de novo Protein Design Presented by Alison Fraser

de novo Protein Design Presented by Alison Fraser, Christine Lee, Pradhuman Jhala, Corban Rivera

Outline n n Introduction Computational methods used for sequence and structure Biophysical and structural characteristics of novel protein Conclusion

Introduction n n Number of protein folds Computational methods for identifying amino acid sequences compatible with target structure – not for protein creation Side Chains as Templates New protein design => more rigorous test of current force fields and optimization methodology than redesign of naturally occurring proteins

Introduction (continued) n n Search of nearby conformational space and sequence space 2 methods of protein redesign (variation of backbone conformation and amino acid sequence) Development of procedure for identifying low free energy sequence-structure pairs that iterates between sequence optimization and structure prediction Result: 93 residue protein with topology not in PDB

Structure to Sequence n n Rosetta. Design predicts a amino acid sequence from a desired structure. Input and Output Input a structure you would like to create n Output a amino acid sequence that will produce the structure with low free energy n

Sequence to Structure n n Rosetta. Design is used to predict protein structure from protein residue sequence. Input and Output Input Amino Acid sequence n Output a Predicted near minimum free energy structure n

How does Top 7 compare to proteins in nature? n n n Folding Stability Structure
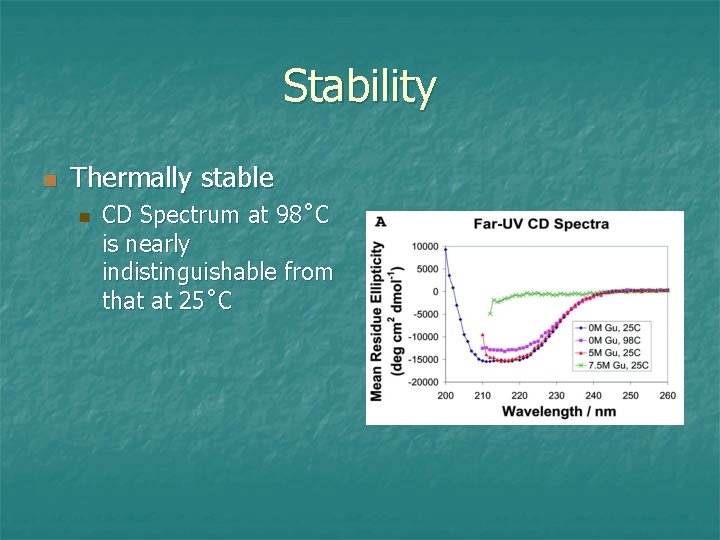
Stability n Thermally stable n CD Spectrum at 98˚C is nearly indistinguishable from that at 25˚C
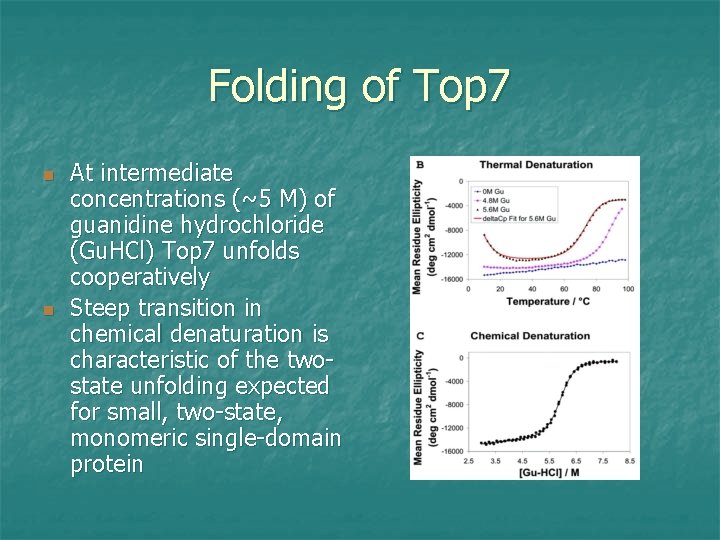
Folding of Top 7 n n At intermediate concentrations (~5 M) of guanidine hydrochloride (Gu. HCl) Top 7 unfolds cooperatively Steep transition in chemical denaturation is characteristic of the twostate unfolding expected for small, two-state, monomeric single-domain protein

Structure n Nuclear Overhauser effect spectroscopy (NOESY) and heteronuclear single quantum coherence (HSQC) exhibit features characteristic of protein with substantial beta-sheet content

Comparing Top 7 to model n Crystallization Top 7 yielded crystals that diffracted to 2. 5 Å n Strong molecular replacement (MR) solution to phase problem n n n This suggest design model very close to true structure Top 7 crystal like the model was also judged to be a novel topology by TOPS server
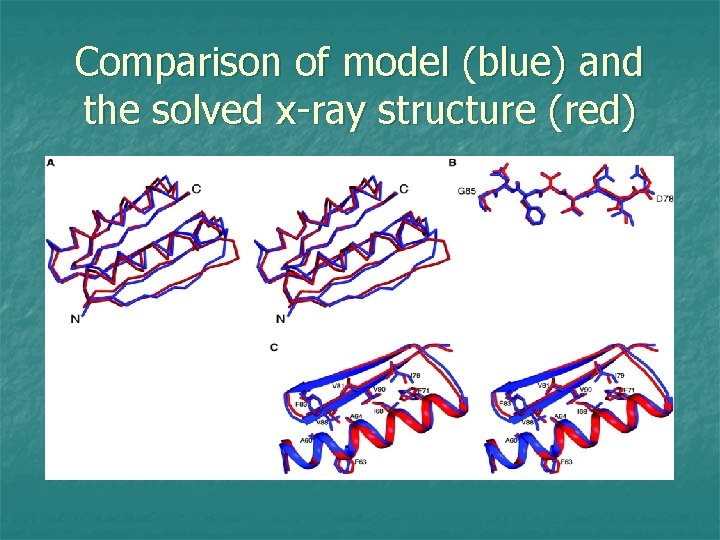
Comparison of model (blue) and the solved x-ray structure (red)

Implications n n Atomic Level Accuracy (RMSD = 1. 17 o. A) in de novo Protein Design Validation of Accuracy of Potential Functions

Reasons n Optimization of Sequence and Structure n No Functional Constraints n Extensive Optimization n No kinetics

Possible Future Impacts n n Synthetic proteins Protein Therapeutics and Molecular Mechanics
- Slides: 15