] • Key observation: c-token uniqueness constraint in c-h code formulation](https://slidetodoc.com/presentation_image_h2/6f013cf02dda9ebbb7d9d30d49c50611/image-16.jpg)










![SBE/SBH Assay [MP 06] Primers T T A T TTGCA AA AC CC CA SBE/SBH Assay [MP 06] Primers T T A T TTGCA AA AC CC CA](https://slidetodoc.com/presentation_image_h2/6f013cf02dda9ebbb7d9d30d49c50611/image-34.jpg)










- Slides: 48

Design and Optimization of Universal DNA Arrays Ion Mandoiu Computer Science & Engineering Department University of Connecticut http: //www. engr. uconn. edu/~ion/ 1

DNA Arrays • Exploit Watson-Crick complementarity to simultaneously perform a large number of substring tests • Used in a variety of high-throughput genomic analyses – Transcription (gene expression) analysis – Single Nucleotide Polymorphism (SNP) genotyping – Alternative splicing, Ch. IP-on-chip, tiling arrays, genomic-based species identification, point-of-service diagnosis, … • Common array formats involve direct hybridization between labeled DNA/RNA sample and DNA probes attached to a glass slide 2

Universal DNA Arrays • “Programable” arrays – Array consists of application independent oligonucleotides – Analysis carried by a sequence of reactions involving application specific primers – Flexible AND cost effective • Universal array architectures: tag arrays, APEX arrays, SBE/SBH arrays, … 5

Overview m Tag Array Design - Tag Set Design - Tag Assignment Algorithms m SBE/SBH Assays - Decoding and Multiplexing Algorithms m Conclusions 6

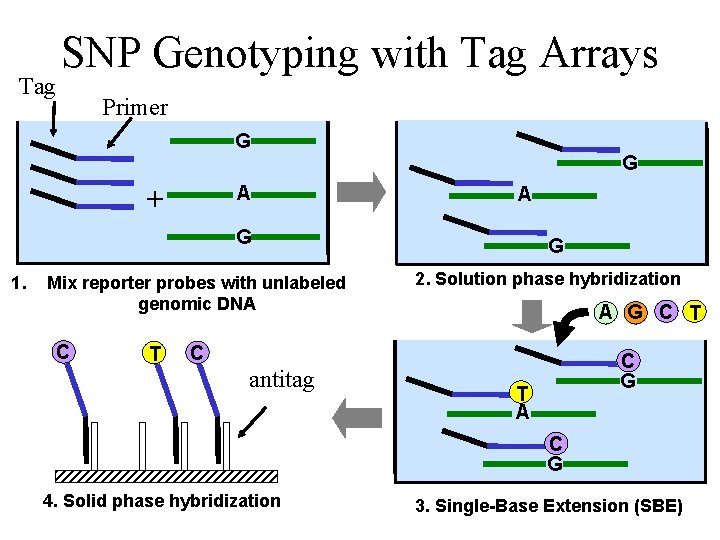
Tag SNP Genotyping with Tag Arrays Primer G + A G 1. Mix reporter probes with unlabeled genomic DNA C T G 2. Solution phase hybridization A G C T C antitag C G T A C G 4. Solid phase hybridization 7 3. Single-Base Extension (SBE)

Tag Array Advantages • Cost effective – Same array used in many analyses can be mass produced • Easy to customize – Only need to synthesize new set of reporter probes • Reliable – Solution phase hybridization better understood than hybridization on solid support 8

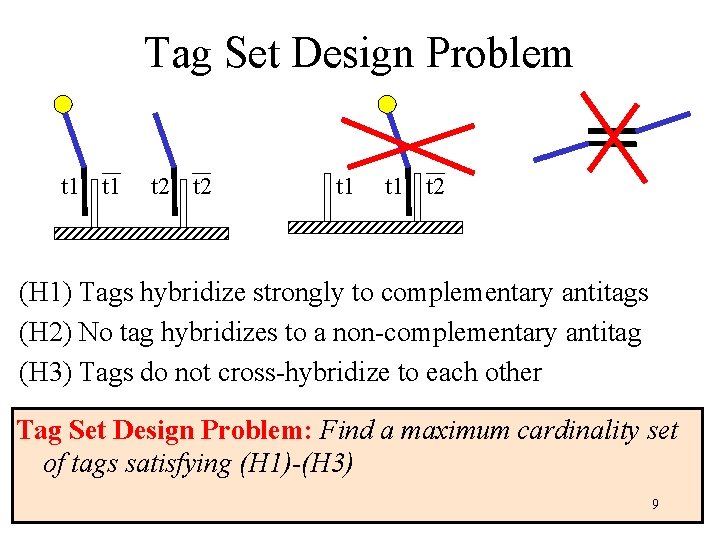
Tag Set Design Problem t 1 t 2 (H 1) Tags hybridize strongly to complementary antitags (H 2) No tag hybridizes to a non-complementary antitag (H 3) Tags do not cross-hybridize to each other Tag Set Design Problem: Find a maximum cardinality set of tags satisfying (H 1)-(H 3) 9

Hybridization Models • Melting temperature Tm: temperature at which 50% of duplexes are in hybridized state • 2 -4 rule Tm = 2 #(As and Ts) + 4 #(Cs and Gs) • More accurate models exist, e. g. , the nearneighbor model 10

Hybridization Models (contd. ) • Hamming distance model, e. g. , [Marathe et al. 01] – Models rigid DNA strands • LCS/edit distance model, e. g. , [Torney et al. 03] – Models infinitely elastic DNA strands • c-token model [Ben-Dor et al. 00]: – Duplex formation requires formation of nucleation complex between perfectly complementary substrings – Nucleation complex must have weight c, where wt(A)=wt(T)=1, wt(C)=wt(G)=2 (2 -4 rule) 11

c-h Code Problem • c-token: left-minimal DNA string of weight c, i. e. , – w(x) c – w(x’) < c for every proper suffix x’ of x • A set of tags is a c-h code if (C 1) Every tag has weight h (C 2) Every c-token is used at most once c-h Code Problem [Ben-Dor et al. 00] Given c and h, find maximum cardinality c-h code 12
![Algorithms for ch Code Problem BenDor et al 00 approximation algorithm based on Algorithms for c-h Code Problem • [Ben-Dor et al. 00] approximation algorithm based on](https://slidetodoc.com/presentation_image_h2/6f013cf02dda9ebbb7d9d30d49c50611/image-11.jpg)
Algorithms for c-h Code Problem • [Ben-Dor et al. 00] approximation algorithm based on De. Bruijn sequences • Alphabetic tree search algorithm - Enumerate candidate tags in lexicographic order, save tags whose ctokens are not used by previously selected tags - Easily modified to handle various combinations of constraints • [MT 05, 06] Optimum c-h codes can be computed in practical time for small values of c by using integer programming • Practical runtime using Garg-Koneman approximation and LP-rounding 13

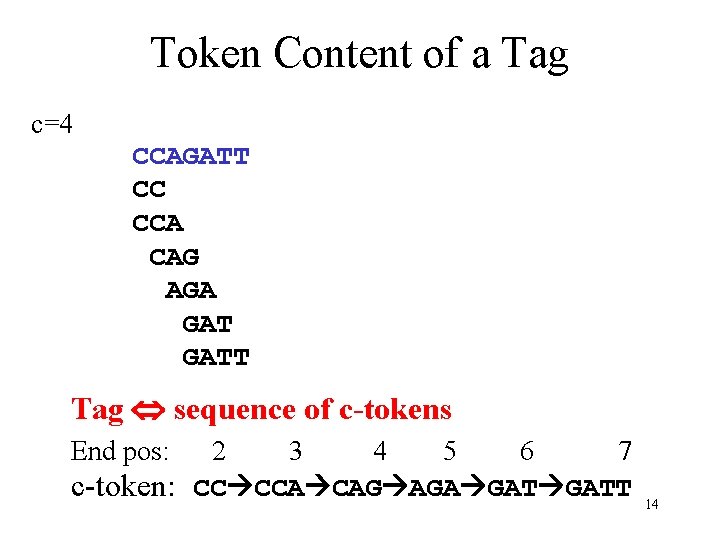
Token Content of a Tag c=4 CCAGATT CC CCA CAG AGA GATT Tag sequence of c-tokens End pos: 2 3 4 5 6 7 c-token: CC CCA CAG AGA GATT 14

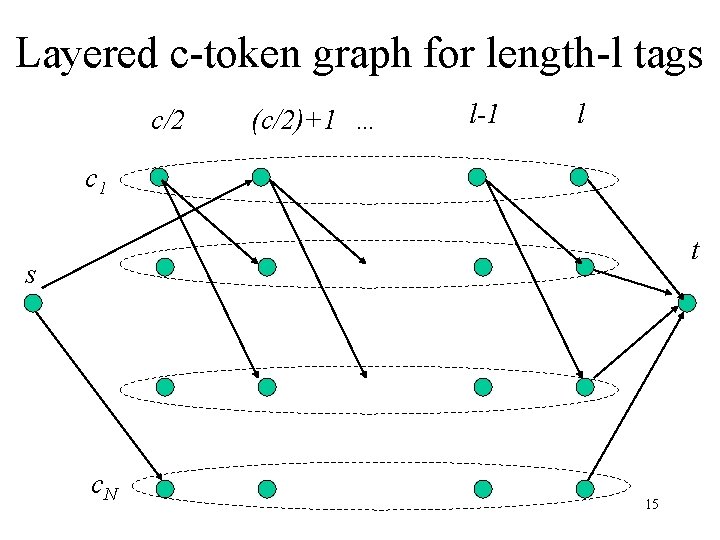
Layered c-token graph for length-l tags c/2 (c/2)+1 … l-1 l c 1 t s c. N 15
![Integer Program Formulation MPT 05 Maximum integer flow problem w set capacity constraints Integer Program Formulation [MPT 05] • Maximum integer flow problem w/ set capacity constraints](https://slidetodoc.com/presentation_image_h2/6f013cf02dda9ebbb7d9d30d49c50611/image-14.jpg)
Integer Program Formulation [MPT 05] • Maximum integer flow problem w/ set capacity constraints • O(h. N) constraints & variables, where N = #c-tokens 16

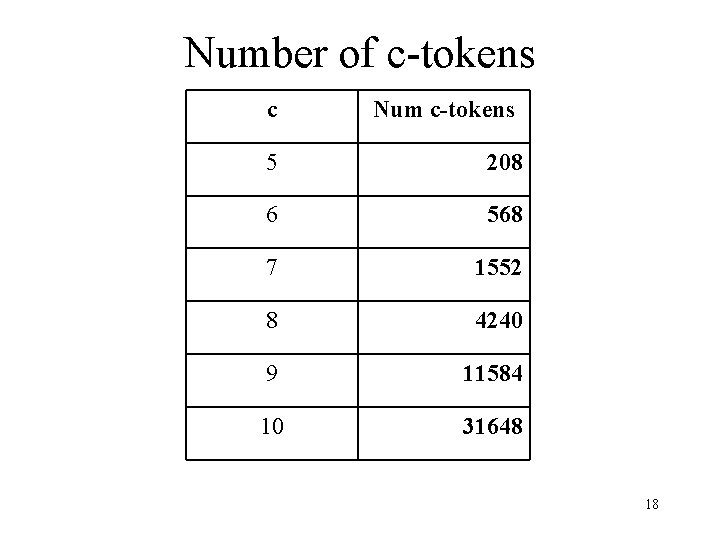
Number of c-tokens c Num c-tokens 5 208 6 568 7 1552 8 4240 9 11584 10 31648 18
![Periodic Tags MT 05 Key observation ctoken uniqueness constraint in ch code formulation Periodic Tags [MT 05] • Key observation: c-token uniqueness constraint in c-h code formulation](https://slidetodoc.com/presentation_image_h2/6f013cf02dda9ebbb7d9d30d49c50611/image-16.jpg)
Periodic Tags [MT 05] • Key observation: c-token uniqueness constraint in c-h code formulation is too strong – A c-token should not appear in two different tags, but can be repeated in a tag • A tag t is called periodic if it is the prefix of ( ) for some “period” – Periodic strings make best use of c-tokens 22

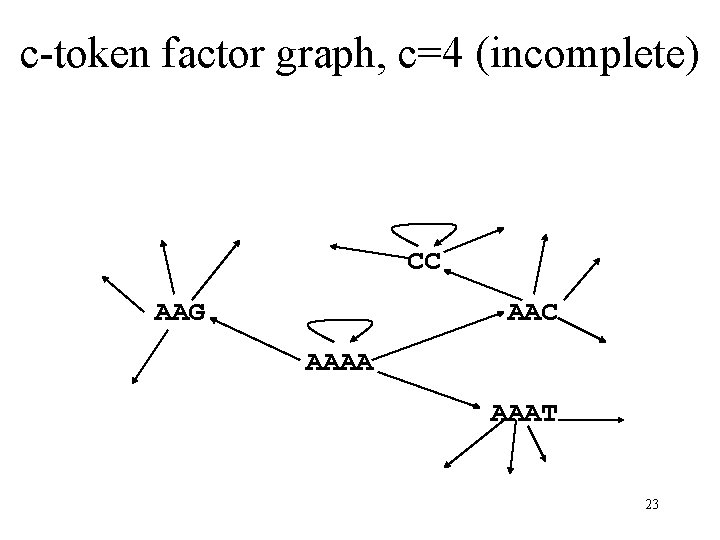
c-token factor graph, c=4 (incomplete) CC AAG AAC AAAA AAAT 23

Vertex-disjoint Cycle Packing Problem • Given directed graph G, find maximum number of vertex disjoint directed cycles in G • [MT 05] APX-hard even for regular directed graphs with in-degree and out-degree 2 – h-c/2+1 approximation factor for tag set design problem • [Salavatipour and Verstraete 05] – Quasi-NP-hard to approximate within (log 1 - n) – O(n 1/2) approximation algorithm 24

Cycle Packing Algorithm 1. Construct c-token factor graph G 2. T {} 3. For all cycles C defining periodic tags, in increasing order of cycle length, • Add to T the tag defined by C • Remove C from G 4. Perform an alphabetic tree search and add to T tags consisting of unused c-tokens 5. Return T – Gives an increase of over 40% in the number of tags compared to previous methods 25

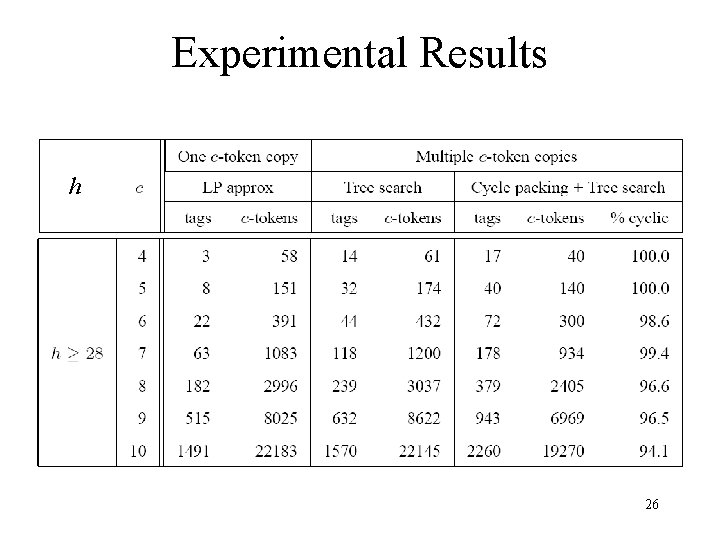
Experimental Results h 26

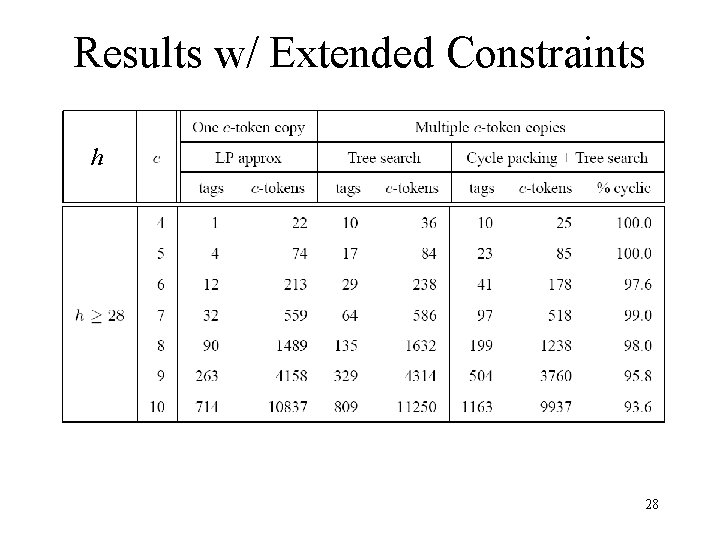
Antitag-to-Antitag Hybridization • Additional constraint: antitags do not cross-hybridize, including self • Formalization in c-token hybridization model: (C 3) No two tags contain complementary substrings of weight c • Cycle packing and tree search extend easily 27

Results w/ Extended Constraints h 28

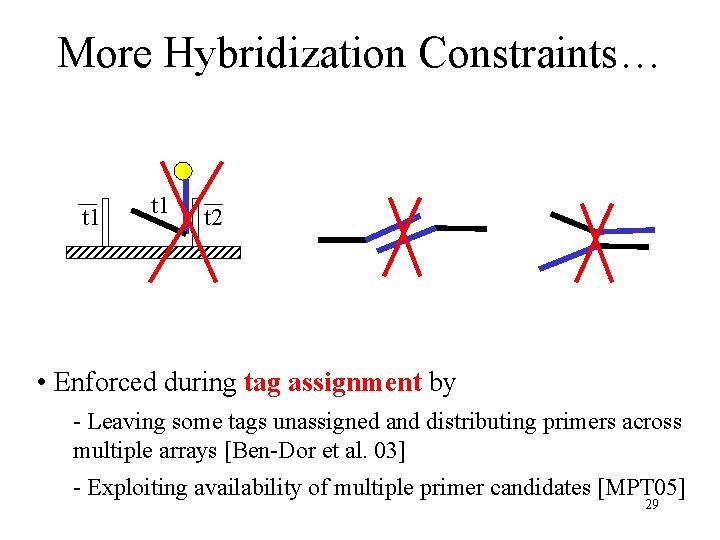
More Hybridization Constraints… t 1 t 2 • Enforced during tag assignment by - Leaving some tags unassigned and distributing primers across multiple arrays [Ben-Dor et al. 03] - Exploiting availability of multiple primer candidates [MPT 05] 29

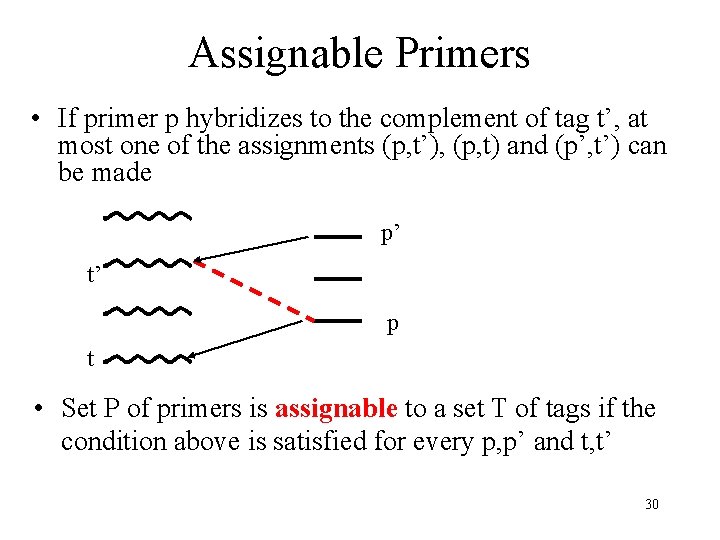
Assignable Primers • If primer p hybridizes to the complement of tag t’, at most one of the assignments (p, t’), (p, t) and (p’, t’) can be made p’ t’ p t • Set P of primers is assignable to a set T of tags if the condition above is satisfied for every p, p’ and t, t’ 30

Characterization of Assignable Sets • conflict graph: – G=(T P, E), where (t, p) ∈ E if t and p hybridize – X = number of primers adjacent to a degree 1 tag – Y = number of degree 0 tags X=1 Y=2 • [Ben-Dor 04] Set P is assignable to T iff X+Y |P| 31

Finding Assignable Primer Sets Multiplexing Problem: given primer set P and tag set T, find partition of P into minimum number of assignable sets Maximum Assignable Primer Set Problem: given primer set P and tag set T, find a maximum size assignable subset of P • Both problems are NP-hard [Ben-Dor 04] 32

Integration with Primer Selection • In practice, several primer candidates with equivalent functionality – In SNP genotyping, can pick primer from either forward and reverse strand – In gene expression/identification applications, many primers have desired length, Tm, etc. 33

Pooled Array Multiplexing Problem Pooled Multiplexing Problem: Given set of primer pools P and tag set T, find a primer from each pool and a partition of selected primers into minimum number of assignable sets 34

Pooled Multiplexing Algorithms 1. Primer-Del = greedy deletion for pools similar to [Ben-Dor et al 04] n Repeatedly delete primer of maximum potential until X+Y #pools, where q q Potential of tag t is 2 -deg(t) Potential of primer p is sum of potentials of conflicting tags q Subtract ½ if primer adjacent to a tag of degree 1 36

Pooled Multiplexing Algorithms 1. Primer-Del = greedy deletion for pools similar to [Ben-Dor et al 04] 2. Primer-Del+ = same but never delete last primer from pool unless no other choice 3. Min-Pot = select primer with min potential from each pool, then run Primer-Del 4. Min-Deg = select primer with min degree, then run Primer-Del 5. Iterative ILP = iteratively find a maximum assignable pool set using integer linear program 37

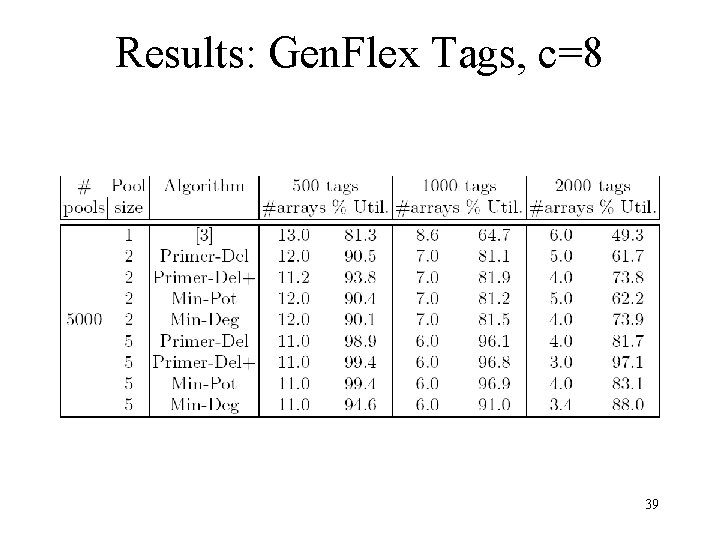
Results: Gen. Flex Tags, c=8 39

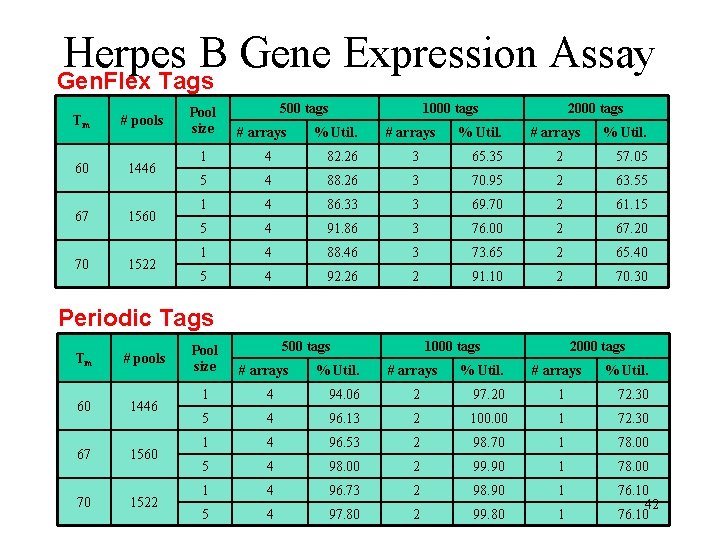
Herpes B Gene Expression Assay Gen. Flex Tags Tm # pools 60 1446 67 1560 70 1522 Pool size 500 tags # arrays % Util. 1000 tags # arrays % Util. 2000 tags # arrays % Util. 1 4 82. 26 3 65. 35 2 57. 05 5 4 88. 26 3 70. 95 2 63. 55 1 4 86. 33 3 69. 70 2 61. 15 5 4 91. 86 3 76. 00 2 67. 20 1 4 88. 46 3 73. 65 2 65. 40 5 4 92. 26 2 91. 10 2 70. 30 Periodic Tags Tm # pools 60 1446 67 1560 70 1522 Pool size 500 tags # arrays % Util. 1000 tags # arrays % Util. 2000 tags # arrays % Util. 1 4 94. 06 2 97. 20 1 72. 30 5 4 96. 13 2 100. 00 1 72. 30 1 4 96. 53 2 98. 70 1 78. 00 5 4 98. 00 2 99. 90 1 78. 00 1 4 96. 73 2 98. 90 1 5 4 97. 80 2 99. 80 1 76. 10 42 76. 10

Overview m Tag Array Design - Tag Set Design - Tag Assignment Algorithms m SBE/SBH Assays - Decoding and Multiplexing Algorithms m Conclusions 43
![SBESBH Assay MP 06 Primers T T A T TTGCA AA AC CC CA SBE/SBH Assay [MP 06] Primers T T A T TTGCA AA AC CC CA](https://slidetodoc.com/presentation_image_h2/6f013cf02dda9ebbb7d9d30d49c50611/image-34.jpg)
SBE/SBH Assay [MP 06] Primers T T A T TTGCA AA AC CC CA T AT AG CG CT TT TG GG GT TA TC GC GA hybridization to k-mer array (SBH) CCATT GATAA A T 44 single-base extension (SBE)

Some notations • P set of primers, X set of probes • Ep ⊆ {A, C, T, G} the set of possible extensions for primer p • The spectrum of primer p, Spec. X(p), is the set of probes hybridizing with p • The extended spectrum of primer p with extension set Ep, 45

Decodable primer sets • Four parallel single-color SBE/SBH experiments one type of extension in each SBE experiment – P is weakly decodable with respect to extension e if for every primer p • One SBE/SBH experiment with 4 colors (4 extensions) – P is weakly decodable if for every primer p and every extension e ∈ Ep 46

Strongly r-decodable primer sets • Hybridization involving labeled nucleotide is less predictable Informative probes should not rely on it • Signal from one SNP may obscure signal from another when read at the same probe due to differences in DNA amplification efficiency Informative probes cannot be shared between SNPs • P is strongly r-decodable if for every primer p where r = redundancy parameter 47

MPPP A set of primer pools P ={P 1, …, Pn } is strongly r-decodable iff there is a primer pi in each pool Pi such that {p 1, …, pn} is strongly r-decodable. n Minimum Pool Partitioning Problem (MPPP) Given: • primer pools set P and extensions sets Ep, for every primer p • probe set X • redundancy r Find: partition of P into the min number of strongly r-decodable subsets 48

MDPSP Maximum r-Decodable Pool Subset Problem (MDPSP) Given: • primer pools set P and extensions sets Ep, for every primer p • probe set X • redundancy r Find: • strongly r-decodable subset of P of maximum size 49

Min-Greedy Algorithm for Maximum Induced Matching in General Graphs • Pick a vertex u of min degree • Pick a vertex v of min degree from among u’s neighbors • Add edge (u, v) to the matching • Delete all neighbors of u and v from the graph • Repeat the above steps until the graph becomes empty • [Duckworth 05] d-1 approximation factor for d-regular graphs 50

Min-Greedy Algorithms for MDPSP • Bipartite hybridization graph G: – Primers in left side, probes in right side – Two types of edges: • N+(p)=Spec. X(p) • N-(p)=Spec. X(p, Ep) Spec. X(p) • Two algorithm variants: – Min. Primer. Greedy: pick primer first – Min. Probe. Greedy: pick probe first • Delete primer/probe if N+ degree drops below r/1 51

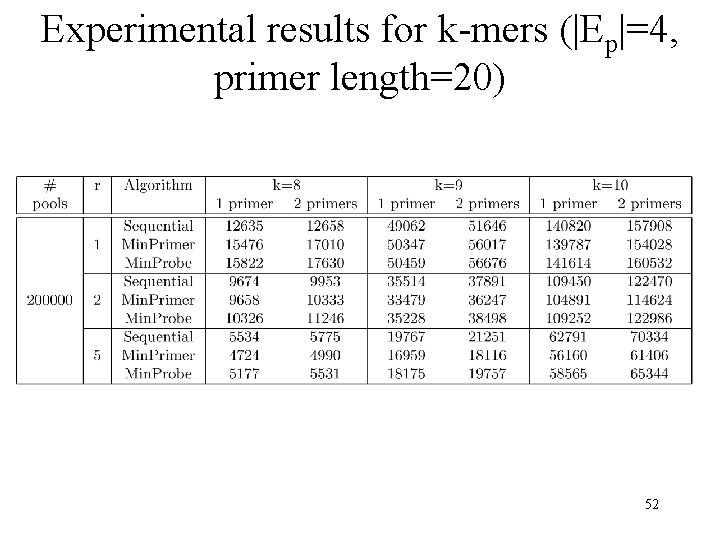
Experimental results for k-mers (|Ep|=4, primer length=20) 52

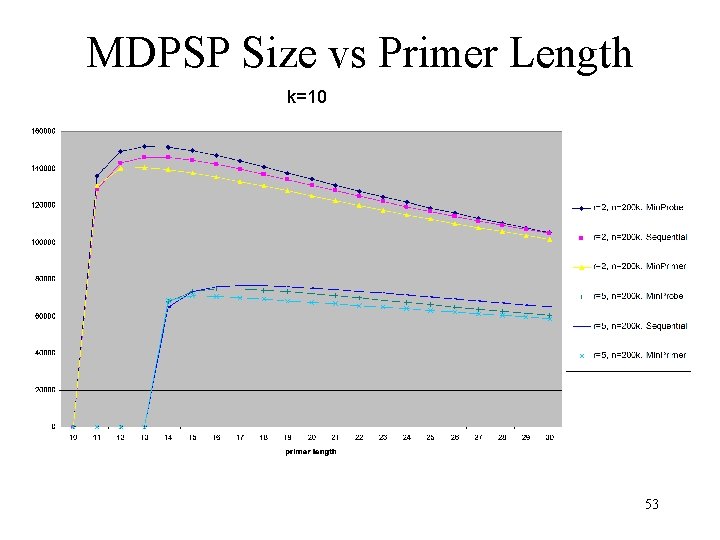
MDPSP Size vs Primer Length k=10 53

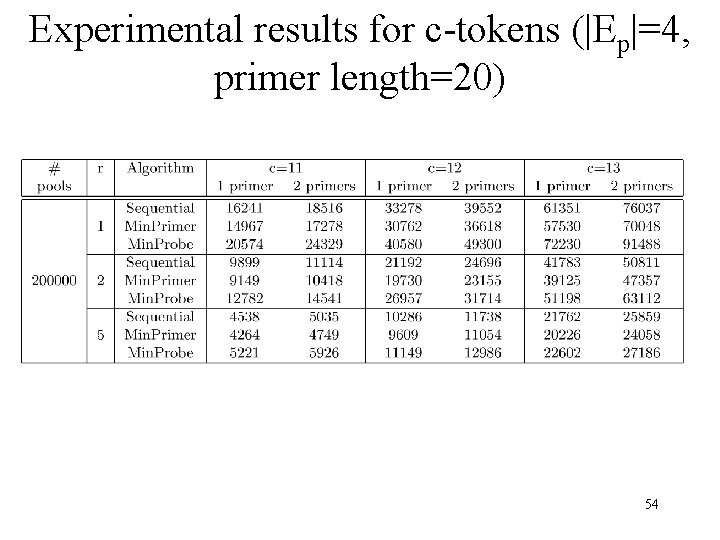
Experimental results for c-tokens (|Ep|=4, primer length=20) 54

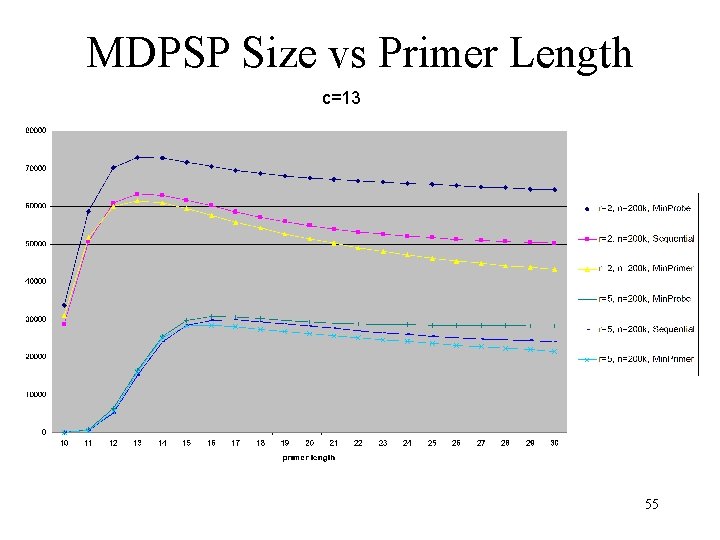
MDPSP Size vs Primer Length c=13 55

Overview m Tag Array Design - Tag Set Design - Tag Assignment Algorithms m SBE/SBH Assays - Decoding and Multiplexing Algorithms m Conclusions 56

Conclusions and Ongoing Work • Combinatorial algorithms yield significant increases in multiplexing rates of universal arrays – New SBE/SBH architecture particularly promising based on preliminary simulation results • Ongoing work: – Extend methods to more accurate hybridization models, e. g. , use NN melting temperature models – More complex (e. g. , temperature dependent) DNA tag set non-interaction requirements for DNA self/mediated assembly – Probabilistic decoding in presence of hybridization errors – Application to novel domains, e. g. , DNA barcoding 57

Acknowledgments • Claudia Prajescu and Dragos Trinca • Funding from NSF (Awards 0546457 and 0543365) and UCONN Research Foundation 58