Design and Optimization of Universal DNA Arrays Ion



![Hybridization Models • Hamming distance model, e. g. , [Marathe et al. 01] – Hybridization Models • Hamming distance model, e. g. , [Marathe et al. 01] –](https://slidetodoc.com/presentation_image/cb8ac03121f32e3f08dcb58462e5f002/image-6.jpg)

![Periodic Tags [MT 05] • Key observation: c-token uniqueness constraint in c-h code formulation Periodic Tags [MT 05] • Key observation: c-token uniqueness constraint in c-h code formulation](https://slidetodoc.com/presentation_image/cb8ac03121f32e3f08dcb58462e5f002/image-8.jpg)




- Slides: 16

Design and Optimization of Universal DNA Arrays Ion Mandoiu CSE Department & BME Program University of Connecticut 1

DNA Microarrays • Exploit Watson-Crick complementarity to simultaneously perform a large number of substring tests • Used in a variety of high-throughput genomic analyses – – Transcription (gene expression) analysis Single Nucleotide Polymorphism (SNP) genotyping Genomic-based microorganism identification Alternative splicing, Ch. IP-on-chip, tiling arrays, … • Common microarray formats involve direct hybridization between labeled DNA/RNA sample and DNA probes attached to a glass slide 2

Universal DNA Arrays • Limitations of direct hybridization formats: – Arrays of c. DNAs: inexpensive, but can only be used for transcription analysis – Oligonucleotide arrays: flexible, but expensive unless produced in large quantities • Universal DNA arrays: “programable” arrays – Array consists of application independent oligonucleotides – Detection carried by a sequence of reactions involving application specific primers – Flexible AND cost effective • Universal array architectures: DNA tag arrays, APEX arrays, SBE/SBH arrays 3

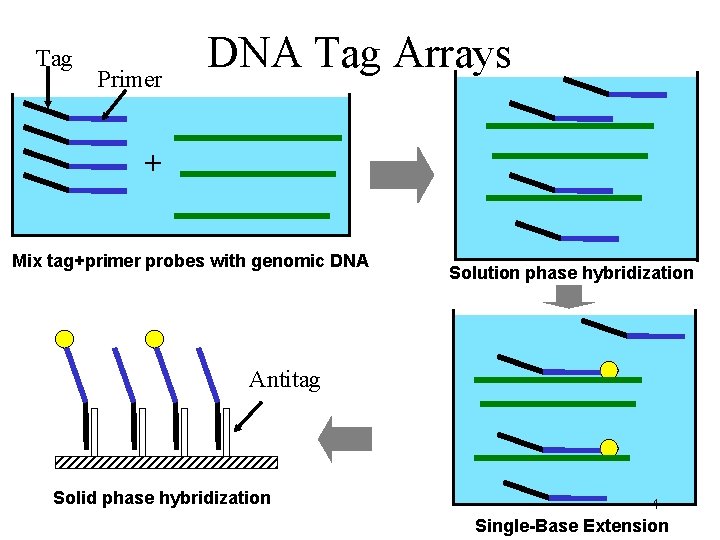
Tag Primer DNA Tag Arrays + Mix tag+primer probes with genomic DNA Solution phase hybridization Antitag Solid phase hybridization 4 Single-Base Extension

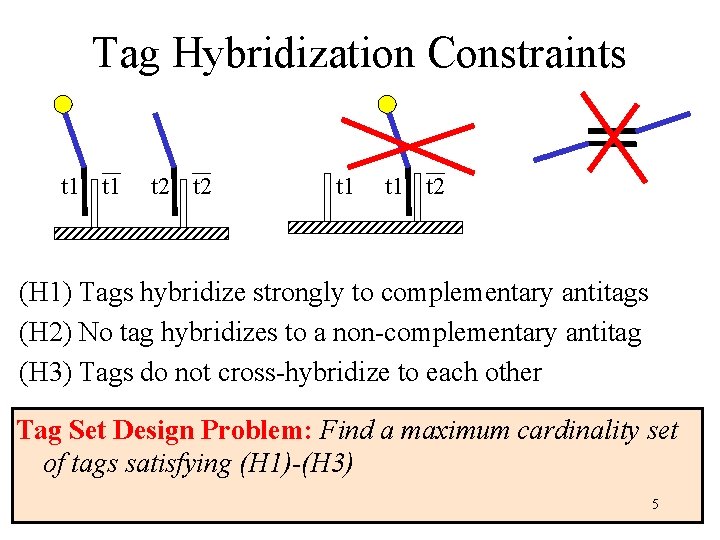
Tag Hybridization Constraints t 1 t 2 (H 1) Tags hybridize strongly to complementary antitags (H 2) No tag hybridizes to a non-complementary antitag (H 3) Tags do not cross-hybridize to each other Tag Set Design Problem: Find a maximum cardinality set of tags satisfying (H 1)-(H 3) 5
![Hybridization Models Hamming distance model e g Marathe et al 01 Hybridization Models • Hamming distance model, e. g. , [Marathe et al. 01] –](https://slidetodoc.com/presentation_image/cb8ac03121f32e3f08dcb58462e5f002/image-6.jpg)
Hybridization Models • Hamming distance model, e. g. , [Marathe et al. 01] – Models rigid DNA strands • LCS/edit distance model, e. g. , [Torney et al. 03] – Models infinitely elastic DNA strands • c-token model [Ben-Dor et al. 00]: – Duplex formation requires formation of nucleation complex between perfectly complementary substrings – Nucleation complex must have weight c, where wt(A)=wt(T)=1, wt(C)=wt(G)=2 (2 -4 rule) 6

c-h Code Problem • c-token: left-minimal DNA string of weight c, i. e. , – w(x) c – w(x’) < c for every proper suffix x’ of x • A set of tags is a c-h code if (C 1) Every tag has weight h (C 2) Every c-token is used at most once c-h Code Problem [Ben-Dor et al. 00] Given c and h, find maximum cardinality c-h code [Ben-Dor et al. 00] give approximation algorithm based on De. Bruijn sequences 7
![Periodic Tags MT 05 Key observation ctoken uniqueness constraint in ch code formulation Periodic Tags [MT 05] • Key observation: c-token uniqueness constraint in c-h code formulation](https://slidetodoc.com/presentation_image/cb8ac03121f32e3f08dcb58462e5f002/image-8.jpg)
Periodic Tags [MT 05] • Key observation: c-token uniqueness constraint in c-h code formulation is too strong – A c-token should not appear in two different tags, but can be repeated in a tag – Periodic tags use fewer c-tokens! Tag set design can be cast as a cycle packing problem 8

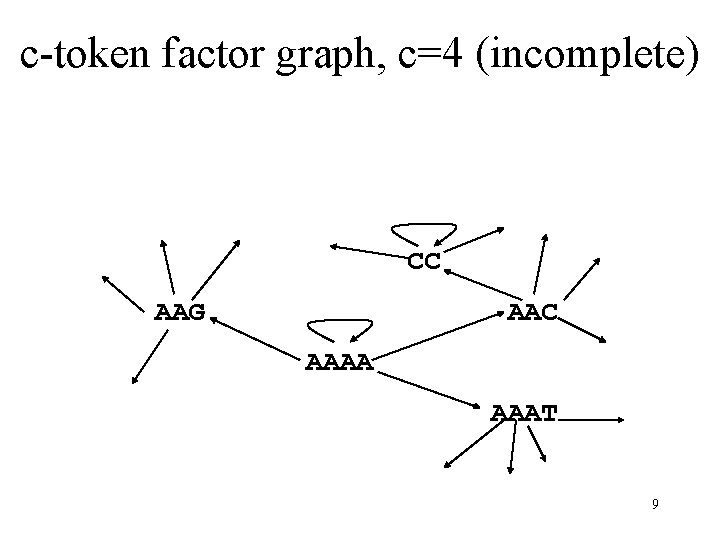
c-token factor graph, c=4 (incomplete) CC AAG AAC AAAA AAAT 9

Cycle Packing Algorithm 1. Construct c-token factor graph G 2. T {} 3. For all cycles C defining periodic tags, in increasing order of cycle length, • Add to T the tag defined by C • Remove C from G 4. Perform an alphabetic tree search and add to T tags consisting of unused c-tokens 5. Return T – Gives an increase of over 40% in the number of tags compared to previous methods 10

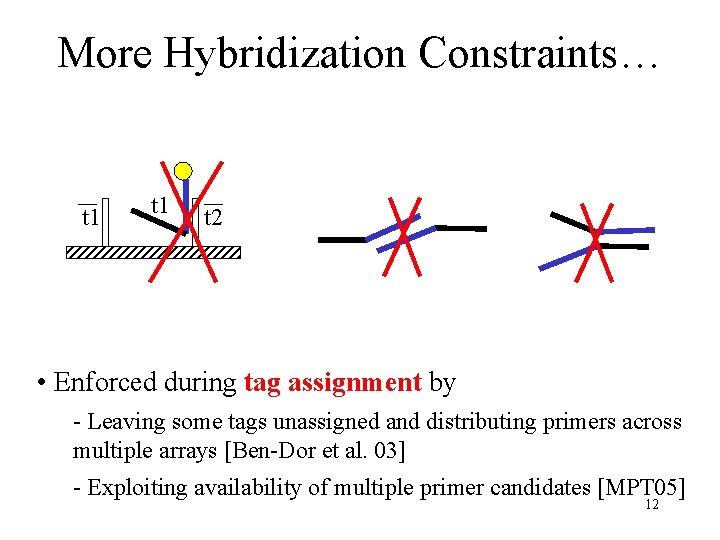
More Hybridization Constraints… t 1 t 2 • Enforced during tag assignment by - Leaving some tags unassigned and distributing primers across multiple arrays [Ben-Dor et al. 03] - Exploiting availability of multiple primer candidates [MPT 05] 12

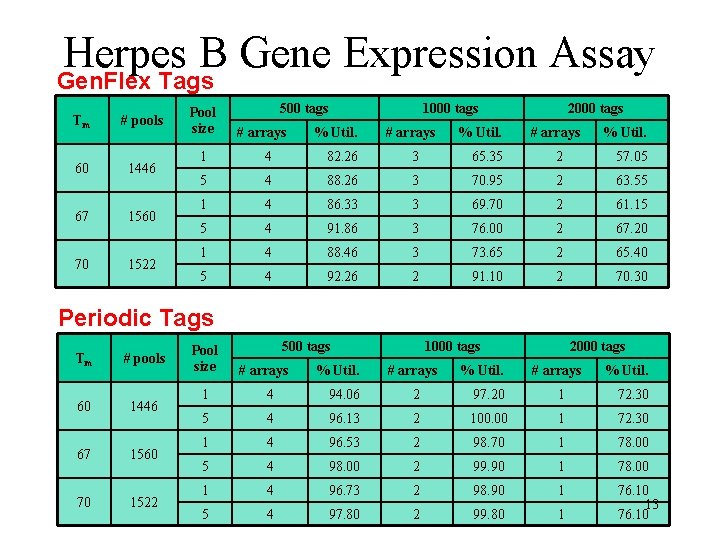
Herpes B Gene Expression Assay Gen. Flex Tags Tm # pools 60 1446 67 1560 70 1522 Pool size 500 tags # arrays % Util. 1000 tags # arrays % Util. 2000 tags # arrays % Util. 1 4 82. 26 3 65. 35 2 57. 05 5 4 88. 26 3 70. 95 2 63. 55 1 4 86. 33 3 69. 70 2 61. 15 5 4 91. 86 3 76. 00 2 67. 20 1 4 88. 46 3 73. 65 2 65. 40 5 4 92. 26 2 91. 10 2 70. 30 Periodic Tags Tm # pools 60 1446 67 1560 70 1522 Pool size 500 tags # arrays % Util. 1000 tags # arrays % Util. 2000 tags # arrays % Util. 1 4 94. 06 2 97. 20 1 72. 30 5 4 96. 13 2 100. 00 1 72. 30 1 4 96. 53 2 98. 70 1 78. 00 5 4 98. 00 2 99. 90 1 78. 00 1 4 96. 73 2 98. 90 1 5 4 97. 80 2 99. 80 1 76. 10 13 76. 10

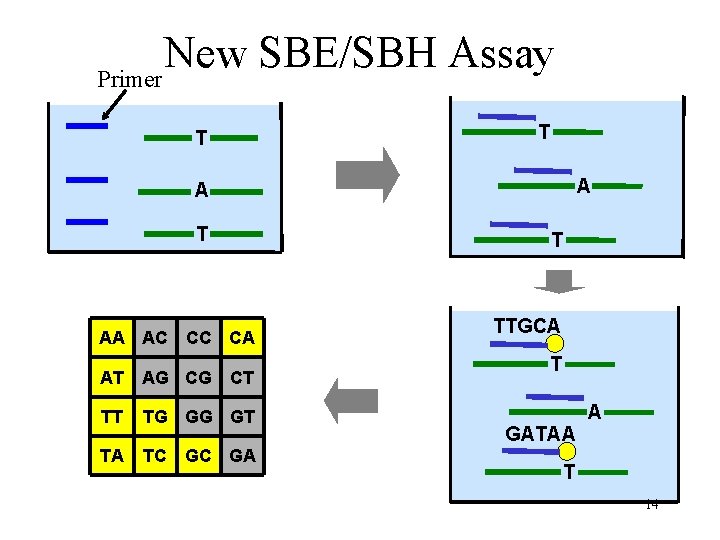
New SBE/SBH Assay Primer T T A A T T AA AC CC CA AT AG CG CT TT TG GG GT TA TC GC GA TTGCA T GATAA A T 14

SBE/SBH Throughput (c=13, r=5) See poster for more details 15

Conclusions and Ongoing Work • Combinatorial algorithms yield significant increases in multiplexing rates of universal DNA arrays – New SBE/SBH architecture particularly promising based on preliminary simulation results • Ongoing work: – Extend methods to more accurate hybridization models, e. g. , use NN melting temperature models – More complex (e. g. , temperature dependent) DNA tag set non-interaction requirements for DNA self/mediated assembly – Probabilistic decoding in presence of hybridization errors 16

Acknowledgments • UCONN Research Foundation • Claudia Prajescu • Dragos Trinca 17