db SNP the NCBI database of genetic variation

db. SNP: the NCBI database of genetic variation S. T. Sherry, M. H. Ward, M. Kholodov, J. Baker, L. Phan, E. M. Smigielski and K. Sirotkin, Nucleic Acids Research, 29: 1, 2001 National Center for Biotechnology Information, National Library of Medicine, National Institute of Health http: //www. ncbi. nlm. nih. gov/SNP

Composition of SNP database Scope: • Disease-causing mutations • Neutral polymorphisms • Sequence info around polymorphism • Experimental conditions • Population description • Frequency Type of variation % composition Single nucleotide substitutions 99. 7 insertion/deletion 0. 21 Invariant regions of 0. 02 sequence Microsatellite repeats 0. 001
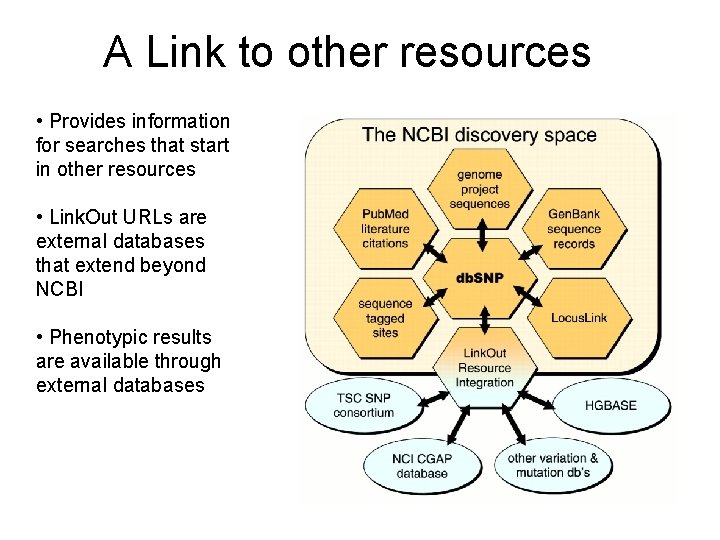
A Link to other resources • Provides information for searches that start in other resources • Link. Out URLs are external databases that extend beyond NCBI • Phenotypic results are available through external databases

Submission Required info: − − − − Contact information of submitter Alleles at locus Flanking sequence surrounding polymorphism Experimental methods Gen. Bank record Population sample and source organism Frequency data * Major contributors to the database are laboratories associated with the National Human Genome Research Institute (NHGRI) grants program

After Submission − ss#: db. SNP accessioning to each submitted variation − rs#: Unique variation in an organism reference genome

Searching the database NCBI resources db. SNP • Gene name • Map location • BLAST • • Accession number Submitter name Local batch ID Method of identification Population type Publication title Chromosome report

Brief summary of Scale • 24 different species have SNP data, including, H. sapiens, A. gambiae, • C. elegans, P. falciparum, Z. mays • 13 million submissions for H. sapiens • 1. 4 million in A. gambiae • 2. 8 millions new submission for H. sapiens * A total of only 5 species and 1. 5 million submissions were present in 2001, at time of publishing
- Slides: 7