Dealing With the Unknown Metabolomics Ben Bowen Metabolite

Dealing With the Unknown Metabolomics & Ben Bowen Metabolite Atlases Pathway Tools Workshop 2010

Acknowledgements Trent Northen Richard Baran Wolfgang Reindl Do Yup Lee Jane Tanamachi Jill Banfield Curt Fisher Paul Wilmes US Department of Energy BER Genome Sciences Program
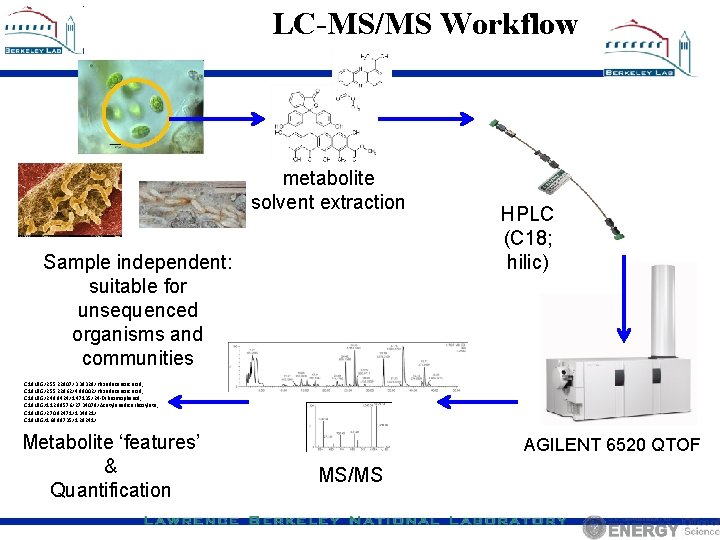
LC-MS/MS Workflow metabolite solvent extraction Sample independent: suitable for unsequenced organisms and communities HPLC (C 18; hilic) C 18 NEG/255. 22807/3. 39329/Hexadecanoic acid; C 18 NEG/255. 22862/4. 89002/Hexadecanoic acid; C 18 NEG/248. 8424/1. 47135/24 -Dibromophenol; C 18 NEG/112. 98576/27. 34079/Acetylenedicarboxylate; C 18 NEG/270. 82471/1. 34821/ C 18 NEG/168. 88735/1. 29241/ Metabolite ‘features’ & Quantification AGILENT 6520 QTOF MS/MS
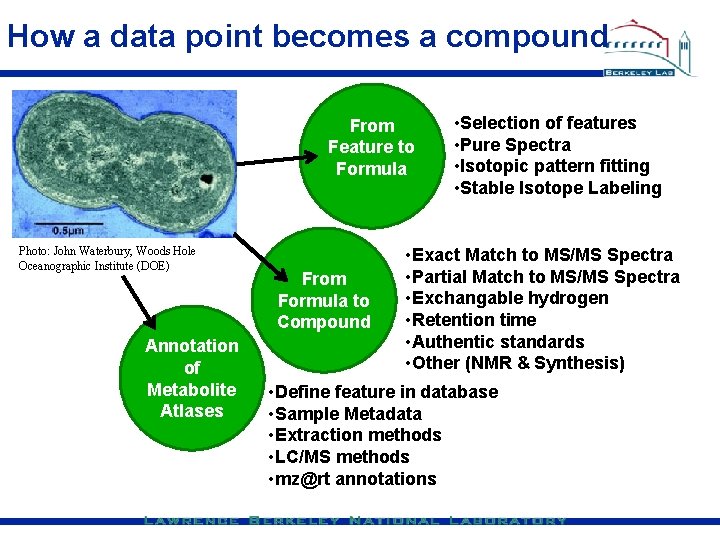
How a data point becomes a compound From Feature to Formula Photo: John Waterbury, Woods Hole Oceanographic Institute (DOE) Annotation of Metabolite Atlases From Formula to Compound • Selection of features • Pure Spectra • Isotopic pattern fitting • Stable Isotope Labeling • Exact Match to MS/MS Spectra • Partial Match to MS/MS Spectra • Exchangable hydrogen • Retention time • Authentic standards • Other (NMR & Synthesis) • Define feature in database • Sample Metadata • Extraction methods • LC/MS methods • mz@rt annotations
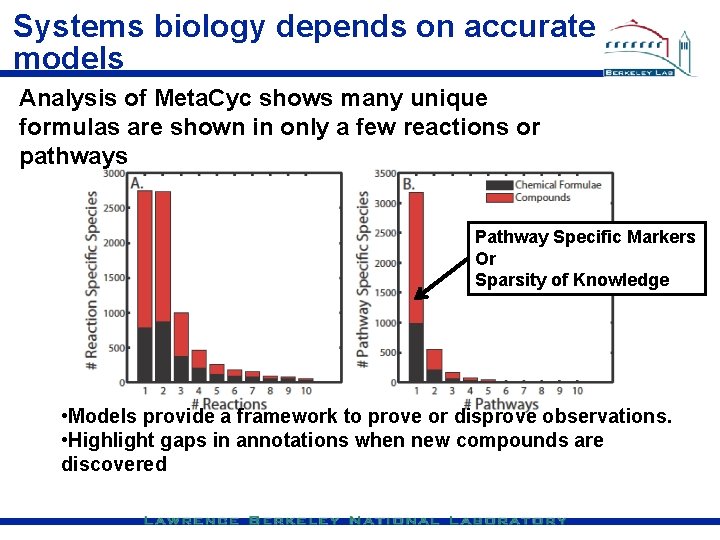
Systems biology depends on accurate models Analysis of Meta. Cyc shows many unique formulas are shown in only a few reactions or pathways Pathway Specific Markers Or Sparsity of Knowledge • Models provide a framework to prove or disprove observations. • Highlight gaps in annotations when new compounds are discovered
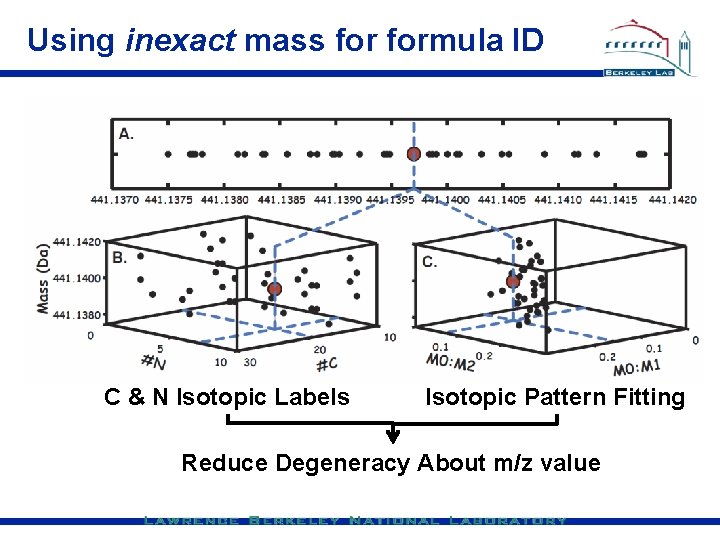
Using inexact mass formula ID C & N Isotopic Labels Isotopic Pattern Fitting Reduce Degeneracy About m/z value
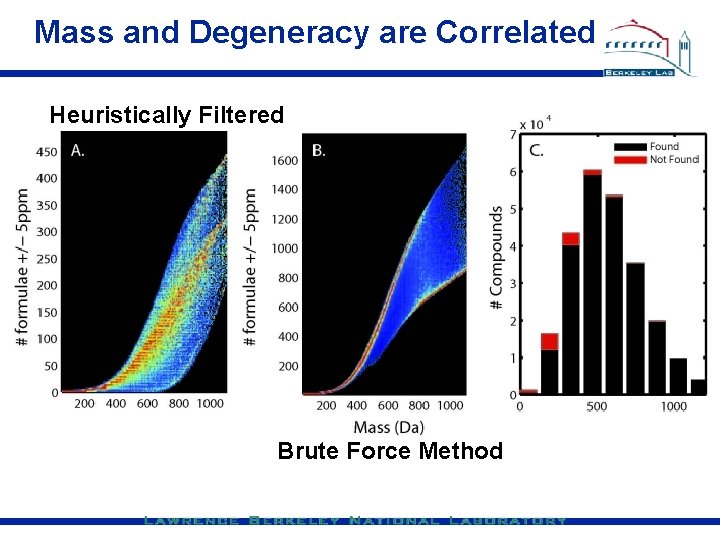
Mass and Degeneracy are Correlated Heuristically Filtered Brute Force Method
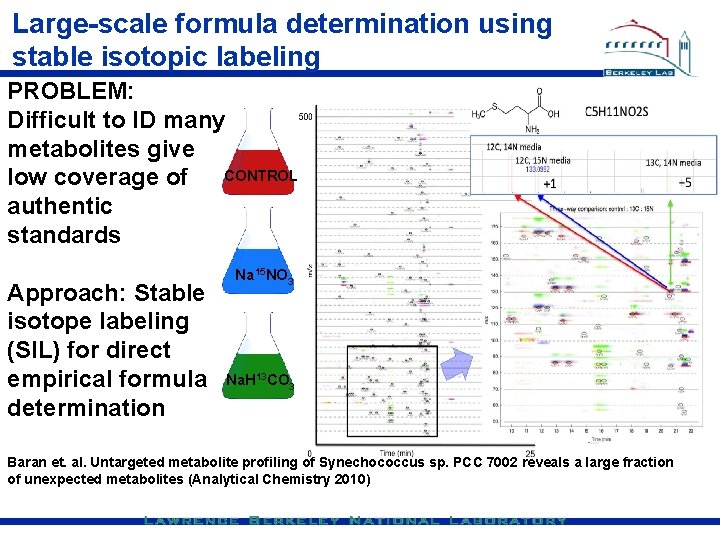
Large-scale formula determination using stable isotopic labeling PROBLEM: Difficult to ID many metabolites give low coverage of CONTROL authentic standards Approach: Stable isotope labeling (SIL) for direct empirical formula determination Na 15 NO 3 Na. H 13 CO 3 Baran et. al. Untargeted metabolite profiling of Synechococcus sp. PCC 7002 reveals a large fraction of unexpected metabolites (Analytical Chemistry 2010)
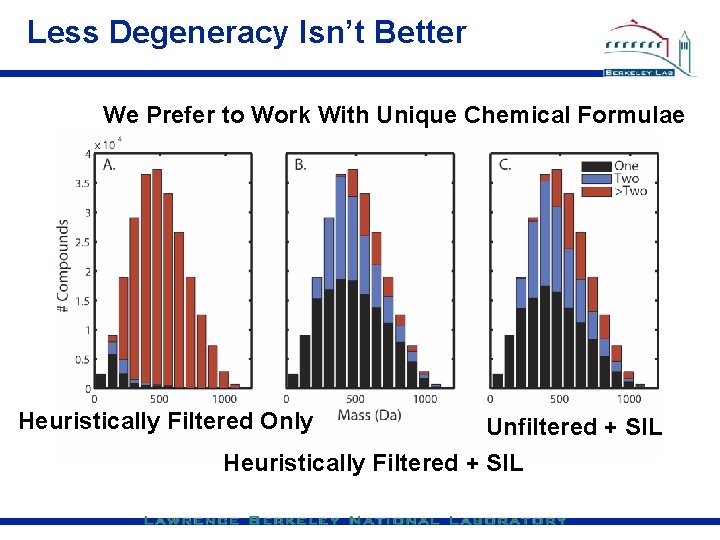
Less Degeneracy Isn’t Better We Prefer to Work With Unique Chemical Formulae Heuristically Filtered Only Unfiltered + SIL Heuristically Filtered + SIL
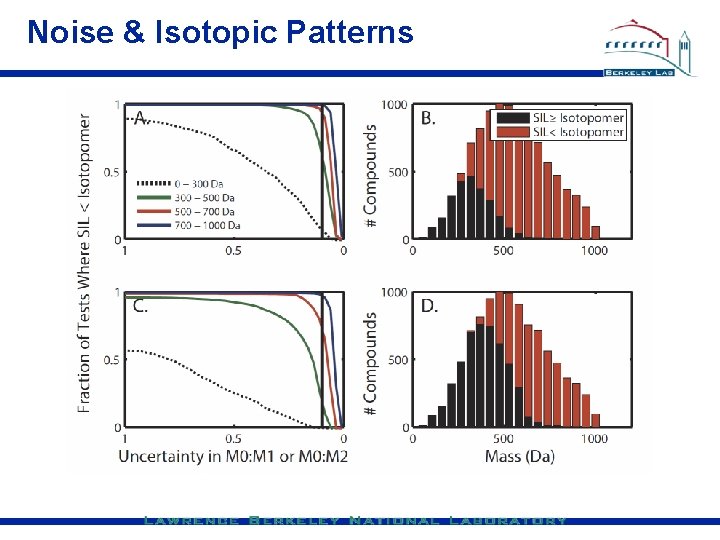
Noise & Isotopic Patterns

Initial focus is on Synechococcus sp a simple yet important model system Simple system For method Widely distributed and globally important in carbon cycling development 1. Photosynthetic bacteria 2. Small genome (3299 ORFs) 3. ~fast growing and easy to grow 4. No metabolite background (salt media) 5. Adaptable: 0 -2 M salt, T up to 45 C

Benefits of Using SIL • Are the signals being measured biological? • What type of ion is the signal? • Has this signal been seen before? • What compound(s) is it? • What else in the sample behaves like that compound? Global Profiling SIL Standards
![Stable isotope labeling Control [15 N]Na. NO 3 15 N [13 C]Na. HCO 3 Stable isotope labeling Control [15 N]Na. NO 3 15 N [13 C]Na. HCO 3](http://slidetodoc.com/presentation_image_h2/85f57791f12f86c05935f69009127b7a/image-13.jpg)
Stable isotope labeling Control [15 N]Na. NO 3 15 N [13 C]Na. HCO 3 13 C
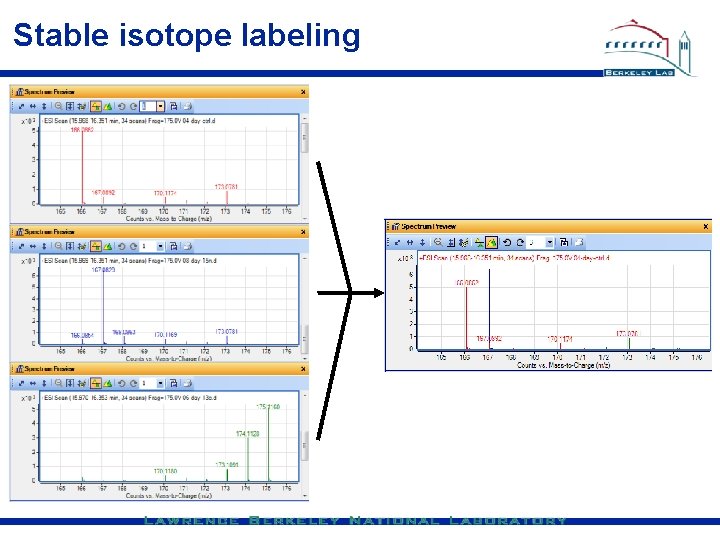
Stable isotope labeling
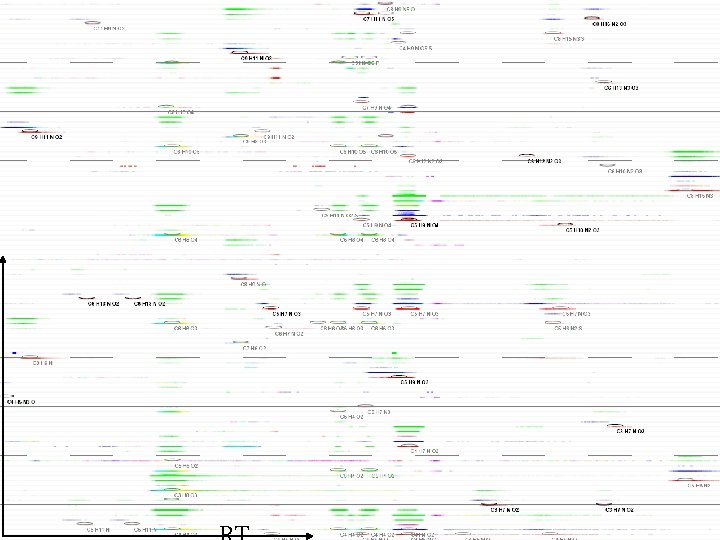
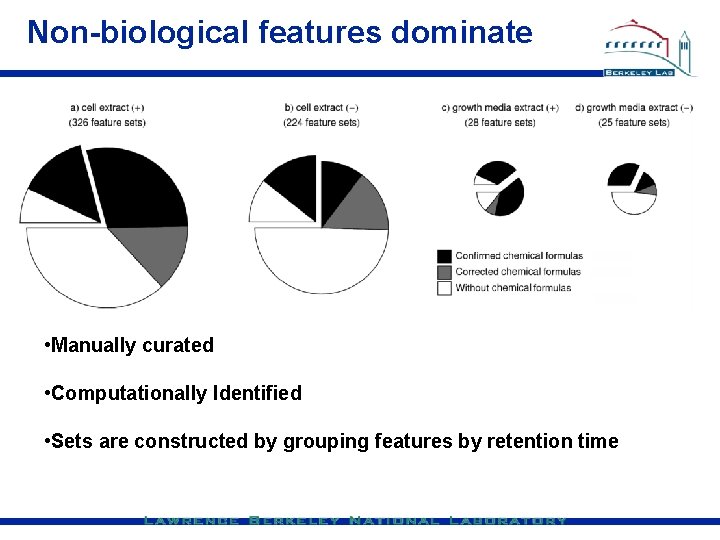
Non-biological features dominate • Manually curated • Computationally Identified • Sets are constructed by grouping features by retention time

Results ~100 distinct metabolites detected 82 assigned chemical formulas 74 unique 45 outside of Syn 7002 Cyc 24 outside of Meta. Cyc or KEGG 54 identified or putatively identified metabolites Using authentic standards or MS/MS

Most dominant biological features Putative hexose(amine)-based trisaccharide:
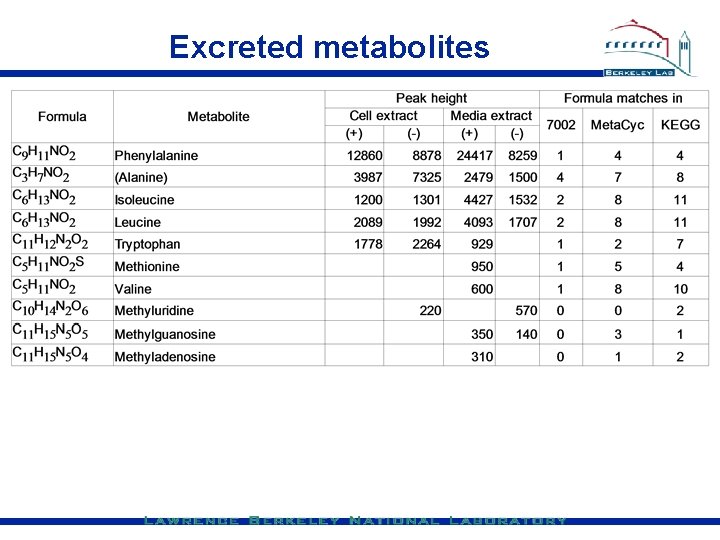
Excreted metabolites
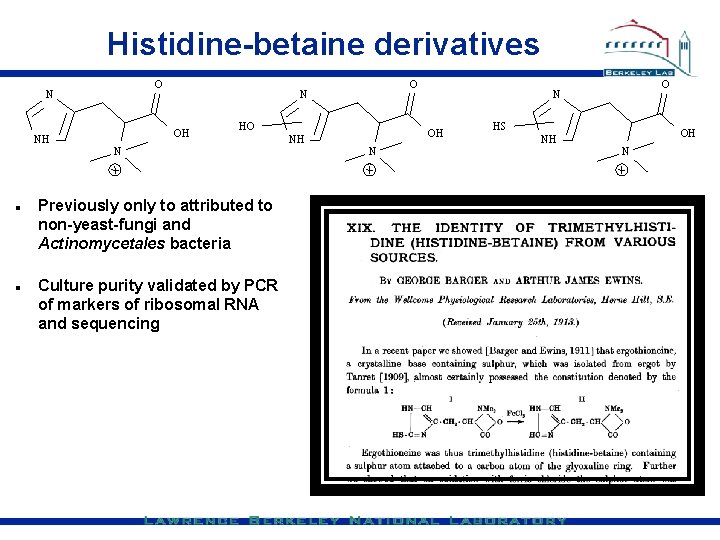
Histidine-betaine derivatives O N NH O N OH HO N Previously only to attributed to non-yeast-fungi and Actinomycetales bacteria Culture purity validated by PCR of markers of ribosomal RNA and sequencing NH OH N O N HS NH OH N
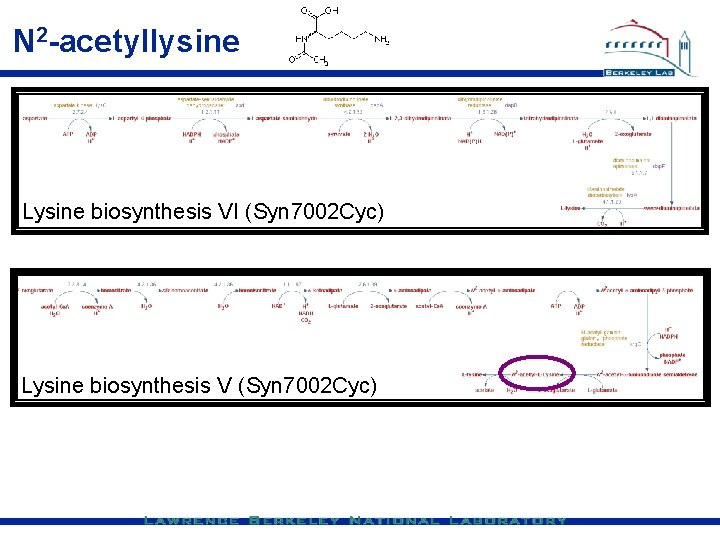
N 2 -acetyllysine Lysine biosynthesis VI (Syn 7002 Cyc) Lysine biosynthesis V (Syn 7002 Cyc)

Analyze selected features by MS/MS Target features at specific m/z & r. t.

MS/MS structural confirmation • Commercial Standards • Metlin • Massbank • Collaborating to expand the number of authentic standards (Siuzdak, Mukhopadhyay) and make these publically available.
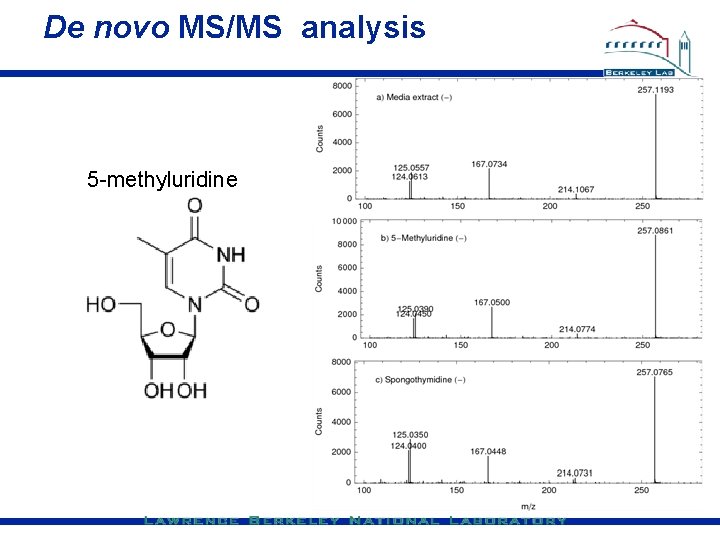
De novo MS/MS analysis 5 -methyluridine
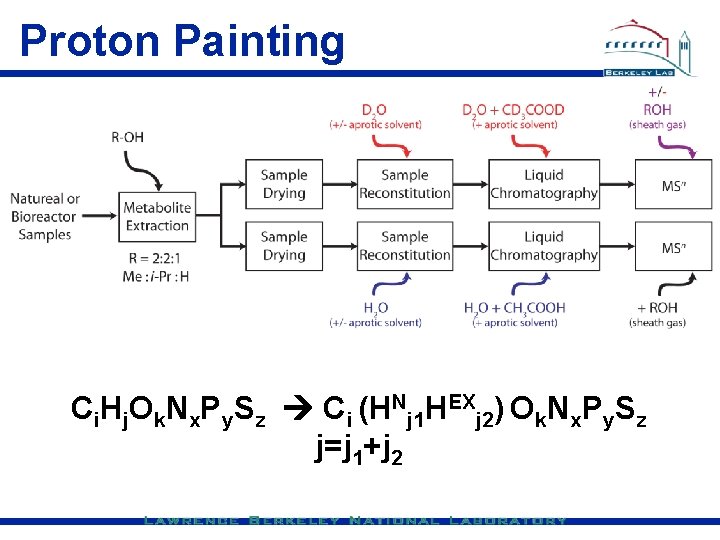
Proton Painting Ci. Hj. Ok. Nx. Py. Sz Ci (HNj 1 HEXj 2) Ok. Nx. Py. Sz j=j 1+j 2
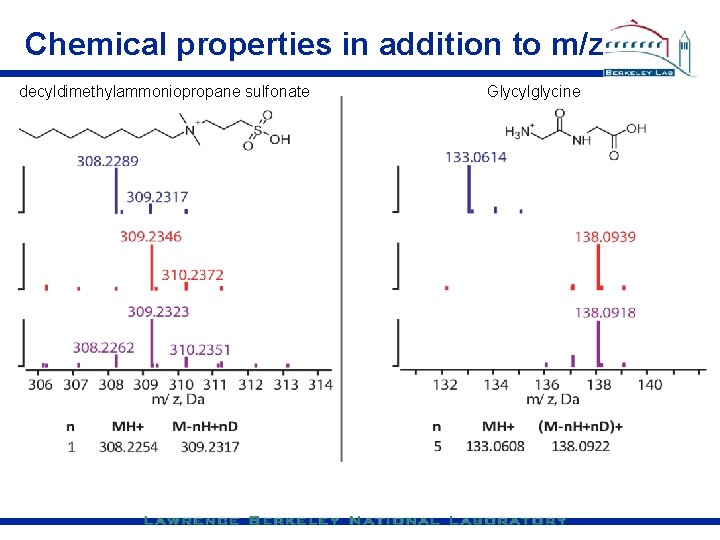
Chemical properties in addition to m/z decyldimethylammoniopropane sulfonate Glycylglycine
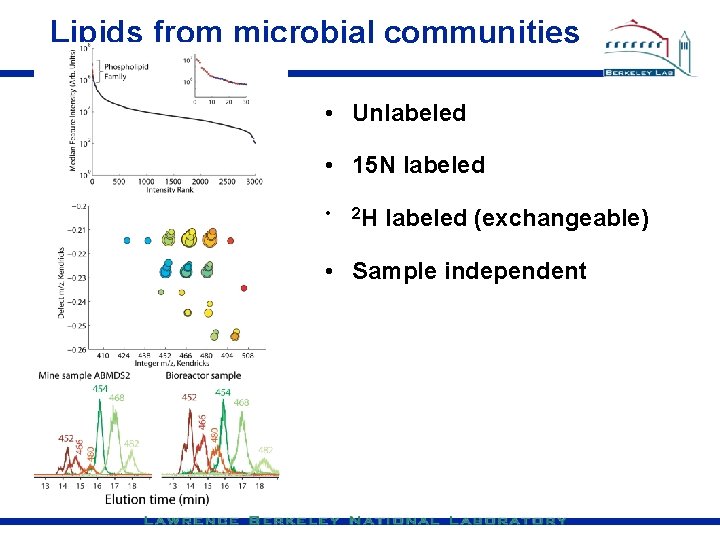
Lipids from microbial communities • Unlabeled • 15 N labeled • 2 H labeled (exchangeable) • Sample independent
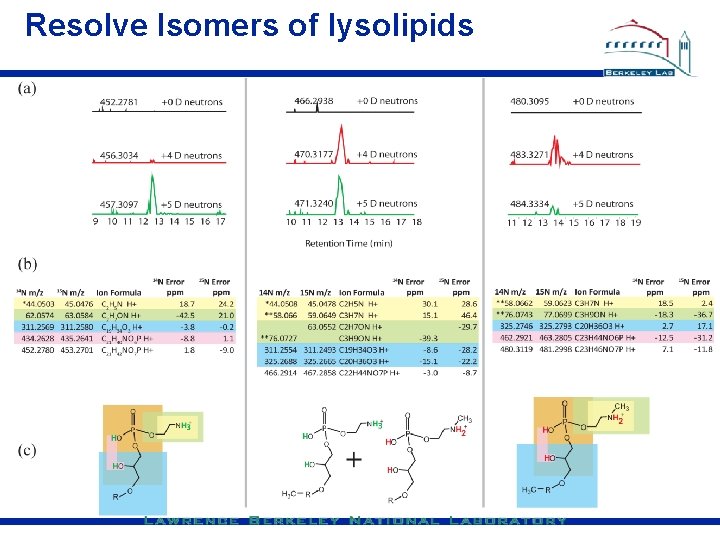
Resolve Isomers of lysolipids
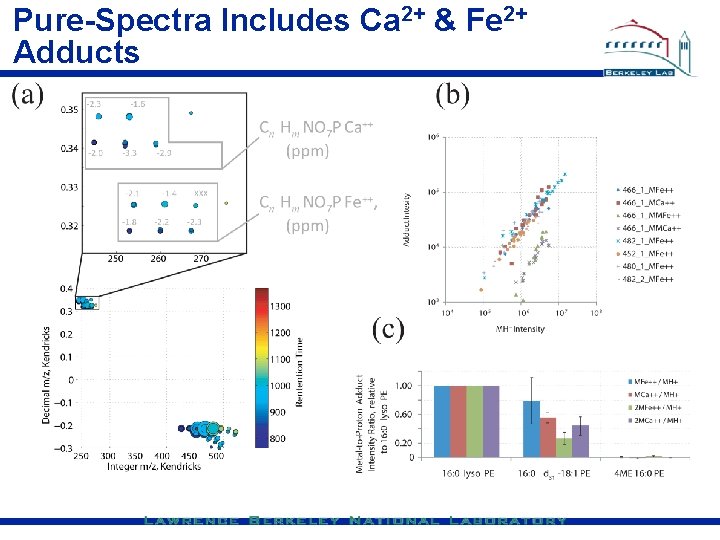
Pure-Spectra Includes Ca 2+ & Fe 2+ Adducts
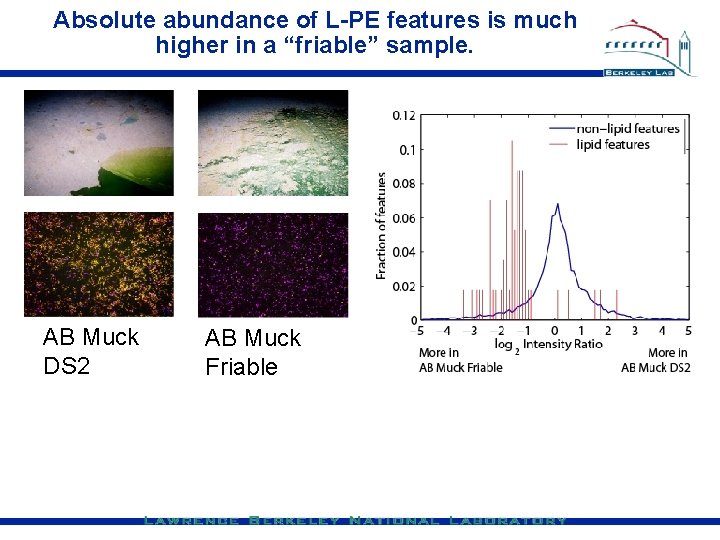
Absolute abundance of L-PE features is much higher in a “friable” sample. AB Muck DS 2 AB Muck Friable
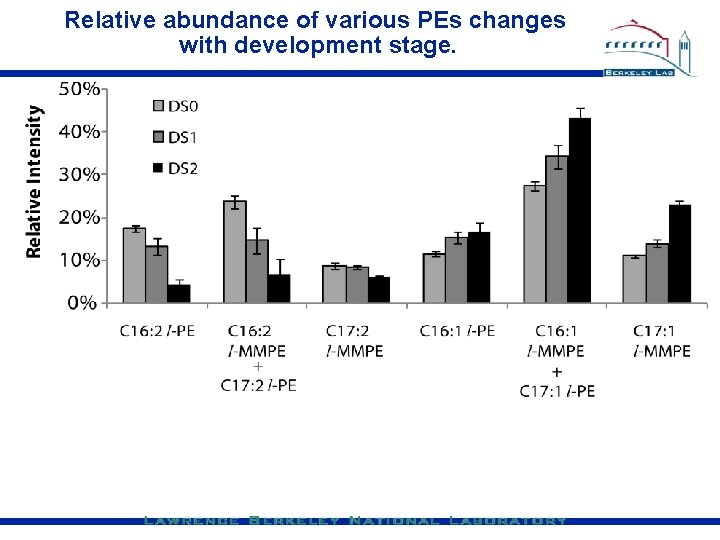
Relative abundance of various PEs changes with development stage.
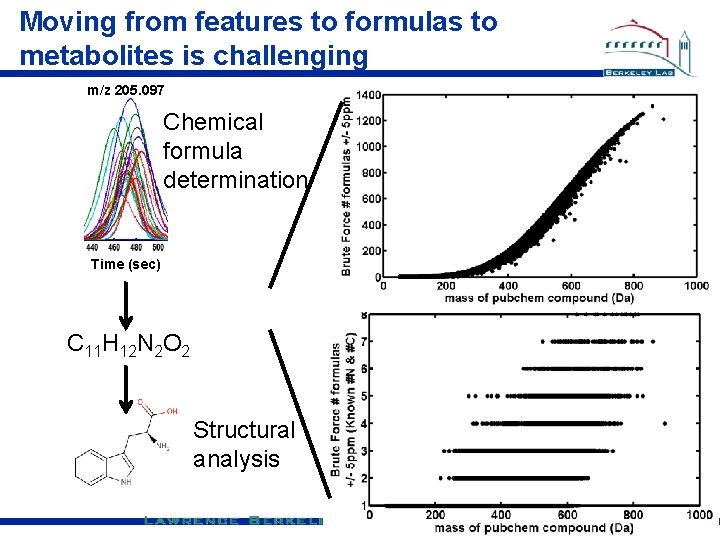
Moving from features to formulas to metabolites is challenging m/z 205. 097 Chemical formula determination Time (sec) C 11 H 12 N 2 O 2 Structural analysis
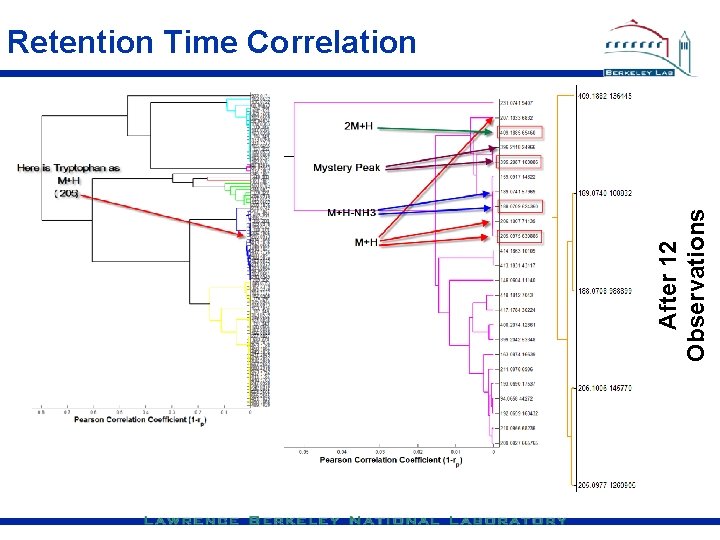
After 12 Observations Retention Time Correlation

Store retention time correlations
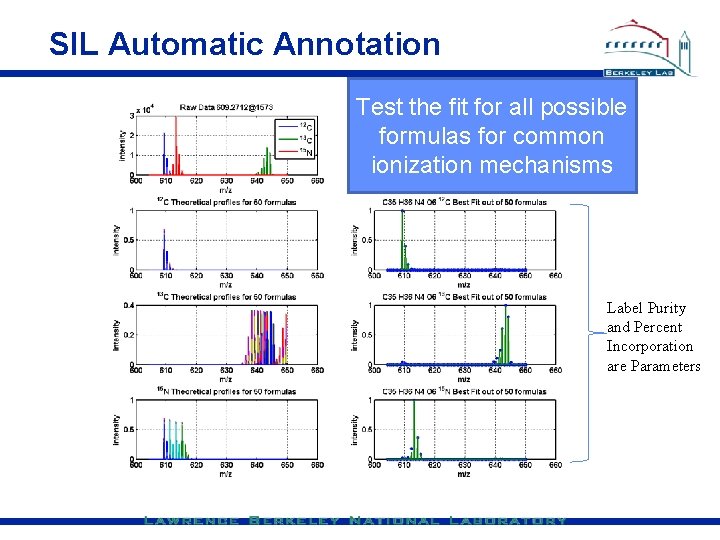
SIL Automatic Annotation Test the fit for all possible formulas for common ionization mechanisms Label Purity and Percent Incorporation are Parameters
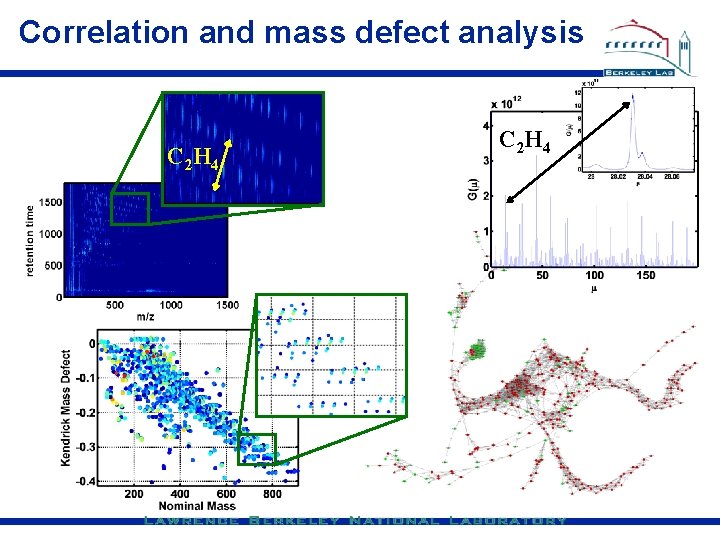
Correlation and mass defect analysis C 2 H 4
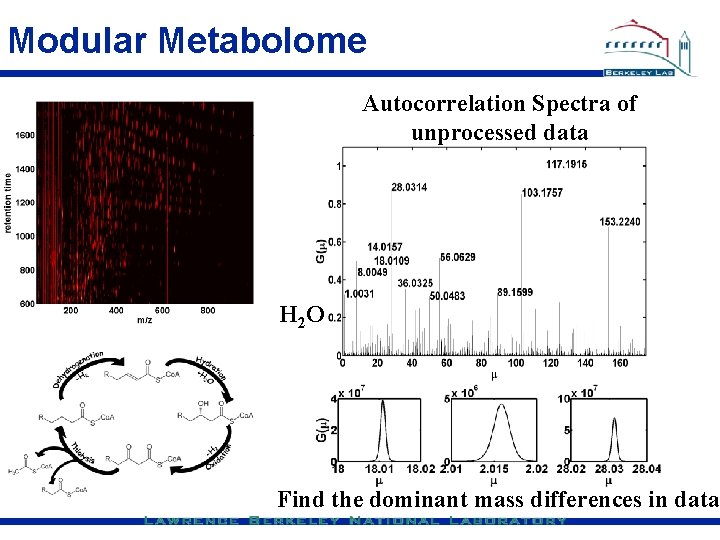
Modular Metabolome Autocorrelation Spectra of unprocessed data H 2 O Find the dominant mass differences in data
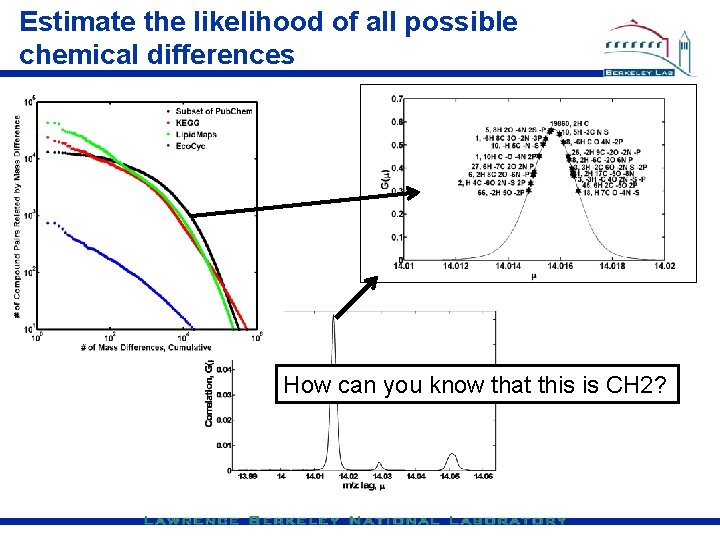
Estimate the likelihood of all possible chemical differences How can you know that this is CH 2?
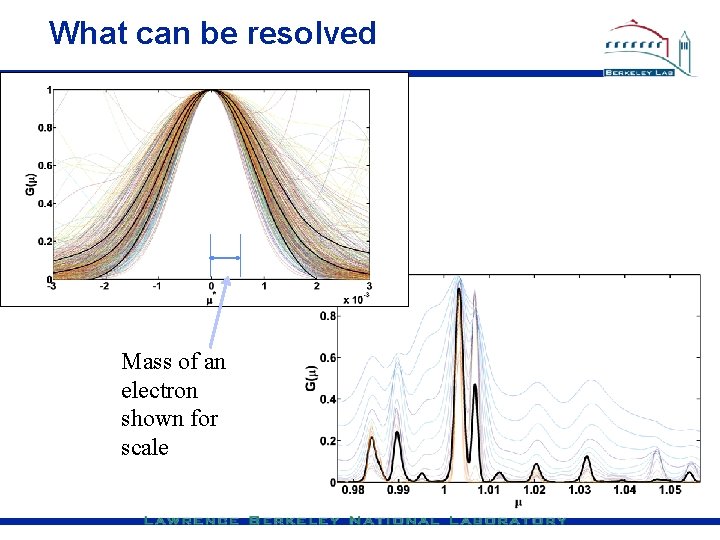
What can be resolved Mass of an electron shown for scale
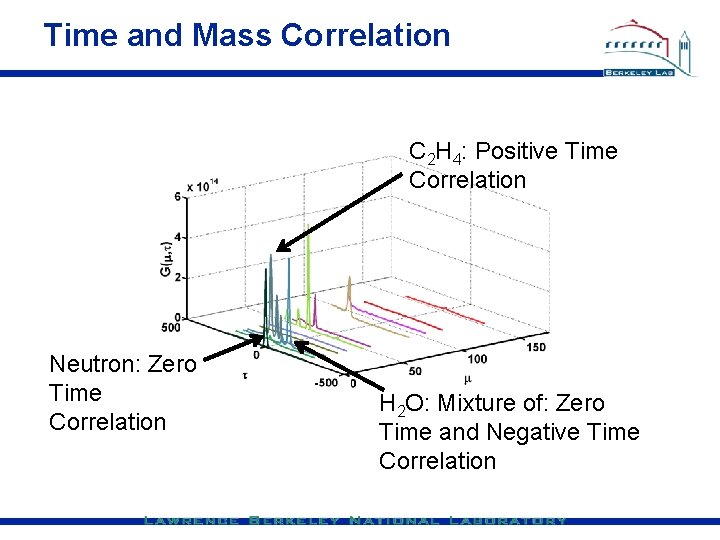
Time and Mass Correlation C 2 H 4: Positive Time Correlation Neutron: Zero Time Correlation H 2 O: Mixture of: Zero Time and Negative Time Correlation
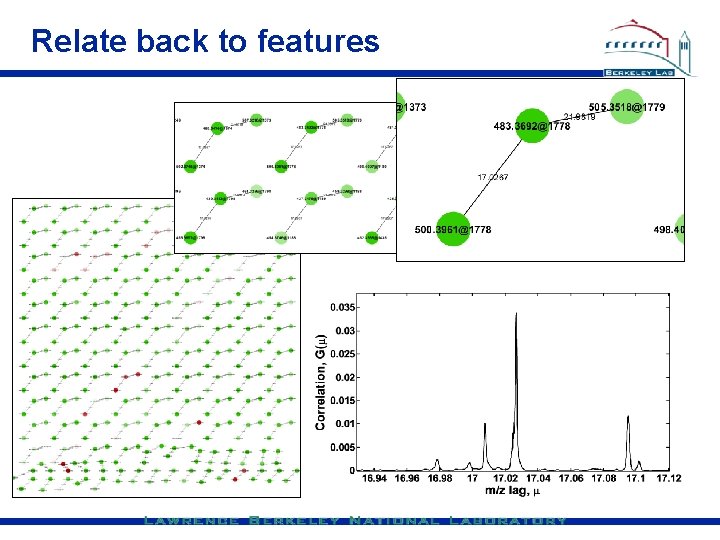
Relate back to features
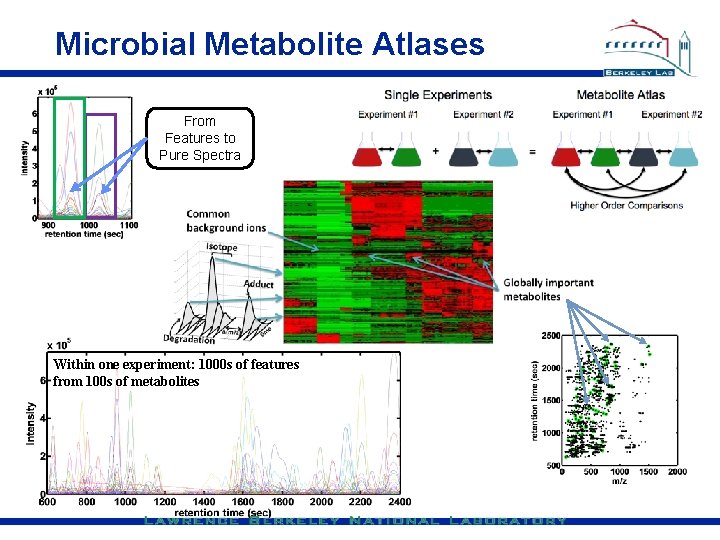
Microbial Metabolite Atlases From Features to Pure Spectra Within one experiment: 1000 s of features from 100 s of metabolites

The End
- Slides: 43