Database Filtering March 2006 Vineet Bafna ProjectExam deadlines

Database Filtering March 2006 Vineet Bafna

Project/Exam deadlines • May 2 – • May 9 – – • Send email to me with a title of your project Each student/group gives a 10 min. presentation on their proposed project. Show preliminary computations. What is the test plan? What is the data like, and how much is there. Last week of classes: – – – A 20 min. presentation from each group A written report on the project A take home exam, due electronically on the date of the final exam March 2006 Vineet Bafna

Building better filters • • Better filters for nc. RNA is an open and relatively unresearched problems. In contrast, filters for sequence searches have been extensively researched – • Some non-intuitive ideas. We will digress into sequence based filters to see if some of the principles can be exported to other domains. March 2006 Vineet Bafna

Large Database Search • Given a query of length m – – • Identify all sub-sequences in a database that aligns with a high score. Imagine the database to be a single long string of length n The straightforward algorithm would employ a scan of the database. How much time would it take? query Sequnce database March 2006 Vineet Bafna

D. P. computation i j • • The entire computation is one large local alignment. S[i, j]: score of the best local alignment of prefix 1. . i of the database against prefix 1. . j of the query. March 2006 Vineet Bafna
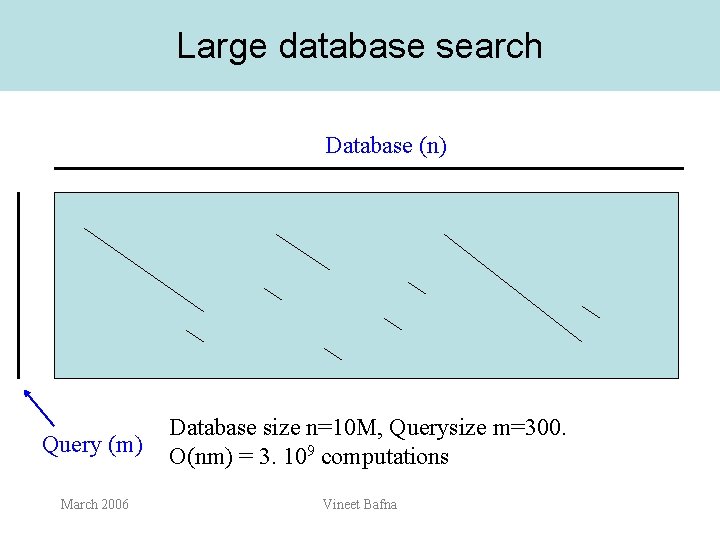
Large database search Database (n) Query (m) March 2006 Database size n=10 M, Querysize m=300. O(nm) = 3. 109 computations Vineet Bafna

Filtering • • The goal of filtering is to reduce the search space to o(nm) using a fast filter How can we filter? March 2006 Vineet Bafna

Observations • • Much of the database is random from the query’s perspective Consider a random DNA string of length n. – • • Pr[A]=Pr[C] = Pr[G]=Pr[T]=0. 25 Assume for the moment that the query is all A’s (length k). What is the probability that an exact match to the query can be found? March 2006 Vineet Bafna
Basic probability • • Probability that there is a match starting at a fixed position i = 0. 25 k What is the probability that some position i has a match. Dependencies confound probability estimates. Related question: What is the expected number of hits? March 2006 Vineet Bafna

Basic Probability: Expectation • Q: Toss a coin: each time it comes up heads, you get a dollar – – What is the money you expect to get after n tosses? Let Xi be the amount earned in the i-th toss § Total money you expect to earn March 2006 Vineet Bafna

Expected number of matches i § Let Xi=1 if there is a match starting at position i, Xi=0 otherwise § Expected number of matches = March 2006 Vineet Bafna

Expected number of exact Matches is small! • Expected number of matches = n*0. 25 k – – – • If n=107, k=10, • Then, expected number of matches = 9. 537 If n=107, k=11 • expected number of hits = 2. 38 n=107, k=12, • Expected number of hits = 0. 5 < 1 Bottom Line: An exact match to a substring of the query is unlikely just by chance. March 2006 Vineet Bafna

Blast filter Take all m-k words of length k. Filter: Consider only those sequences that match at least one of these words. Expected number of matches in a random database? • • • =(m-k)(n-k) (1/4)k • • • Efficiency = (1/4)k A small increase in k decreases efficiency considerably What can we say about accuracy? March 2006 Vineet Bafna

Observation 2: Pigeonhole principle § Suppose we are looking for a database string with greater than 90% identity to the query (length 100) § Partition the query into size 10 substrings. At least one must match the database string exactly March 2006 Vineet Bafna

Why is this important? • • Suppose we are looking for sequences that are 80% identical to the query sequence of length 100. Assume that the mismatches are randomly distributed. What is the probability that there is no stretch of 10 bp, where the query and the subject match exactly? Rough calculations show that it is very low. Exact match of a short query substring to a truly similar subject is very high. – – The above equation does not take dependencies into account Reality is better because the matches are not randomly distributed March 2006 Vineet Bafna

Combining the Facts • Consider the set of all substrings of the query string of fixed length W. – – Prob. of exact match to a random database string is very low. Prob. of exact match to a true homolog is very high. This filter is efficient and accurate. What about speed? Keyword Search (exact matches) is MUCH faster than sequence alignment March 2006 Vineet Bafna
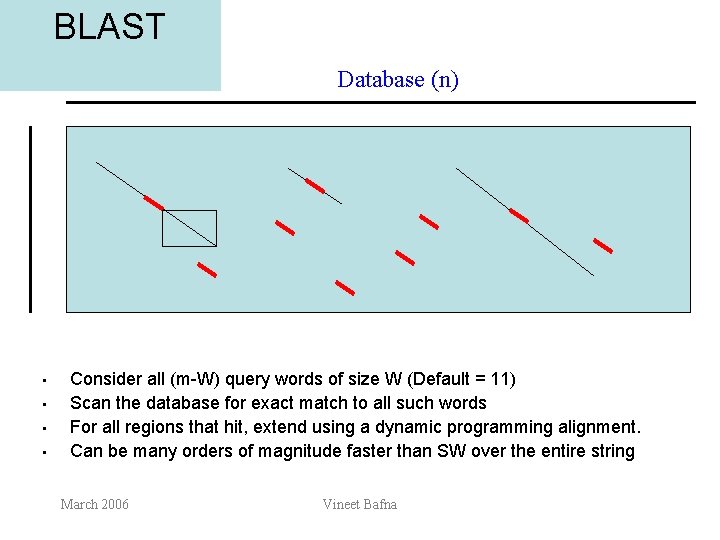
BLAST Database (n) • • Consider all (m-W) query words of size W (Default = 11) Scan the database for exact match to all such words For all regions that hit, extend using a dynamic programming alignment. Can be many orders of magnitude faster than SW over the entire string March 2006 Vineet Bafna

Why is BLAST fast? • • Assume that keyword searching does not consume any time and that alignment computation the expensive step. Query m=1000, random Db n=107, no TP SW = O(nm) = 1000*107 = 1010 computations 50 BLAST, W=11 50 • • E(#11 -mer hits)= 1000* (1/4)11 * 107=2384 Number of computations = 2384*100*50=1. 292*107 Ratio=1010/(1. 292*107)=774 Further speed improvements are possible March 2006 Vineet Bafna
Filter Speed: Keyword Matching • • How fast can we match keywords? Hash table/Db index? What is the size of the hash table, for m=11 Suffix trees? What is the size of the suffix trees? Trie based search. We will do this in class. March 2006 Vineet Bafna AATCA 567

Dictionary Matching 1: POTATO 2: POTASSIUM 3: TASTE P O T A S T P O T A T O database dictionary • • Q: Given k words (si has length li), and a database of size n, find all matches to these words in the database string. How fast can this be done? March 2006 Vineet Bafna

Dict. Matching & string matching • How fast can you do it, if you only had one word of length m? – – • Dictionary matching – – – • Trivial algorithm O(nm) time Pre-processing O(m), Search O(n) time. Trivial algorithm (l 1+l 2+l 3…)n Using a keyword tree, lpn (lp is the length of the longest pattern) Aho-Corasick: O(n) after preprocessing O(l 1+l 2. . ) We will consider the most general case March 2006 Vineet Bafna

Direct Algorithm P O P O T A S T P O T A T O P T O A TA A O P O T A T O P A O T TO O Observations: • When we mismatch, we (should) know something about where the next match will be. • When there is a mismatch, we (should) know something about other patterns in the dictionary as well. March 2006 Vineet Bafna
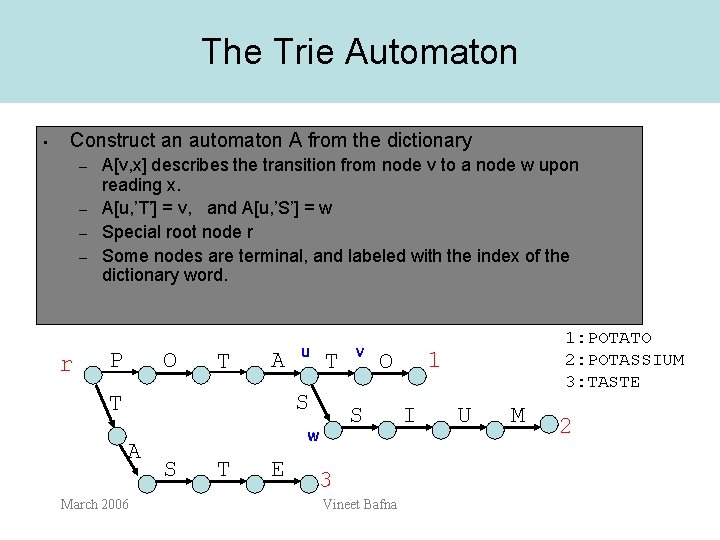
The Trie Automaton • Construct an automaton A from the dictionary – – r A[v, x] describes the transition from node v to a node w upon reading x. A[u, ’T’] = v, and A[u, ’S’] = w Special root node r Some nodes are terminal, and labeled with the index of the dictionary word. P O T A u T S T A March 2006 T E 1 O S w S v 3 Vineet Bafna 1: POTATO 2: POTASSIUM 3: TASTE I U M 2

An O(lpn) algorithm for keyword matching • • Start with the first position in the db, and the root node. If successful transition – – – • Else – – – March 2006 Vineet Bafna Increment current pointer Move to a new node If terminal node “success” Retract ‘current’ pointer Increment ‘start’ pointer Move to root & repeat
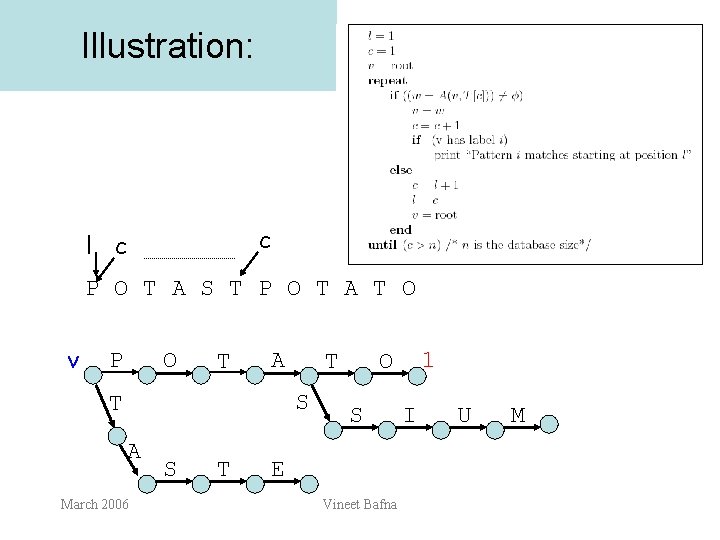
Illustration: c l c P O T A S T P O T A T O v P O T A T S T A March 2006 S T 1 O S E Vineet Bafna I U M
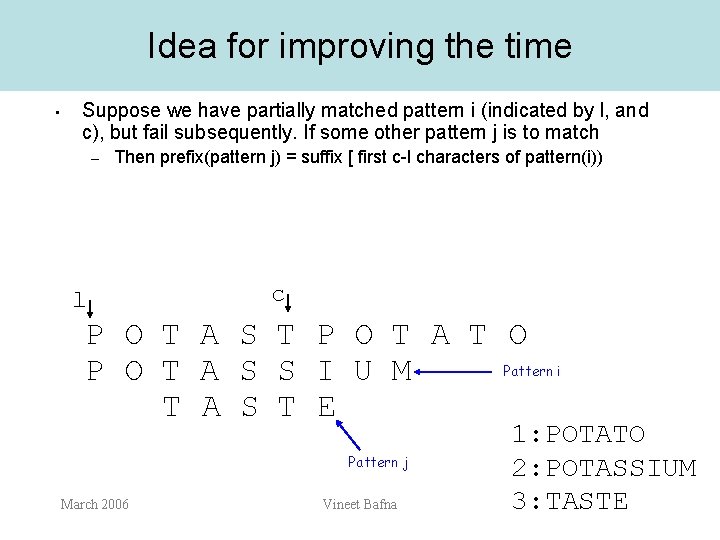
Idea for improving the time • Suppose we have partially matched pattern i (indicated by l, and c), but fail subsequently. If some other pattern j is to match – Then prefix(pattern j) = suffix [ first c-l characters of pattern(i)) l c P O T A S T P O T A T O Pattern i P O T A S S I U M T A S T E Pattern j March 2006 Vineet Bafna 1: POTATO 2: POTASSIUM 3: TASTE

Improving speed of dictionary matching • • Every node v corresponds to a string sv that is a prefix of some pattern. Define F[v] to be the node u such that su is the longest suffix of sv If we fail to match at v, we should jump to F[v], and commence matching from there Let lp[v] = |su| P 2 O 3 T 4 A 5 T 1 7 S 6 T A March 20068 S 9 T 10 E O S 11 I Vineet Bafna U M
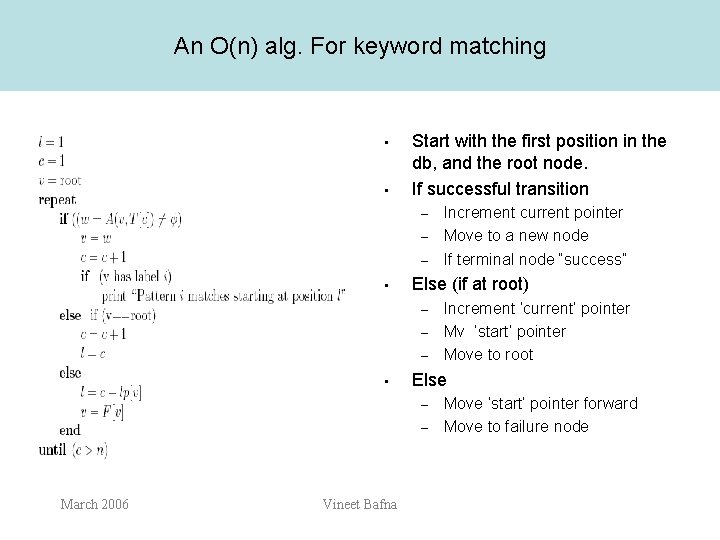
An O(n) alg. For keyword matching • • Start with the first position in the db, and the root node. If successful transition – – – • Else (if at root) – – – • – Vineet Bafna Increment ‘current’ pointer Mv ‘start’ pointer Move to root Else – March 2006 Increment current pointer Move to a new node If terminal node “success” Move ‘start’ pointer forward Move to failure node
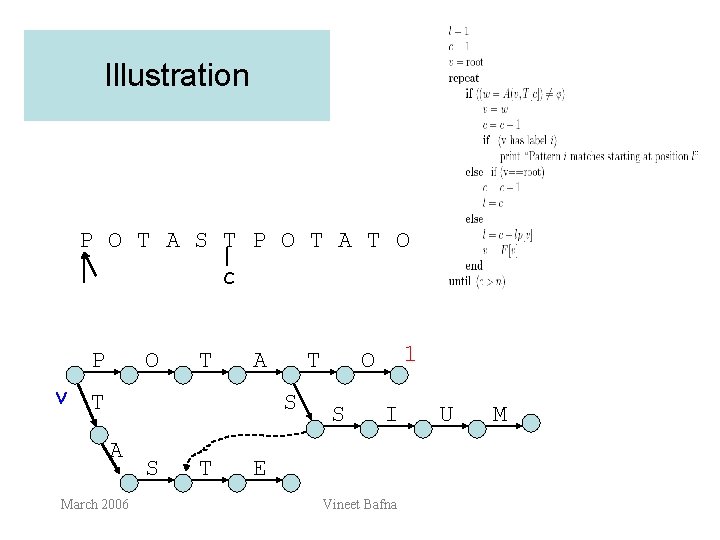
Illustration P O T A S T P O T A T O l c P v O T A T T S A March 2006 S T 1 O S I E Vineet Bafna U M

Time analysis • • In each step, either c is incremented, or l is incremented Neither pointer is ever decremented (lp[v] < c-l). l and c do not exceed n Total time <= 2 n l c P O T A S T P O T A T O March 2006 Vineet Bafna

Blast: Putting it all together • • Input: Query of length m, database of size n Select word-size, scoring matrix, gap penalties, E-value cutoff March 2006 Vineet Bafna

Blast Steps 1. 2. 3. 4. 5. Generate an automaton of all query keywords. Scan database using a “Dictionary Matching” algorithm (O(n) time). Identify all hits. Extend each hit using a variant of “local alignment” algorithm. Use the scoring matrix and gap penalties. For each alignment with score S, compute the bitscore, E-value, and the P-value. Sort according to increasing E-value until the cut-off is reached. Output results. March 2006 Vineet Bafna

Can we improve the filter? • For a query word of size M, – – Consider a binary string Q of length M with W<=M ones. Q ‘matches’ a substring as long as the ‘ones’ match 11010011010 ACCGTCACGTA ACCATAAACAGAUACTTAATTTGGGA March 2006 Vineet Bafna M=11 W=6 W = weight of spaced seed

Can Spaced seeds help? • • The ‘spaced seed’ for BLAST has W consecutive 1 s. Efficiency? – – • Blast Expected(hits) = n p. W For any (M, W), expected hits =~ np. W Accuracy? March 2006 Vineet Bafna

Accuracy • • • Consider a 64 bp sequence that is 70% similar to the query. Pr(an 11 mer matches) = 0. 3 Pr(A spaced seed 11101001. . Matches) = 0. 466 This non-intuitive result leads to selection of spaced words that are an order of magnitude faster for identical specificity and sensitivity Implemented in PATTERNHUNTER March 2006 Vineet Bafna

How to compute a spaced seed • • No good algorithm is known. Iterate over all (M choose W) seeds. – – Use a computation to decide Pr(match) Choose the seed that maximizes probability. March 2006 Vineet Bafna

Prob. Computation for Spaced Seeds • • • Given a specific seed Q(M, W), compute the probability of a hit in a sequence of length L. We can assume that there is a probability p of match. The match mismatch string is a binary string with probability p of 1 1 L 11101110111100 March 2006 Vineet Bafna

Prob. Computation for Spaced Seeds • Given a specific seed Q(M, W), compute the probability of a hit in a sequence of length L. – • Q is a binary string of length M, with W 1 s We try to match the binary ‘match string’ S which is a random binary string with probability p of success. M 1 L 110… 0. 1… 1. . 0 • • PQ = Prob. (Q matches random S at some location) How can we compute PQ? March 2006 Vineet Bafna

Computing F(i, b) • • For a specific string b, define F(i, b) = Prob. (Q matches a random string S of length i, s. t. S ends in B) i 1 b March 2006 Vineet Bafna

Why is it sufficient to compute f(i, b) • PQ = f(L, ) b • • • We have two possibilities: b B 1 : b is consistent with a suffix of Q. b B 0 = B-B 1 110001 Q March 2006 Vineet Bafna 110001
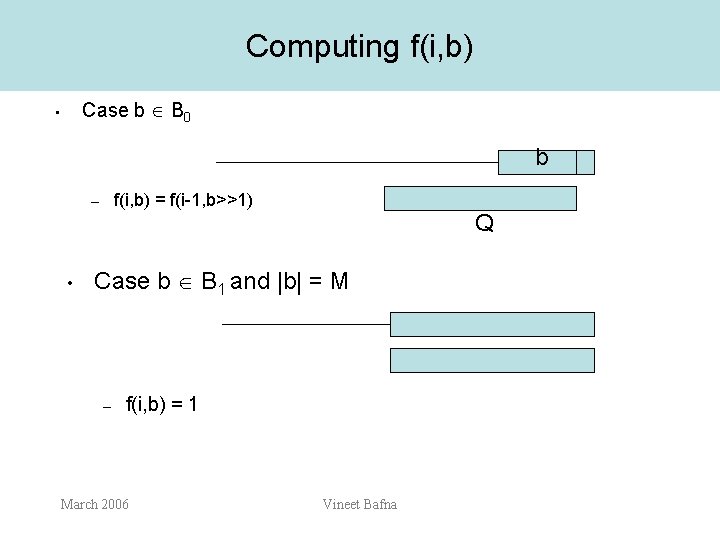
Computing f(i, b) Case b B 0 • b f(i, b) = f(i-1, b>>1) – • Q Case b B 1 and |b| = M – f(i, b) = 1 March 2006 Vineet Bafna

Computing f(i, b) • Case b B 1 b – f(i, b) = pf(i-1, 1 b) + (1 -p) pf(i-1, 0 b) March 2006 Vineet Bafna Q
- Slides: 42