Database Publishing at Nature Timo Hannay Nature Publishing

Database Publishing at Nature Timo Hannay Nature Publishing Group 7 October 2005

Overview l Publishing collaborations: Making databases more like journals l NPG New Technology: Making journals more like databases l Tagging and social bookmarking: New methods of annotation and navigation

Database publishing at NPG l The Af. CS-Nature Signaling Gateway (http: //www. signaling-gateway. org/) l The CMC-Nature Cell Migration Gateway (http: //www. cellmigration. org/) l Forthcoming collaborations with NCI and several other groups

The Af. CS-Nature Signaling Gateway l A freely available online resource for anyone interested in cellular signalling l A collaboration with the research community through the Alliance for Cellular Signaling l An experiment in the next generation of online, database-driven scientific publications
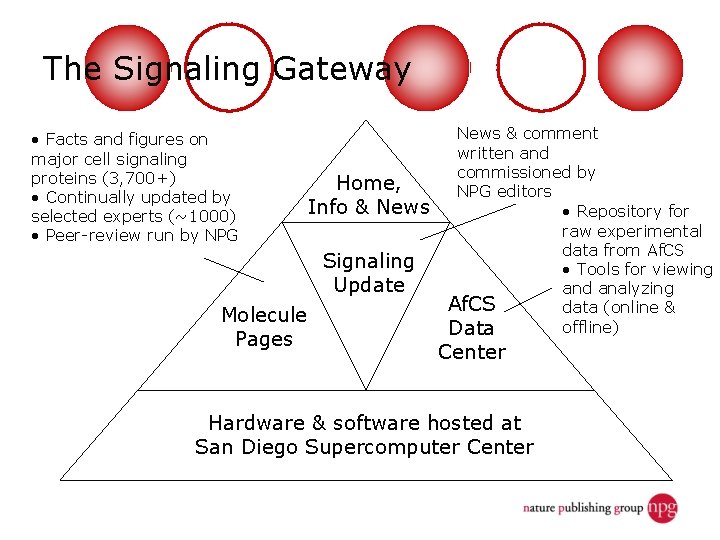
The Signaling Gateway • Facts and figures on major cell signaling proteins (3, 700+) • Continually updated by selected experts (~1000) • Peer-review run by NPG Home, Info & News Signaling Update Molecule Pages News & comment written and commissioned by NPG editors • Repository for raw experimental data from Af. CS • Tools for viewing and analyzing Af. CS data (online & offline) Data Center Hardware & software hosted at San Diego Supercomputer Center

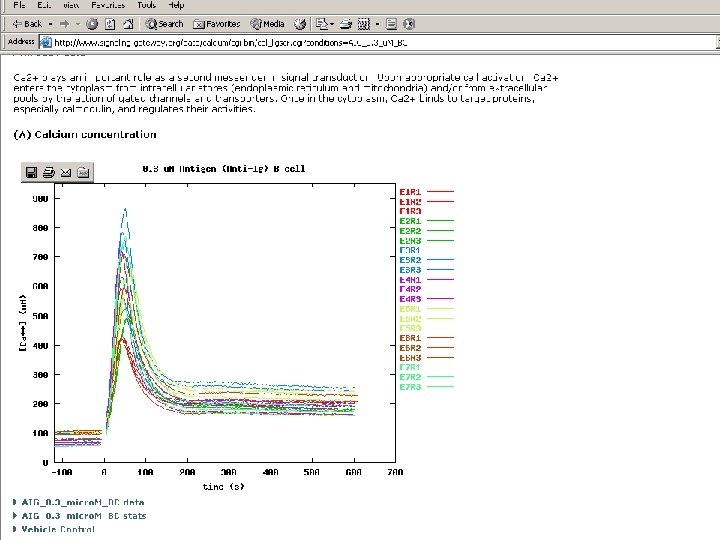
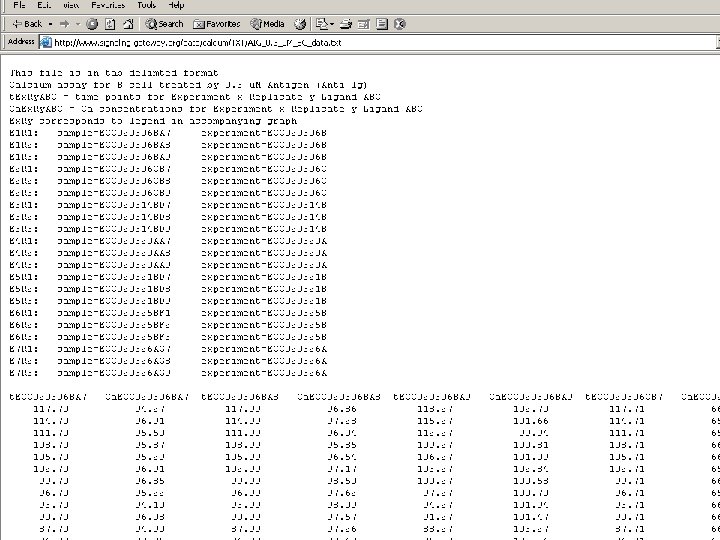

The Molecule Pages l l l Comprehensive, structured data for 3, 700+ proteins involved in cellular signalling Some information automatically fed in from other online databases and updated monthly Other information entered by selected expert authors and updated annually Author-entered data peer-reviewed by NPG Fully citable using digital object identifiers (DOIs)
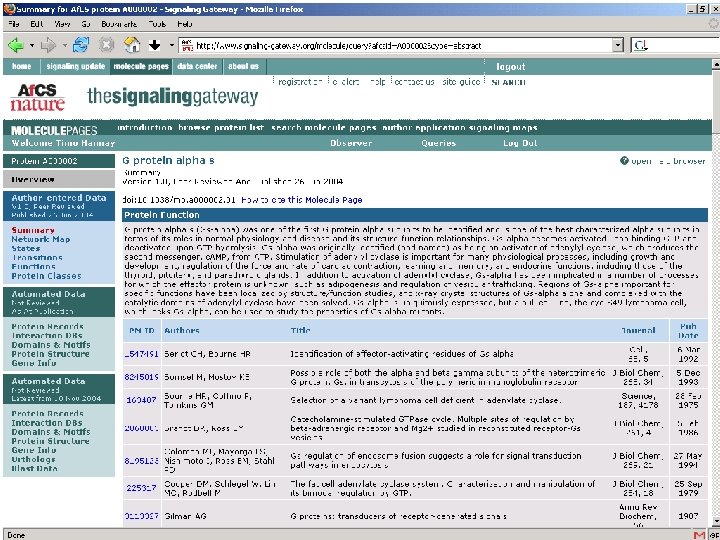
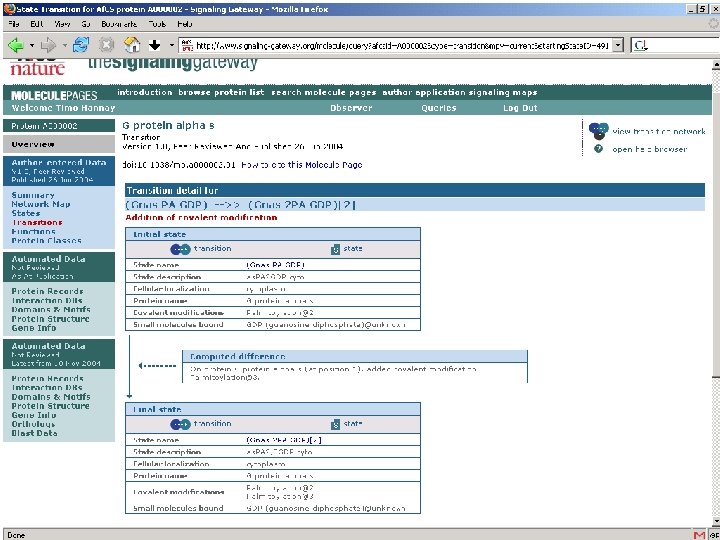

Using Digital Object Identifiers http: //dx. doi. org/10. 1038/35057062 Nature 409, 860 - 921 (2001) doi: 10. 1038/35057062 Correct URL at publisher’s website IDF/Cross. Ref databases • Allows unambiguous identification of paper • Allows readers to find the paper online • Allows publishers to cross-link reference lists • Guaranteed not to change (even if the publisher changes)
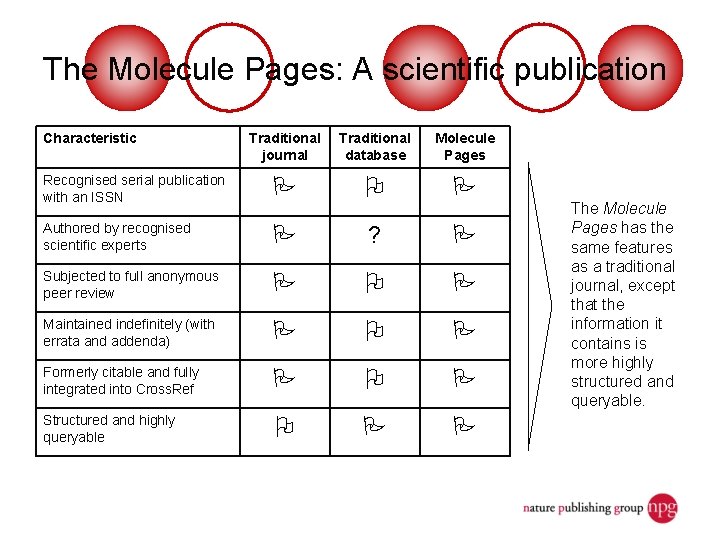
The Molecule Pages: A scientific publication Characteristic Traditional journal Traditional database Molecule Pages Recognised serial publication with an ISSN Authored by recognised scientific experts ? Subjected to full anonymous peer review Maintained indefinitely (with errata and addenda) Formerly citable and fully integrated into Cross. Ref Structured and highly queryable The Molecule Pages has the same features as a traditional journal, except that the information it contains is more highly structured and queryable.

Overview l Publishing collaborations: Making databases more like journals l NPG New Technology: Making journals more like databases l Tagging and social bookmarking: New methods of annotation and navigation
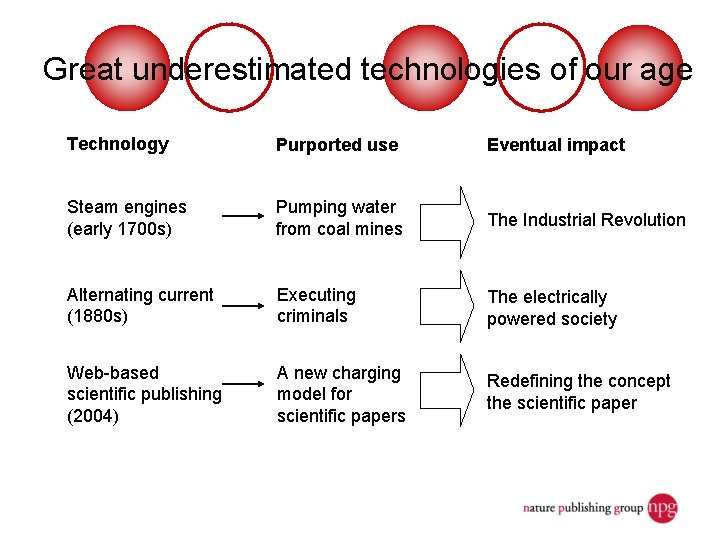
Great underestimated technologies of our age Technology Purported use Eventual impact Steam engines (early 1700 s) Pumping water from coal mines The Industrial Revolution Alternating current (1880 s) Executing criminals The electrically powered society Web-based scientific publishing (2004) A new charging model for scientific papers Redefining the concept the scientific paper

Scientific papers as structured data objects Print journal Article metadatabase Structured data sets <rdf> Online facsimile <svg> </rdf> circa 2000 Structured, interactive and queryable figures and text circa 2006

Experimental article metadatabase Initial data to be included: Author and institute details l Scientific: ¡ Molecules (In. Ch. I) ¡ Genes (Entrez Gene) ¡ Proteins (Uni. Prot) ¡ Cellular processes, functions, locations (GO) ¡ Species (NCBI) l Citation annotations (controlled vocabulary) l
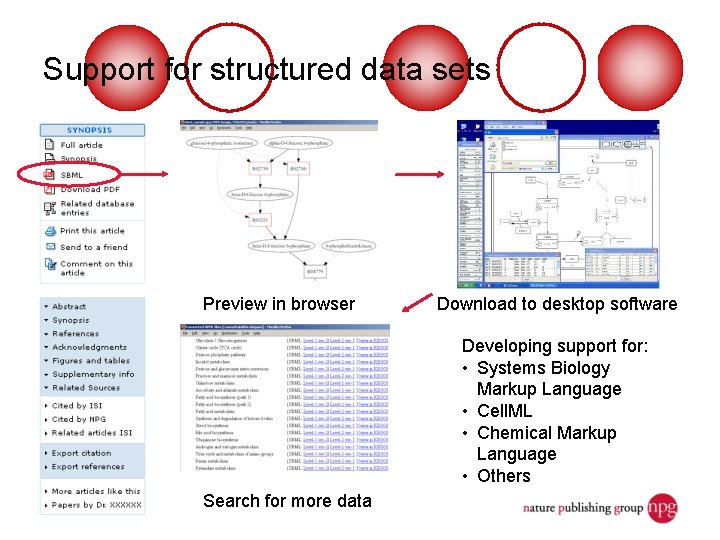
Support for structured data sets Preview in browser Download to desktop software Developing support for: • Systems Biology Markup Language • Cell. ML • Chemical Markup Language • Others Search for more data
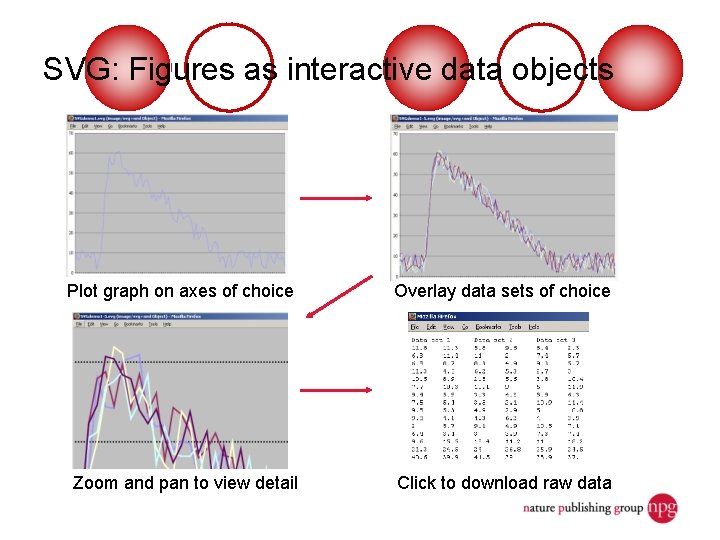
SVG: Figures as interactive data objects Plot graph on axes of choice Overlay data sets of choice Zoom and pan to view detail Click to download raw data
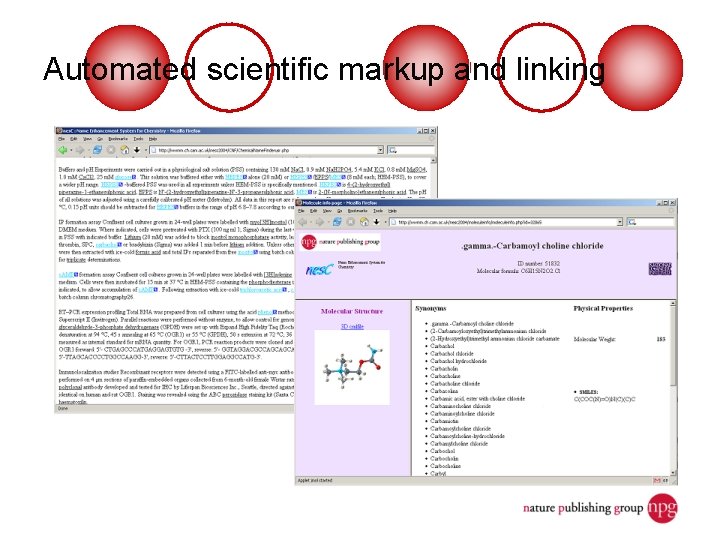
Automated scientific markup and linking

Increasing structure in text markup (1) The old way (no semantic markup): “<p>. . . gp 120 binding to CXCR 4 or CCR 5 activates PYK 2 and FAK…</p>” Now (key entities and concepts marked up): “<p>. . . <protein id="urn: lsid: uniprot. org: uniprot: P 03378">gp 120</protein> <action id="urn: lsid: geneontology. org: go: 000548">binding</action> to <protein id="urn: lsid: uniprot. org: uniprot: P 48061">CXCR 4</protein> or <protein id="urn: lsid: uniprot. org: uniprot: P 10147">CCR 5</protein> <action id="urn: lsid: geneontology. org: go: 0008047">activates</action> <protein id="urn: lsid: uniprot. org: uniprot: O 43150">PYK 2</protein> and <protein id="urn: lsid: uniprot. org: uniprot: Q 05397">FAK</protein>…</p>”

Increasing structure in text markup (2) The new way (full RDF/XML): <p>. . . <rdf: Graph xmlns: rdf="http: //www. w 3. org/1999/02/22 -rdf-syntax-ns#" xmlns: go="urn: lsid: geneontology. org: go: " xmlns: uniprot="urn: lsid: uniprot. org: uniprot: "> <go: 000548> <uniprot: Protein rdf: resource="urn: lsid: uniprot. org: uniprot: P 03378"/> <uniprot: Protein rdf: resource="urn: lsid: uniprot. org: uniprot: P 48061"/> <go: 0008047 rdf: resource="urn: lsid: uniprot. org: uniprot: O 43150"/> <go: 0008047 rdf: resource="urn: lsid: uniprot. org: uniprot: Q 05397"/> </go: 000548> <uniprot: Protein rdf: resource="urn: lsid: uniprot. org: uniprot: P 03378"/> <uniprot: Protein rdf: resource="urn: lsid: uniprot. org: uniprot: P 10147"/> <go: 0008047 rdf: resource="urn: lsid: uniprot. org: uniprot: O 43150"/> <go: 0008047 rdf: resource="urn: lsid: uniprot. org: uniprot: Q 05397"/> </go: 000548> <rdf: label>gp 120 binding to CXCR 4 or CCR 5 activates PYK 2 and FAK</rdf: label> </rdf: Graph> …</p> With RDF markup, the article XML itself literally becomes a relational database

Why go to all this effort? Discoverability and recontextualisation “Show me statements about the hedgehog gene. ” “Find claims that disagree with this. ” Transparency and flexibility “Plot this graph on a different scale, with error bars added and with these two extra data sets overlaid. ” Specificity and completeness “Give me a full description of this mathematical model that I can run on my own computer. ” Reuse and interoperability “Provide the raw data set used in this analysis in a form that allows me to merge it with my own data. ”

Views from the database side “Before the end of the next decade, pathway databases will become scientific journals and journals will become databases. Biologists will be greatly empowered, and bioinformatics will continue its long evolution. ” Lincoln Stein (Reactome) “Is a biological database any different than a biological journal? I am working toward reaching an answer of, no, there is no difference. ” Phil Bourne (Protein Data Bank)

Overview l Publishing collaborations: Making databases more like journals l NPG New Technology: Making journals more like databases l Tagging and social bookmarking: New methods of annotation and navigation
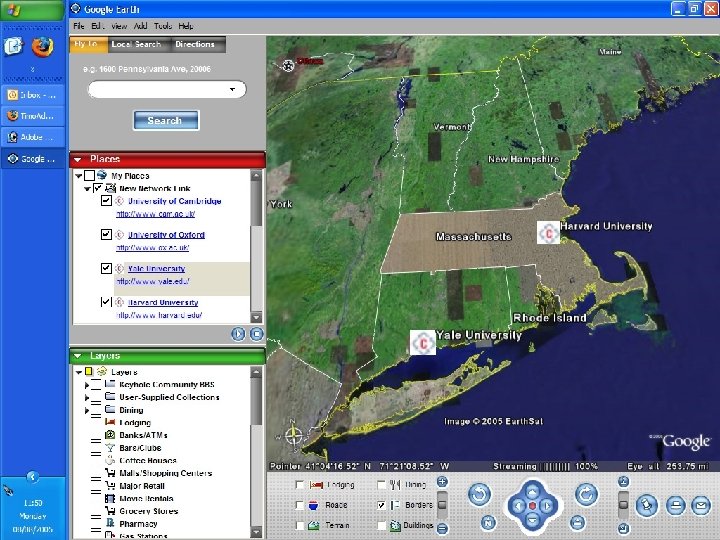

A few uses for Connotea Keeping bookmarks and references in order l Sharing links and ideas within a team (perhaps geographically dispersed) l Providing readers with a (dynamic) list of further or related reading l Encouraging readers to share relevant links with the author and with each other l
- Slides: 49