Department of Animal Science Animal Breeding and Genetics

Department of Animal Science – Animal Breeding and Genetics Characterization of chicken cecal microbiome during acute and chronic heat stress John C. F. Hsieh, MPH Bioinformatics and Computational Biology Ph. D Candidate June 11, 2018 Metagenomics Mondays

Outline Thesis Project Introduction & Overview 2. 16 S r. RNA Analysis Considerations & Comparison 3. Metatranscriptome Analysis Considerations 1. Fayoumi Advanced Intercross Line Leghorn 2 Department of Animal Science Animal Breeding and Genetics

Thesis Project Introduction & Overview 3 Department of Animal Science Animal Breeding and Genetics
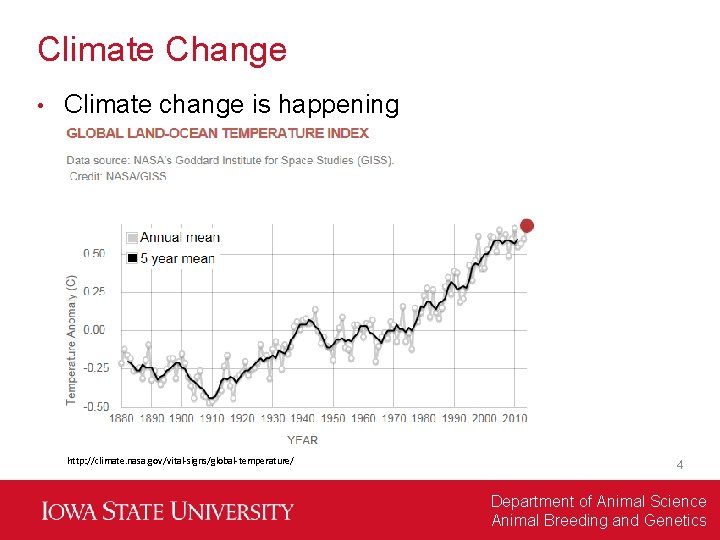
Climate Change • Climate change is happening http: //climate. nasa. gov/vital-signs/global-temperature/ 4 Department of Animal Science Animal Breeding and Genetics

Heat Stress in Chicken • Chickens don’t like it too hot… http: //www. heatstress. info/Heat. Stress. Explained/Howtorecognise. Heat. Stressinpoultry. aspx http: //www. heatstress. info/portals/38/poutry%20 heated. JPG 5 Department of Animal Science Animal Breeding and Genetics

Heat Stress and Microbiome Heat stress-induced dysfunction of the gut may lead to endotoxemia and poor feed efficiency • Dietary supplementation is one approach to alleviate the negative impacts of heat stress • Probiotics • Prebiotics • Essential Oils • Organic Acids • • However, little is known of the effect of heat on the gut microbiome https: //www. nurymartinez. com/facts/taking-probiotics-with-antibiotics-is-it-a-good-idea. html 6 Department of Animal Science Animal Breeding and Genetics

Heat Stress and Intestinal Health • Current studies of heat stress on intestinal health in chickens focuses on 3 aspects: 1. Gut Morphology 2. Mucosal Immunity Deng et al. , 2012 Brown et al. , 2013 3. Microbial Community Author (Year) Technique Stress Location Lan et. al. (2004) TRFLP AHS Jejunum/Cecum Burkholder et. al. (2008) PCR-DGGE AHS Ileum Sohail et. al. (2015) 16 S r. RNA CHS Trachea/Cecum Note: All studies were done on Broilers 7 Department of Animal Science Animal Breeding and Genetics
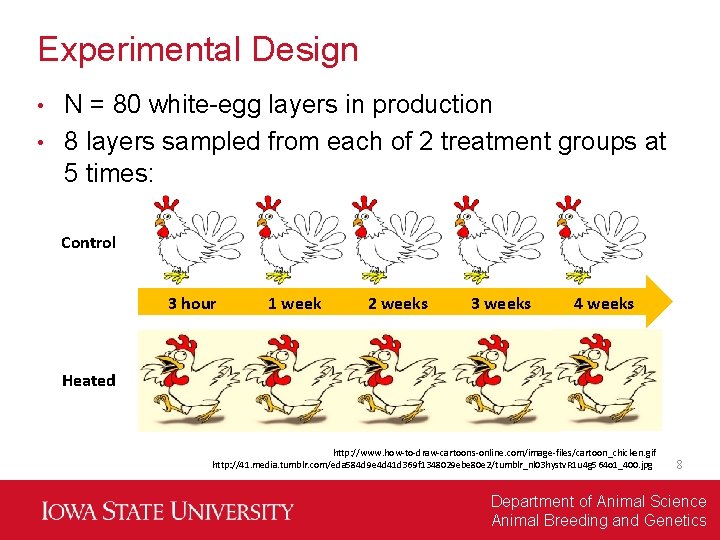
Experimental Design N = 80 white-egg layers in production • 8 layers sampled from each of 2 treatment groups at 5 times: • Control 3 hour 1 week 2 weeks 3 weeks 4 weeks Heated http: //www. how-to-draw-cartoons-online. com/image-files/cartoon_chicken. gif http: //41. media. tumblr. com/eda 584 d 9 e 4 d 41 d 369 f 1348029 ebe 80 e 2/tumblr_nl 03 hystv. R 1 u 4 g 564 o 1_400. jpg 8 Department of Animal Science Animal Breeding and Genetics
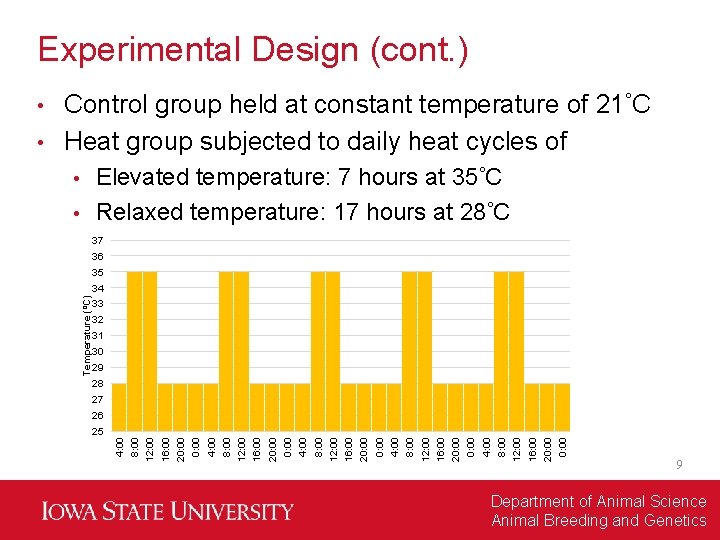
Experimental Design (cont. ) Control group held at constant temperature of 21 C • Heat group subjected to daily heat cycles of • Elevated temperature: 7 hours at 35 C • Relaxed temperature: 17 hours at 28 C • 37 36 35 33 32 31 30 29 28 27 0: 00 20: 00 16: 00 12: 00 8: 00 4: 00 0: 00 20: 00 16: 00 12: 00 25 8: 00 26 4: 00 Temperature (⁰C) 34 9 Department of Animal Science Animal Breeding and Genetics
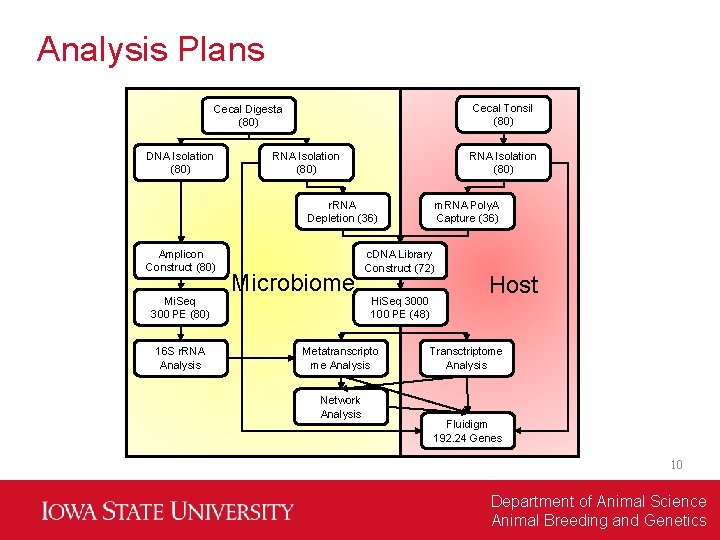
Analysis Plans Cecal Tonsil (80) Cecal Digesta (80) DNA Isolation (80) RNA Isolation (80) r. RNA Depletion (36) Amplicon Construct (80) Mi. Seq 300 PE (80) 16 S r. RNA Analysis Microbiome c. DNA Library Construct (72) Hi. Seq 3000 100 PE (48) Metatranscripto me Analysis Network Analysis m. RNA Poly. A Capture (36) Host Transctriptome Analysis Fluidigm 192. 24 Genes 10 Department of Animal Science Animal Breeding and Genetics

16 S r. RNA Analysis Considerations & Comparison 11 Department of Animal Science Animal Breeding and Genetics

Introduction • A little history of 16 S r. RNA 1990, Woese et al. suggested the use of r. RNA to classify organisms • 1997, Liu et al. used terminal restriction fragment length polymorphisms (TRFLP) to characterize micrbiome based on 16 S r. RNA • 2006, Sogin et al. used 454 pyrosequencing of V 6 variable region from ocean samples • • Significance of 16 S r. RNA Before: culturing (with problems…) • Now: extract DNA and PCR amplicons for sequencing • 12 Department of Animal Science Animal Breeding and Genetics

Analysis Consideration #1: Database • 16 S r. RNA variable regions V 1 -V 9 regions • Conserved regions between variable regions • Databases: • RDP: Ribosomal Database Project • GG: Green. Genes • SILVA: (from Latin silva = forest) • Adapted from Yarza et al. , 2014 13 Department of Animal Science Animal Breeding and Genetics
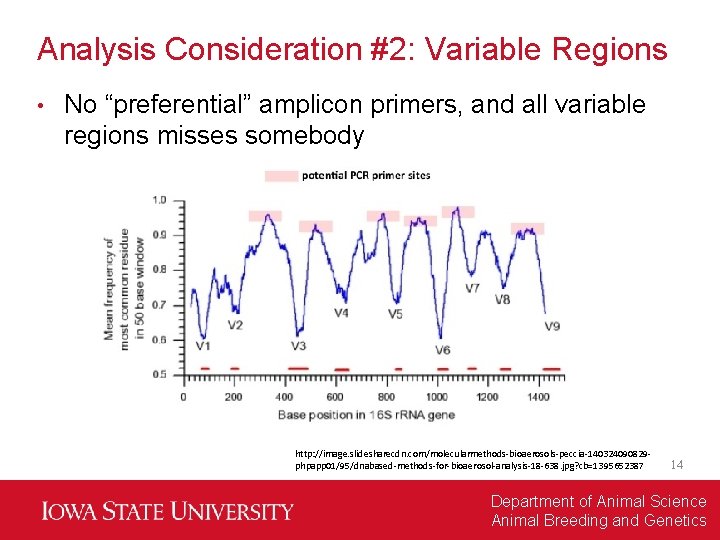
Analysis Consideration #2: Variable Regions • No “preferential” amplicon primers, and all variable regions misses somebody http: //image. slidesharecdn. com/molecularmethods-bioaerosols-peccia-140324090829 phpapp 01/95/dnabased-methods-for-bioaerosol-analysis-18 -638. jpg? cb=1395652387 14 Department of Animal Science Animal Breeding and Genetics

Analysis Consideration #3: Analysis Pipeline • “Best” analysis tool/pipeline doesn’t exist, and each gives a slightly different result QIIME (http: //qiime. org/) • mothur (http: //www. mothur. org/) • MG-RAST (http: //metagenomics. anl. gov/) • 15 Department of Animal Science Animal Breeding and Genetics

Analysis Consideration #4: Resolution • A little more history… 2007, Janda et al. pointed out Genus/species identification problems with 16 S r. RNA sequencing data, suggested improvement of databases • 2010 s, numerous shotgun metagenomic studies showed discrepancies of Genus/species identification • • How to get higher resolution? More reads (Shotgun WGS) • Longer reads (Pac. Bio SMRT/Nanopore) • Better targets (Affymetrix Array? ) • 16 Department of Animal Science Animal Breeding and Genetics
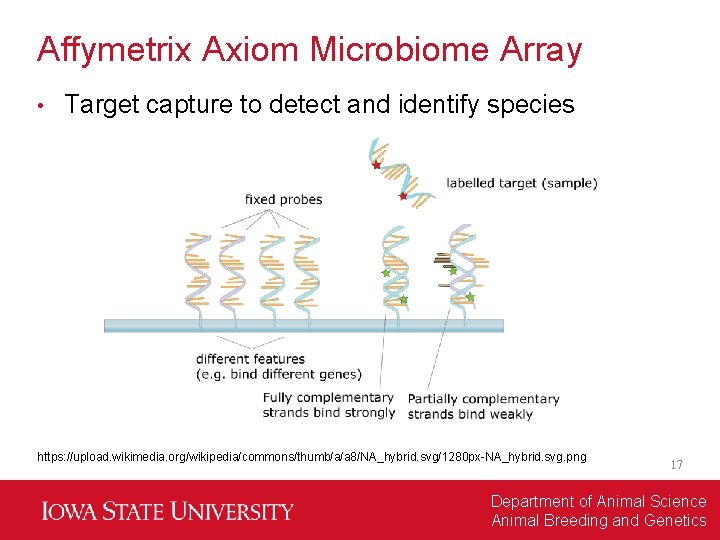
Affymetrix Axiom Microbiome Array • Target capture to detect and identify species https: //upload. wikimedia. org/wikipedia/commons/thumb/a/a 8/NA_hybrid. svg/1280 px-NA_hybrid. svg. png 17 Department of Animal Science Animal Breeding and Genetics
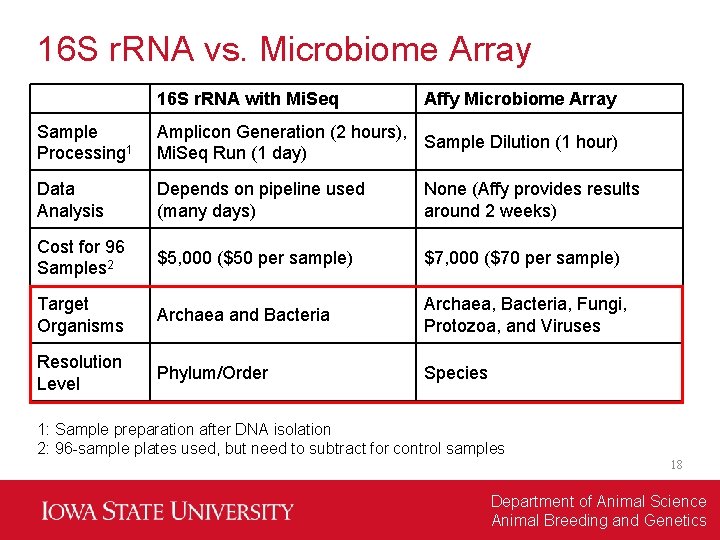
16 S r. RNA vs. Microbiome Array 16 S r. RNA with Mi. Seq Affy Microbiome Array Sample Processing 1 Amplicon Generation (2 hours), Sample Dilution (1 hour) Mi. Seq Run (1 day) Data Analysis Depends on pipeline used (many days) None (Affy provides results around 2 weeks) Cost for 96 Samples 2 $5, 000 ($50 per sample) $7, 000 ($70 per sample) Target Organisms Archaea and Bacteria Archaea, Bacteria, Fungi, Protozoa, and Viruses Resolution Level Phylum/Order Species 1: Sample preparation after DNA isolation 2: 96 -sample plates used, but need to subtract for control samples 18 Department of Animal Science Animal Breeding and Genetics
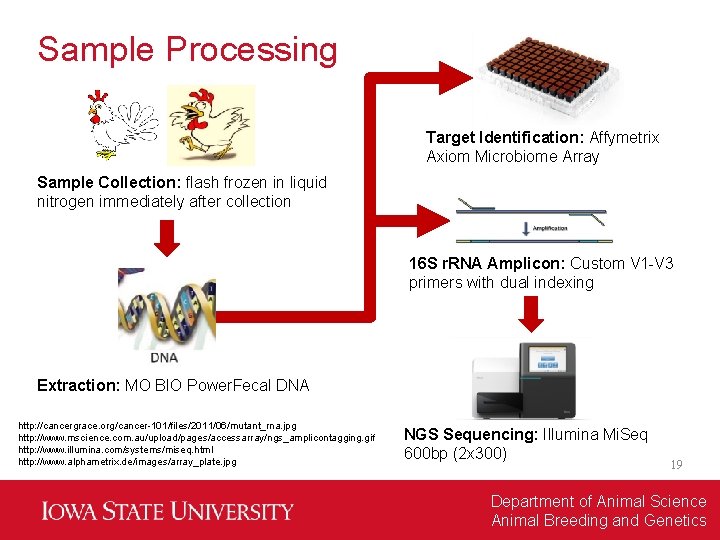
Sample Processing Target Identification: Affymetrix Axiom Microbiome Array Sample Collection: flash frozen in liquid nitrogen immediately after collection 16 S r. RNA Amplicon: Custom V 1 -V 3 primers with dual indexing Extraction: MO BIO Power. Fecal DNA http: //cancergrace. org/cancer-101/files/2011/06/mutant_rna. jpg http: //www. mscience. com. au/upload/pages/accessarray/ngs_amplicontagging. gif http: //www. illumina. com/systems/miseq. html http: //www. alphametrix. de/images/array_plate. jpg NGS Sequencing: Illumina Mi. Seq 600 bp (2 x 300) 19 Department of Animal Science Animal Breeding and Genetics

Analysis Pipelines • Pipeline 1: Affymetrix microarray results are analyzed with Mi. DAS (Microbial Detection Analysis Software) • Pipeline 2: QIIME (https: //nephele. niaid. nih. gov) • • Pipeline 3: mothur (https: //www. mothur. org) • • Database: SILVA Pipeline 4: MG-RAST (https: //metagenomics. anl. gov) • Database: M 5 nr 20 Department of Animal Science Animal Breeding and Genetics

Microbiome Array Replicability • DNA samples (n=6) were profiled twice on the Affymetrix Axiom Microbiome Array • A C B • • • Agreement % = C / (A+B+C) • N = 6 pairs of technical replicates Min: 54. 84% Average: 62. 86% Max: 76. 00% SD: 0. 0849 Technical replicates only had 60% agreement based on the microbiome array 21 Department of Animal Science Animal Breeding and Genetics
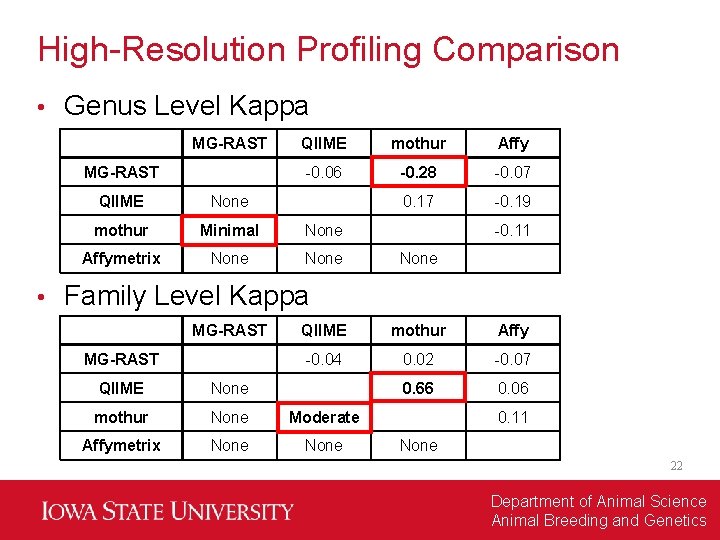
High-Resolution Profiling Comparison • Genus Level Kappa MG-RAST • QIIME mothur Affy -0. 06 -0. 28 -0. 07 0. 17 -0. 19 QIIME None mothur Minimal None Affymetrix None -0. 11 None Family Level Kappa MG-RAST QIIME mothur Affy -0. 04 0. 02 -0. 07 0. 66 0. 06 QIIME None mothur None Moderate Affymetrix None 0. 11 None 22 Department of Animal Science Animal Breeding and Genetics

Metatranscriptome Analysis 23 Department of Animal Science Animal Breeding and Genetics
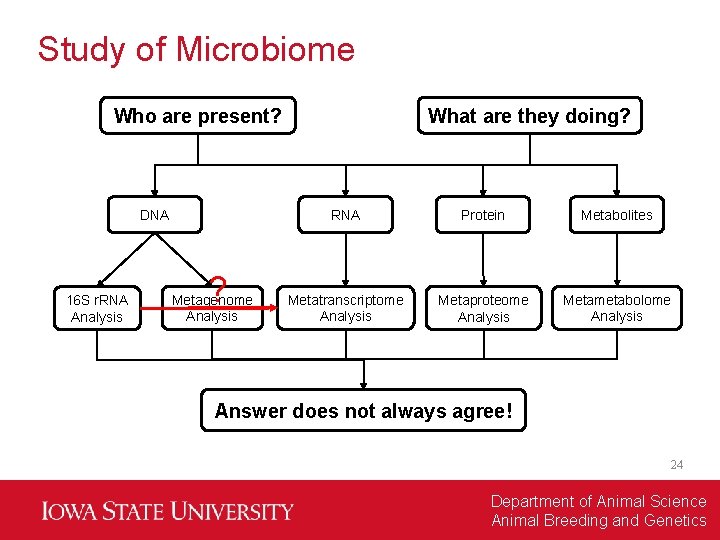
Study of Microbiome Who are present? DNA 16 S r. RNA Analysis ? Metagenome Analysis What are they doing? RNA Protein Metabolites Metatranscriptome Analysis Metaproteome Analysis Metametabolome Analysis Answer does not always agree! 24 Department of Animal Science Animal Breeding and Genetics
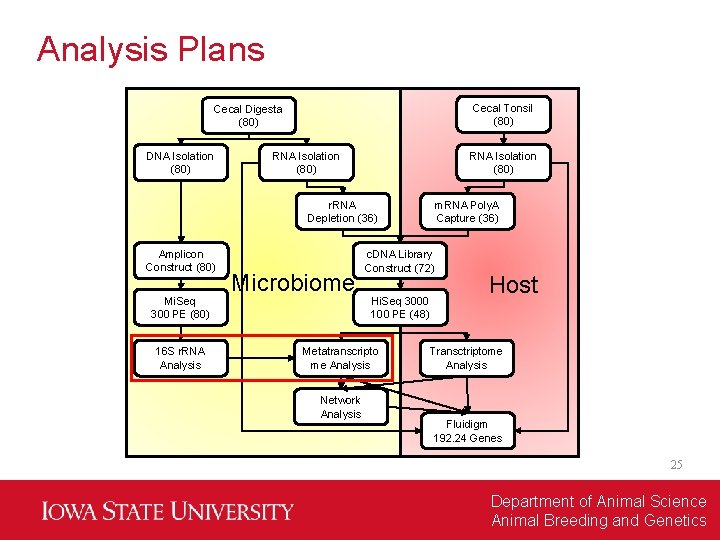
Analysis Plans Cecal Tonsil (80) Cecal Digesta (80) DNA Isolation (80) RNA Isolation (80) r. RNA Depletion (36) Amplicon Construct (80) Mi. Seq 300 PE (80) 16 S r. RNA Analysis Microbiome c. DNA Library Construct (72) Hi. Seq 3000 100 PE (48) Metatranscripto me Analysis Network Analysis m. RNA Poly. A Capture (36) Host Transctriptome Analysis Fluidigm 192. 24 Genes 25 Department of Animal Science Animal Breeding and Genetics

Major Crisis… Where are the protocols for doing metatranscriptome analysis? 26 Department of Animal Science Animal Breeding and Genetics
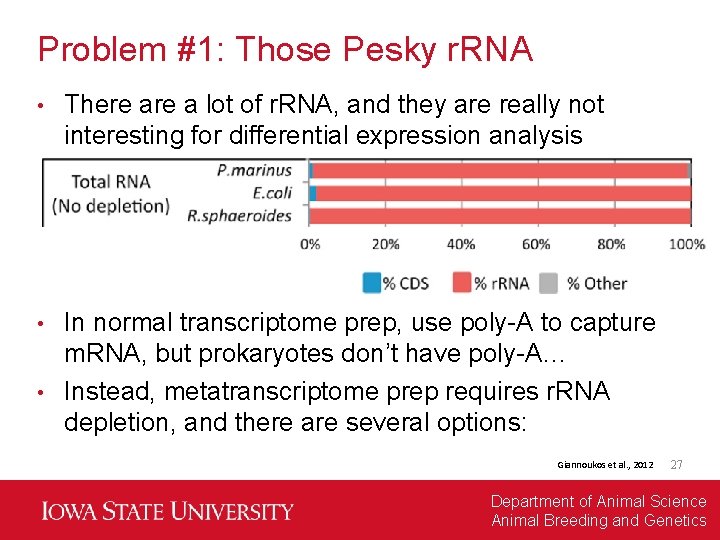
Problem #1: Those Pesky r. RNA • There a lot of r. RNA, and they are really not interesting for differential expression analysis In normal transcriptome prep, use poly-A to capture m. RNA, but prokaryotes don’t have poly-A… • Instead, metatranscriptome prep requires r. RNA depletion, and there are several options: • Giannoukos et al. , 2012 27 Department of Animal Science Animal Breeding and Genetics
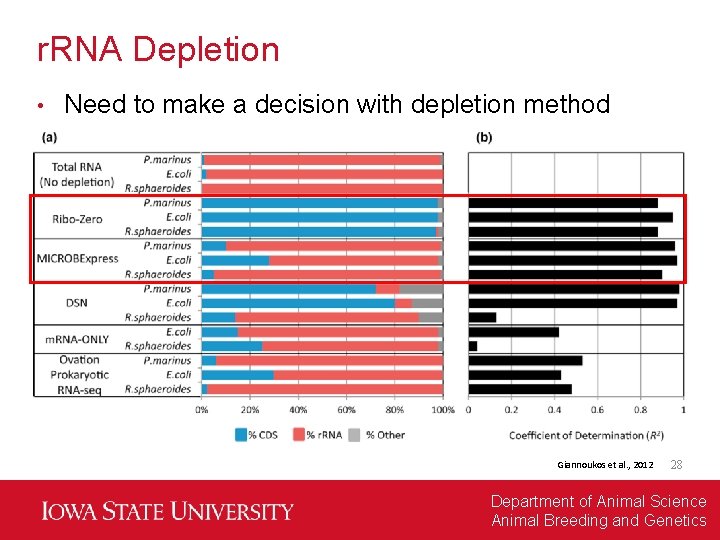
r. RNA Depletion • Need to make a decision with depletion method Giannoukos et al. , 2012 28 Department of Animal Science Animal Breeding and Genetics
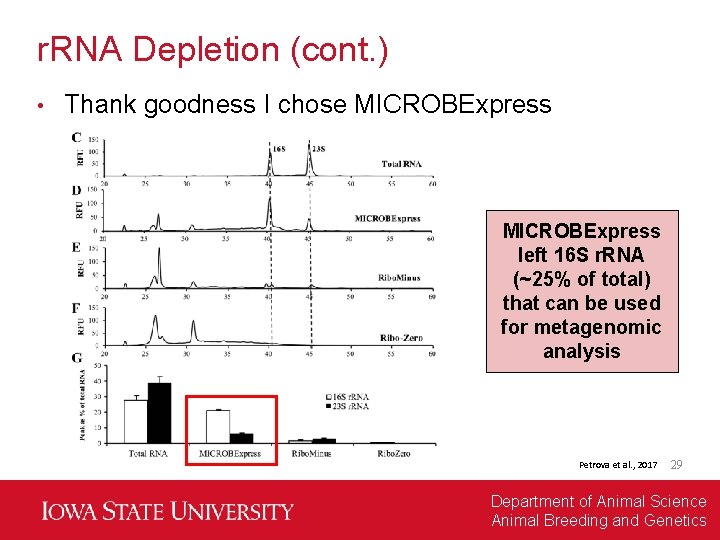
r. RNA Depletion (cont. ) • Thank goodness I chose MICROBExpress left 16 S r. RNA (~25% of total) that can be used for metagenomic analysis Petrova et al. , 2017 29 Department of Animal Science Animal Breeding and Genetics
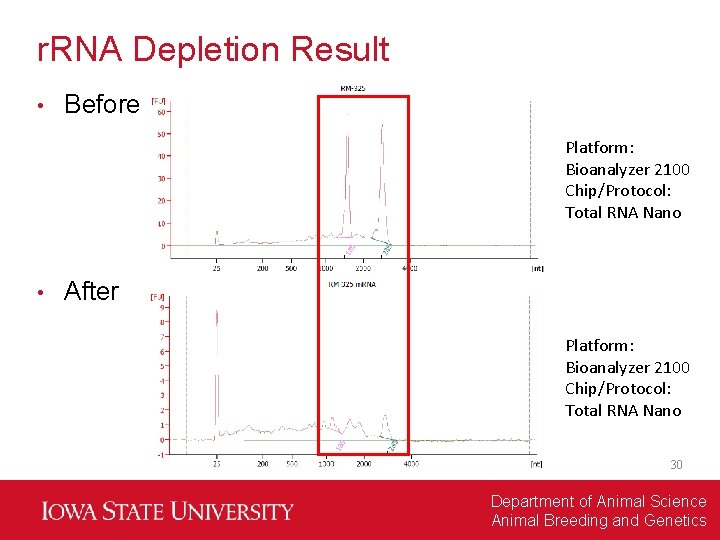
r. RNA Depletion Result • Before Platform: Bioanalyzer 2100 Chip/Protocol: Total RNA Nano • After Platform: Bioanalyzer 2100 Chip/Protocol: Total RNA Nano 30 Department of Animal Science Animal Breeding and Genetics
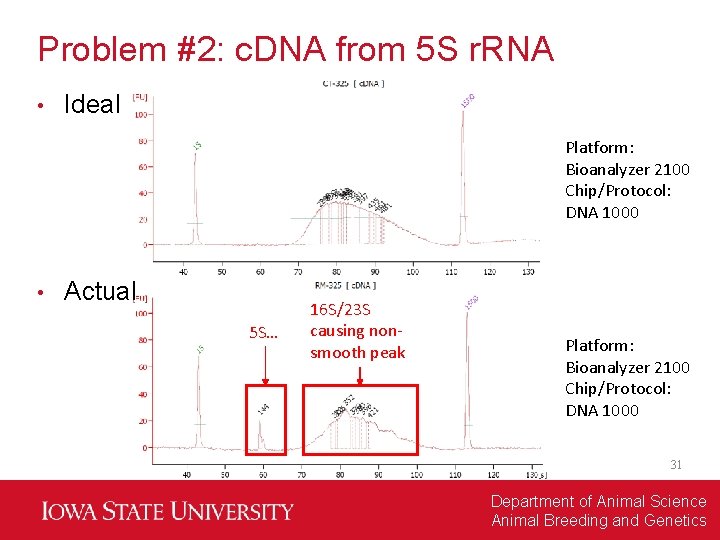
Problem #2: c. DNA from 5 S r. RNA • Ideal Platform: Bioanalyzer 2100 Chip/Protocol: DNA 1000 • Actual 5 S… 16 S/23 S causing nonsmooth peak Platform: Bioanalyzer 2100 Chip/Protocol: DNA 1000 31 Department of Animal Science Animal Breeding and Genetics
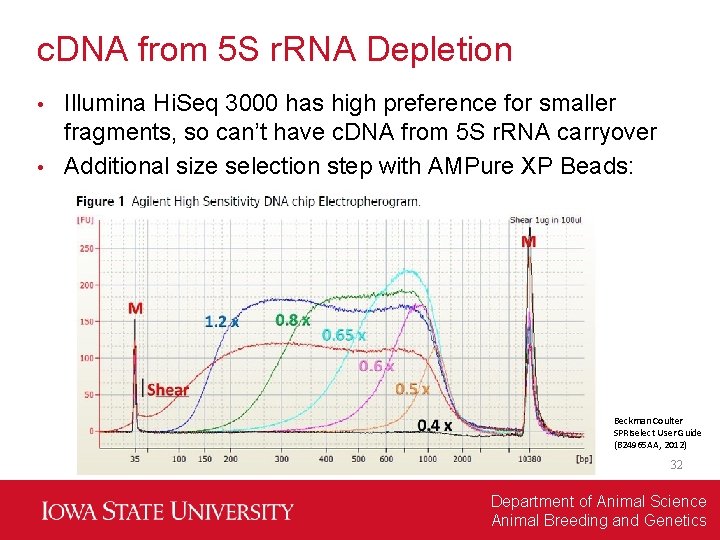
c. DNA from 5 S r. RNA Depletion Illumina Hi. Seq 3000 has high preference for smaller fragments, so can’t have c. DNA from 5 S r. RNA carryover • Additional size selection step with AMPure XP Beads: • Beckman Coulter SPRIselect User Guide (B 24965 AA, 2012) 32 Department of Animal Science Animal Breeding and Genetics
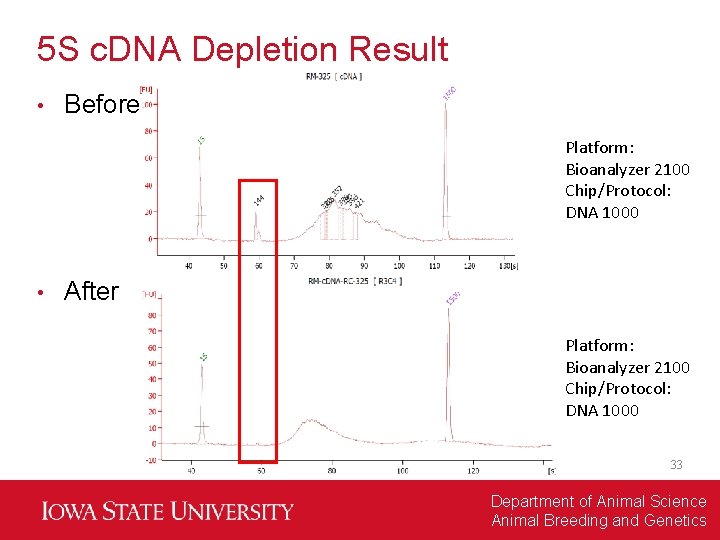
5 S c. DNA Depletion Result • Before Platform: Bioanalyzer 2100 Chip/Protocol: DNA 1000 • After Platform: Bioanalyzer 2100 Chip/Protocol: DNA 1000 33 Department of Animal Science Animal Breeding and Genetics
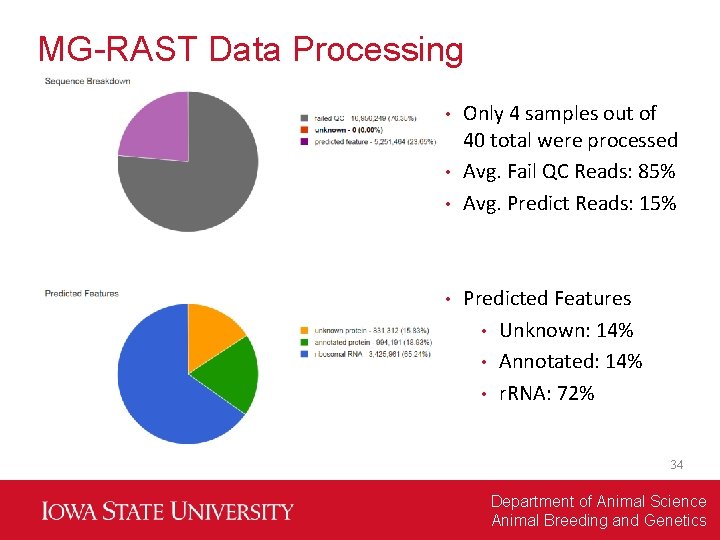
MG-RAST Data Processing Only 4 samples out of 40 total were processed • Avg. Fail QC Reads: 85% • Avg. Predict Reads: 15% • • Predicted Features • Unknown: 14% • Annotated: 14% • r. RNA: 72% 34 Department of Animal Science Animal Breeding and Genetics
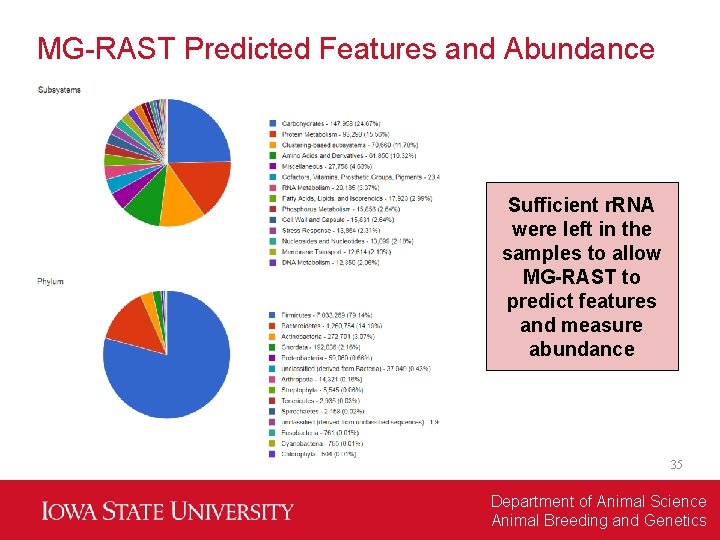
MG-RAST Predicted Features and Abundance Sufficient r. RNA were left in the samples to allow MG-RAST to predict features and measure abundance 35 Department of Animal Science Animal Breeding and Genetics

Problem #3: Differential Expression Analysis • Bioinformatics problem with metatranscriptome differential expression analysis Membership Change Expression Change vs. • Potential solution: incorporate microbiome membership information from 16 S r. RNA 36 Department of Animal Science Animal Breeding and Genetics
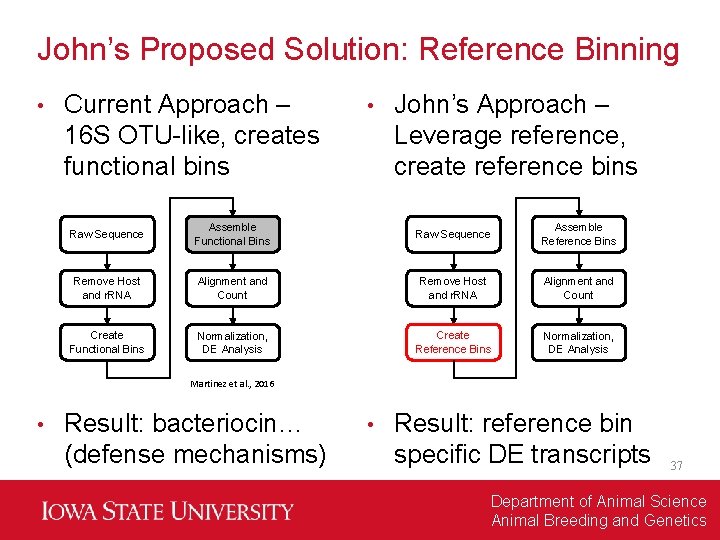
John’s Proposed Solution: Reference Binning • Current Approach – 16 S OTU-like, creates functional bins • John’s Approach – Leverage reference, create reference bins Raw Sequence Assemble Functional Bins Raw Sequence Assemble Reference Bins Remove Host and r. RNA Alignment and Count Create Functional Bins Normalization, DE Analysis Create Reference Bins Normalization, DE Analysis Martinez et al. , 2016 • Result: bacteriocin… (defense mechanisms) • Result: reference bin specific DE transcripts 37 Department of Animal Science Animal Breeding and Genetics

Reference Binning Method for DE Analysis • • Step 1: QC Fasta Files (Fast. QC, Trimmomatic) Step 2: Remove Host m. RNA (STAR) Step 3: Remove r. RNA (Sort. Me. RNA) Step 4: Pool Samples and Create Reference Bins (Kraken) Step 5: Assemble Reference Bins (Trinity) Step 6: Align Samples to Each Assembly (BWA) Step 7: Count Features by Sample (HT-Count) Step 8: Normalization (both abundance and library size) and DE Analysis (DESeq 2) Raw Sequence Remove Host and r. RNA Create Reference Bins Assemble Reference Bins Alignment and Count Normalization, DE Analysis 38 Department of Animal Science Animal Breeding and Genetics

Summary (Take-Home Messages) Bioinformatics for microbiome analysis is still being researched and developed, so be aware that there is no “one-size fits all” solution (as with most bioinformatics) • 16 S r. RNA analysis has a lot of tools and resources currently available, but be cognizant of choices made for 16 S r. RNA database, variable region sequenced, analysis pipeline, and interpretation of results • Metatranscriptome analysis is still at its infancy, but can provide immensely valuable information if the data is generated analyzed carefully • 39 Department of Animal Science Animal Breeding and Genetics

Acknowledgements • POSC Members • Susan J. Lamont • Drena L. Dobbs • James M. Reecy • Karin Dorman • Peng Liu Project PIs • Carl J. Schmidt • Michael E. Persia • Chris M. Ashwell • Max F. Rothschild • Special Thanks • Torey Looft • This work is supported by USDA Agriculture and Food Research Initiative Competitive Grant #2011 -67003 -30228 Department of Animal Science Animal Breeding and Genetics
- Slides: 40