Data analysis methods for nextgeneration sequencing technologies Gabor

Data analysis methods for nextgeneration sequencing technologies Gabor T. Marth Boston College Biology Department Epigenomics & Sequencing Meeting July 14 -15, 2008, Boston, MA

T 1. Roche / 454 FLX system • pyrosequencing technology • variable read-length • the only new technology with >100 bp reads • tested in many published applications • supports paired-end read protocols with up to 10 kb separation size
T 2. Illumina / Solexa Genome Analyzer • fixed-length short-read sequencer • read properties are very close traditional capillary sequences • very low INDEL error rate • tested in many published applications • paired-end read protocols support short (<600 bp) separation
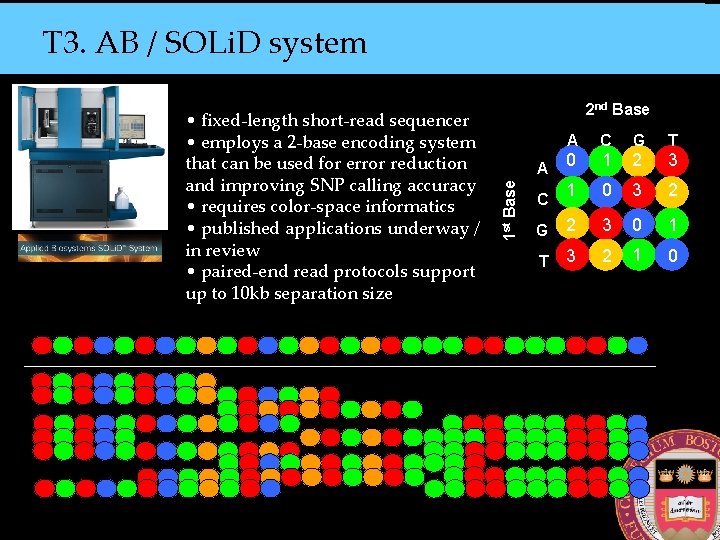
T 3. AB / SOLi. D system A C G T 0 1 2 3 1 0 3 2 G 2 3 0 1 T 3 2 1 0 A 1 st Base • fixed-length short-read sequencer • employs a 2 -base encoding system that can be used for error reduction and improving SNP calling accuracy • requires color-space informatics • published applications underway / in review • paired-end read protocols support up to 10 kb separation size 2 nd Base C
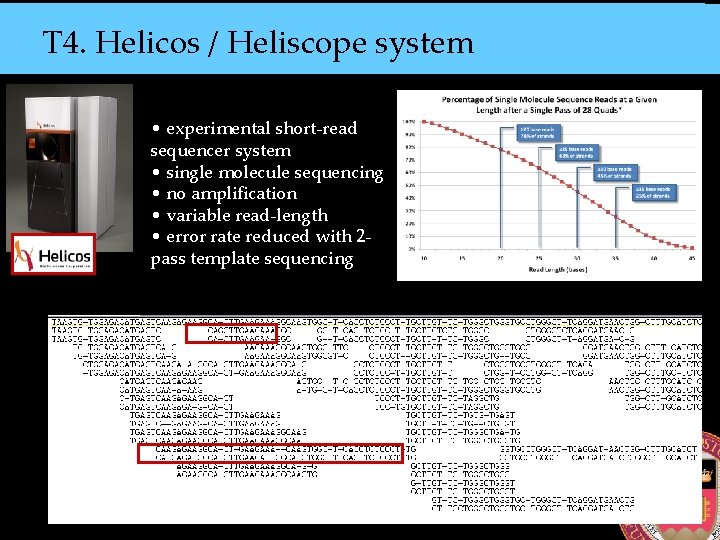
T 4. Helicos / Heliscope system • experimental short-read sequencer system • single molecule sequencing • no amplification • variable read-length • error rate reduced with 2 pass template sequencing

A 1. Variation discovery: SNPs and short-INDELs 1. sequence alignment 2. dealing with non-unique mapping 3. looking for allelic differences
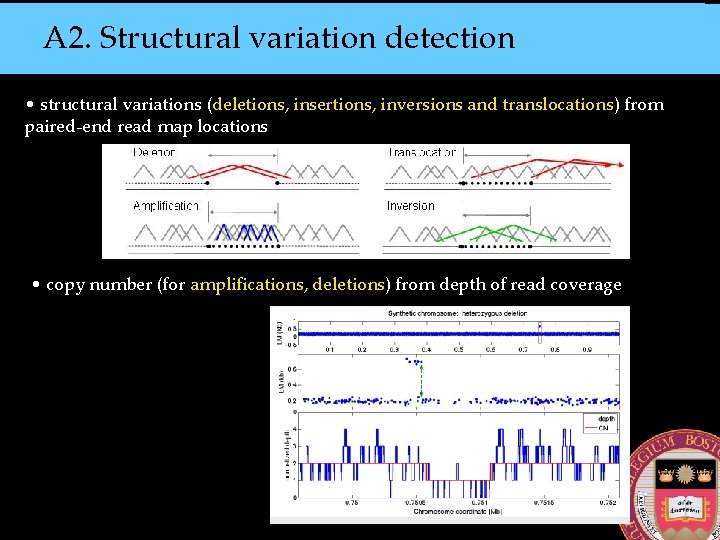
A 2. Structural variation detection • structural variations (deletions, insertions, inversions and translocations) from paired-end read map locations • copy number (for amplifications, deletions) from depth of read coverage

A 3. Identification of protein-bound DNA genome sequence aligned reads Chromatin structure (CHIP-SEQ) (Mikkelsen et al. Nature 2007) Transcription binding sites. Robertson et al. Nature Methods, 2007
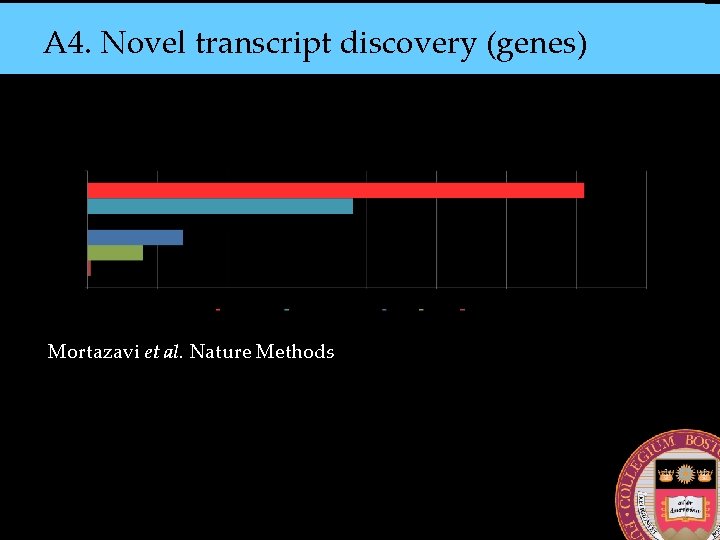
A 4. Novel transcript discovery (genes) Mortazavi et al. Nature Methods

A 5. Novel transcript discovery (mi. RNAs) Ruby et al. Cell, 2006

A 6. Expression profiling by tag counting gene aligned reads Jones-Rhoads et al. PLo. S Genetics, 2007
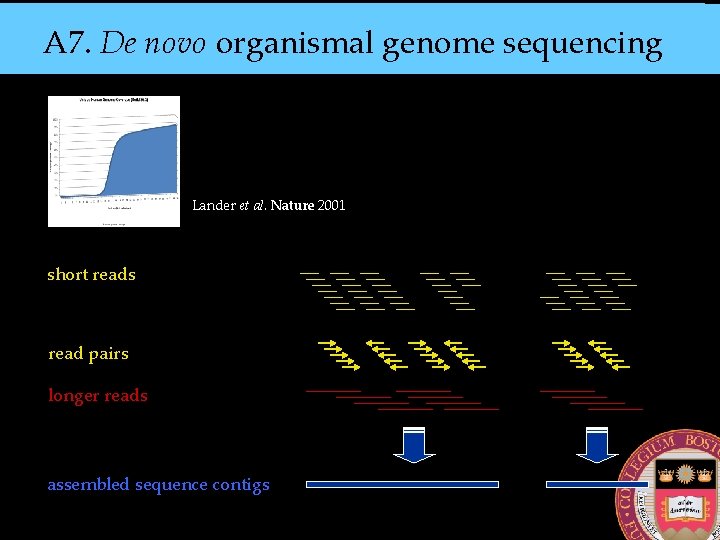
A 7. De novo organismal genome sequencing Lander et al. Nature 2001 short reads read pairs longer reads assembled sequence contigs
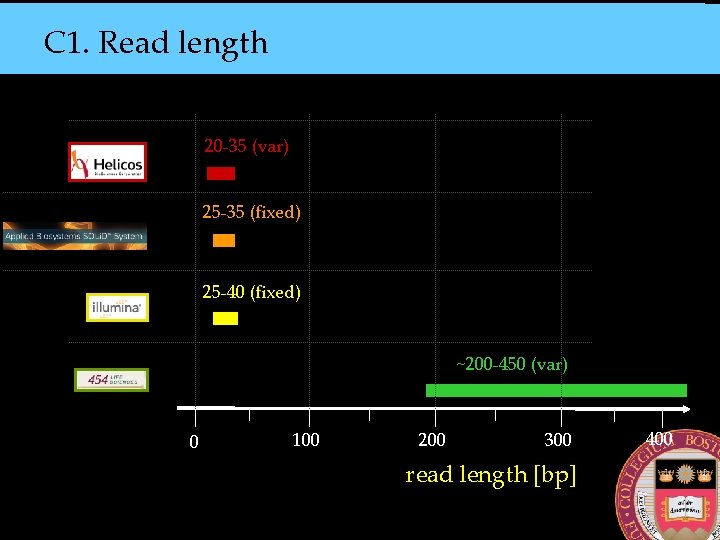
C 1. Read length 20 -35 (var) 25 -35 (fixed) 25 -40 (fixed) ~200 -450 (var) 0 100 200 300 read length [bp] 400

When does read length matter? • short reads often sufficient where the entire read length can be used for mapping: SNPs, short-INDELs, SVs CHIP-SEQ short RNA discovery counting (m. RNA mi. RNA) • longer reads are needed where one must use parts of reads for mapping: de novo sequencing novel transcript discovery aacttagacttacatacgta Known exon 1 Known exon 2 accgattacta
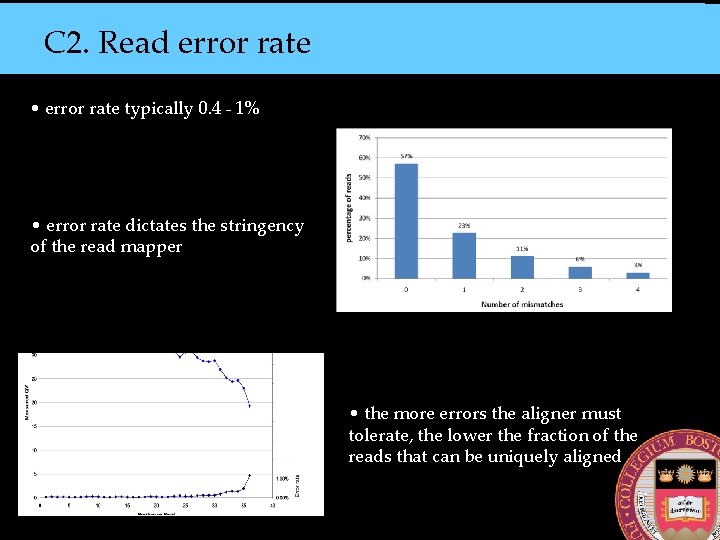
C 2. Read error rate • error rate typically 0. 4 - 1% • error rate dictates the stringency of the read mapper • the more errors the aligner must tolerate, the lower the fraction of the reads that can be uniquely aligned
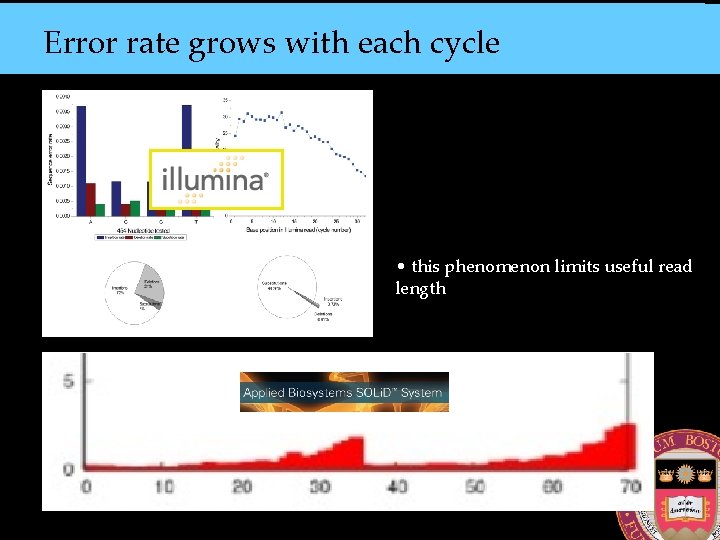
Error rate grows with each cycle • this phenomenon limits useful read length

Substitutions vs. INDEL errors
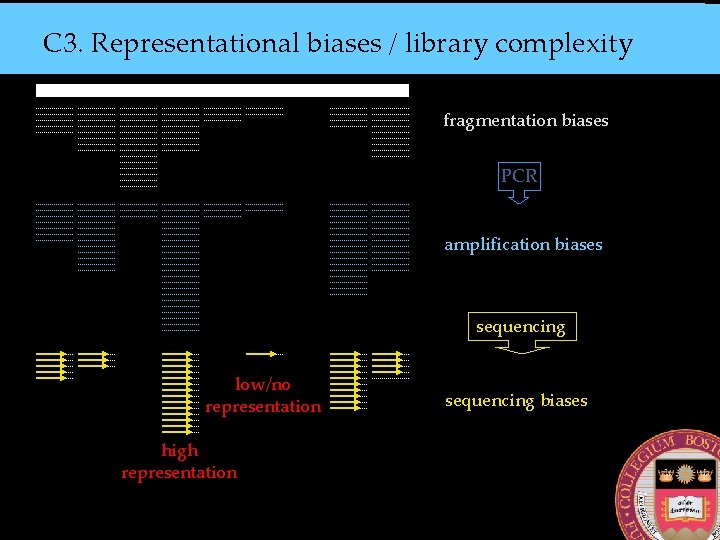
C 3. Representational biases / library complexity fragmentation biases PCR amplification biases sequencing low/no representation high representation sequencing biases
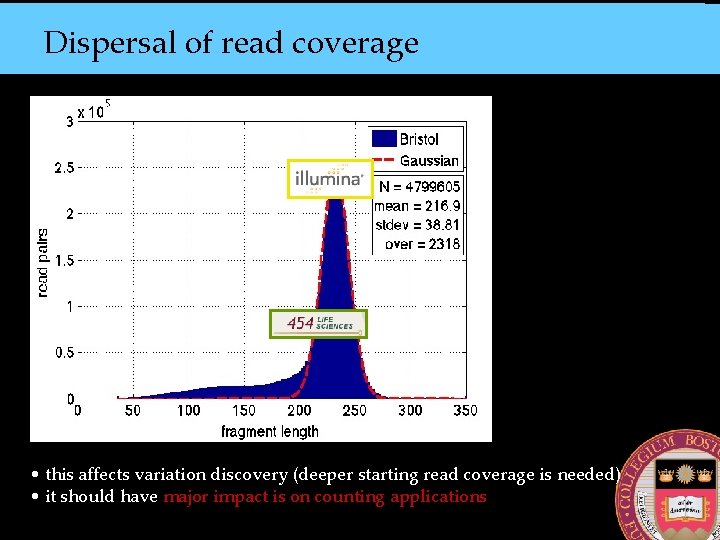
Dispersal of read coverage • this affects variation discovery (deeper starting read coverage is needed) • it should have major impact is on counting applications
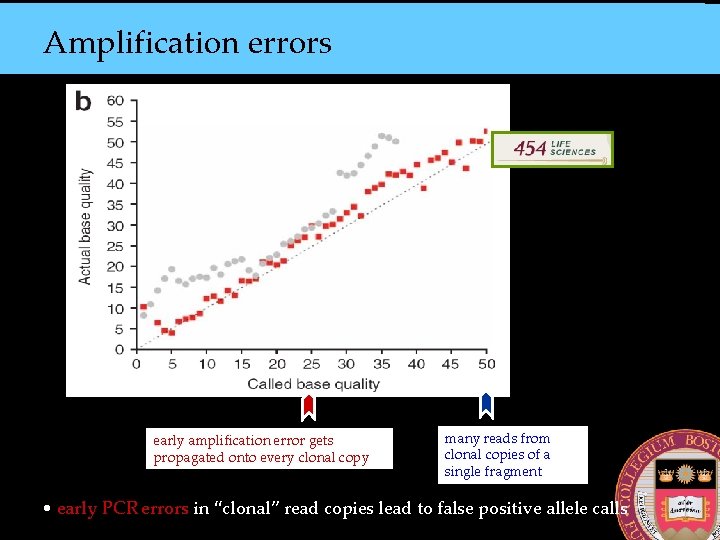
Amplification errors early amplification error gets propagated onto every clonal copy many reads from clonal copies of a single fragment • early PCR errors in “clonal” read copies lead to false positive allele calls
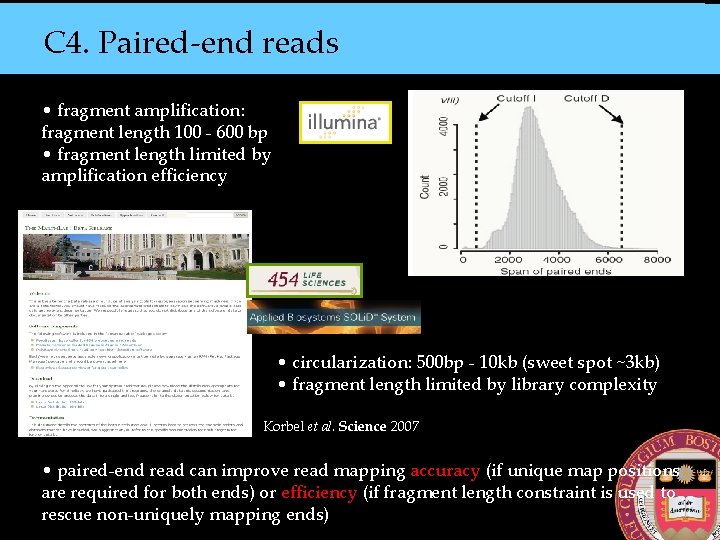
C 4. Paired-end reads • fragment amplification: fragment length 100 - 600 bp • fragment length limited by amplification efficiency • circularization: 500 bp - 10 kb (sweet spot ~3 kb) • fragment length limited by library complexity Korbel et al. Science 2007 • paired-end read can improve read mapping accuracy (if unique map positions are required for both ends) or efficiency (if fragment length constraint is used to rescue non-uniquely mapping ends)
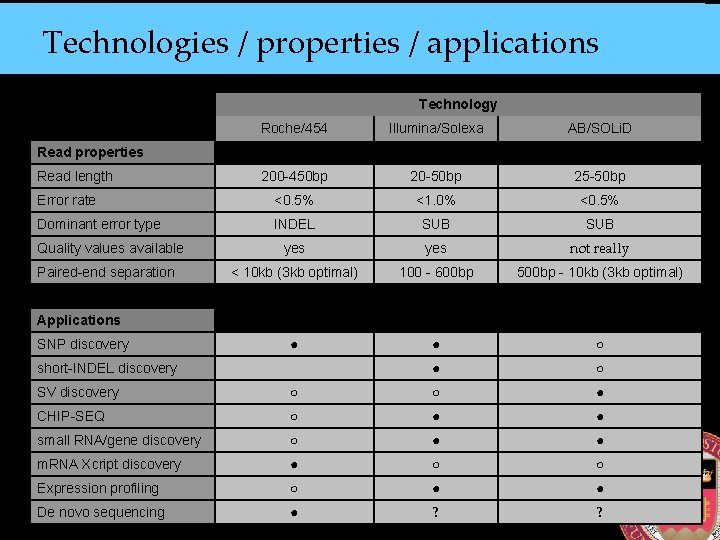
Technologies / properties / applications Technology Roche/454 Illumina/Solexa AB/SOLi. D 200 -450 bp 20 -50 bp 25 -50 bp Error rate <0. 5% <1. 0% <0. 5% Dominant error type INDEL SUB yes not really < 10 kb (3 kb optimal) 100 - 600 bp 500 bp - 10 kb (3 kb optimal) Applications SNP discovery ● ● ○ short-INDEL discovery ● ○ SV discovery ○ ○ ● CHIP-SEQ ○ ● ● small RNA/gene discovery ○ ● ● m. RNA Xcript discovery ● ○ ○ Expression profiling ○ ● ● De novo sequencing ● ? ? Read properties Read length Quality values available Paired-end separation
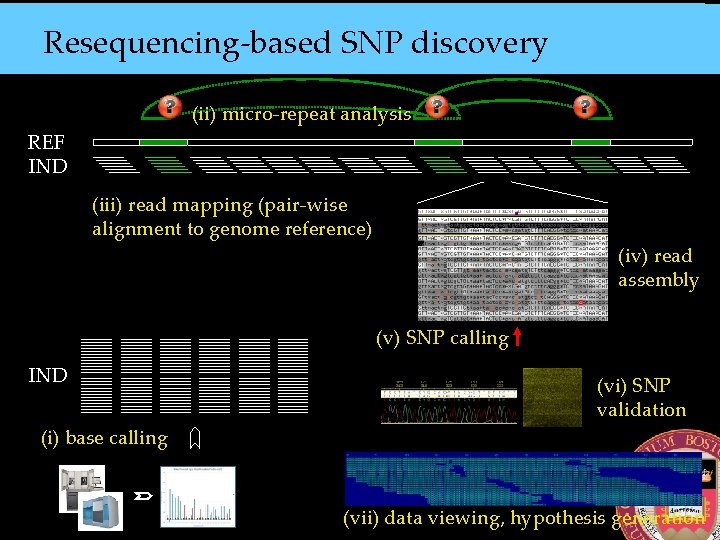
Resequencing-based SNP discovery (ii) micro-repeat analysis REF IND (iii) read mapping (pair-wise alignment to genome reference) (iv) read assembly (v) SNP calling IND (vi) SNP validation (i) base calling (vii) data viewing, hypothesis generation
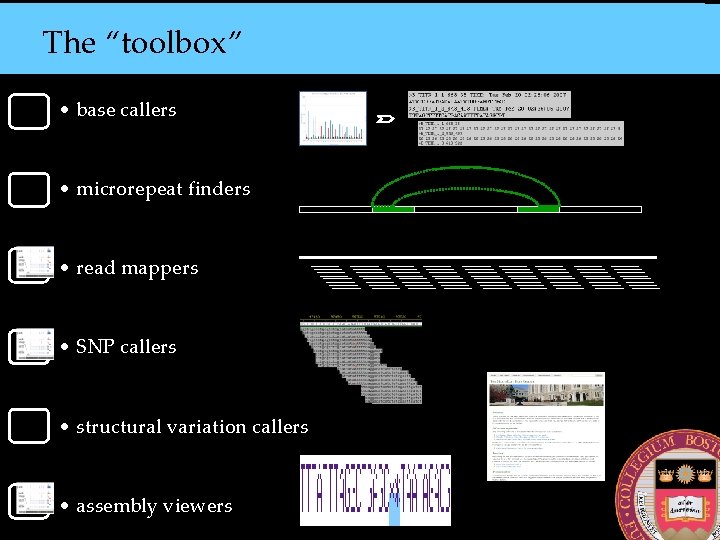
The “toolbox” • base callers • microrepeat finders • read mappers • SNP callers • structural variation callers • assembly viewers
Reference guided read mapping Reference-sequence guided mapping: …you get the pieces… …AND they give you the cover on the box Some pieces are more unique than others

MOSAIK: an anchored aligner / assembler Step 1. initial short-hash scan for possible read locations Step 2. evaluation of candidate locations with SW method Michael Stromberg

Non-unique mapping, gapped alignments 1. Non-unique read mapping: optionally either only report uniquely mapped reads or report all map locations for each read (mapping quality values for all mapped reads are being implemented) 2. Gapped alignments: allow for mapping reads with insertion or deletion sequencing errors, and reads with bona fide INDEL alleles
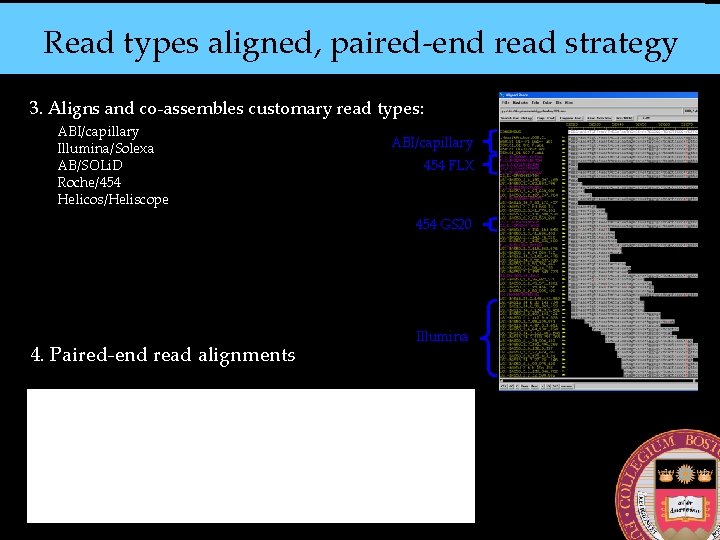
Read types aligned, paired-end read strategy 3. Aligns and co-assembles customary read types: ABI/capillary Illumina/Solexa AB/SOLi. D Roche/454 Helicos/Heliscope ABI/capillary 454 FLX 454 GS 20 4. Paired-end read alignments Illumina

Other mainstream read mappers • ELAND (Tony Cox, Illumina) -- the “official” read mapper supplied by Illumina, fast • MAQ (Li Heng + Richard Durbin, Sanger) -- the most widely used read mapper, low RAM footprint • SOAP (Beijing Genomics Institute) -- a new mapper developed for human next-gen reads • SHRIMP (Michael Brudno, University of Toronto) -- full Smith-Waterman
Speed
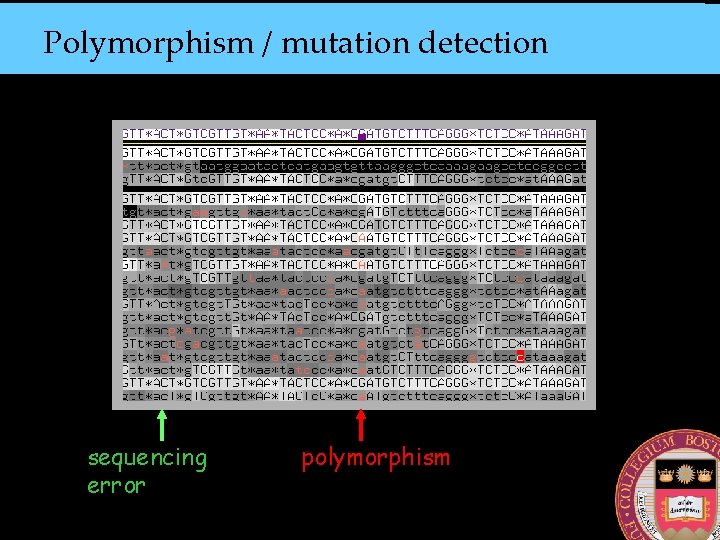
Polymorphism / mutation detection sequencing error polymorphism
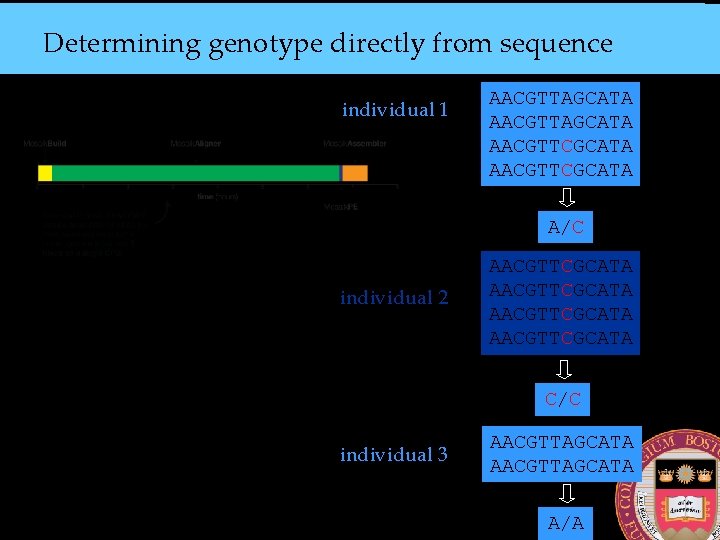
Determining genotype directly from sequence individual 1 AACGTTAGCATA AACGTTCGCATA A/C individual 2 AACGTTCGCATA C/C individual 3 AACGTTAGCATA A/A
Software SNP INS
Data visualization 1. aid software development: integration of trace data viewing, fast navigation, zooming/panning 2. facilitate data validation (e. g. SNP validation): co-viewing of multiple read types, quality value displays 3. promote hypothesis generation: integration of annotation tracks Weichun Huang

Applications 1. SNP discovery in shallow, single-read 454 coverage (Drosophila melanogaster) 2. SNP and INDEL discovery in deep Illumina short-read coverage (Caenorhabditis elegans) 3. Mutational profiling in deep 454 and Illumina read data (Pichia stipitis) (image from Nature Biotech. )
Our software is available for testing http: //bioinformatics. bc. edu/marthlab/Beta_Release

Credits Elaine Mardis (Washington University) Andy Clark (Cornell University) Doug Smith (Agencourt) Research supported by: NHGRI (G. T. M. ) BC Presidential Scholarship (A. R. Q. ) Michael Stromberg Michele Busby Eric Tsung Aaron Quinlan Derek Barnett Chip Stewart Damien Croteau-Chonka Weichun Huang http: //bioinformatics. bc. edu/marthlab
Accuracy • As is the case for all heuristic alignment algorithms accuracy and speed are option- and parameter-dependent

C 3. Quality values are important for allele calling • PHRED base quality values represent the estimated likelihood of sequencing error and help us pick out true alternate alleles • inaccurate or not well calibrated base quality values hinder allele calling Q-values should be accurate … and high!

Software tools for next-gen sequence analysis

Next-generation sequencing technologies and applications
- Slides: 41