Dairy Cattle Breeding in the United States John

Dairy Cattle Breeding in the United States John B. Cole Animal Improvement Programs Laboratory Agricultural Research Service, USDA, Beltsville, MD jcole@aipl. arsusda. gov 2006 2004

U. S. dairy statistics (2004) w 9. 0 million cows w 67, 000 herds w 135 cows/herd w 19, 000 lb (8600 kg)/cow w ~93% Holsteins, ~5% Jerseys w ~75% bred AI w 46% milk recorded through Dairy Herd Improvement (DHI) CSU 2006 – Departmental Seminar (2) Cole 2006 2004
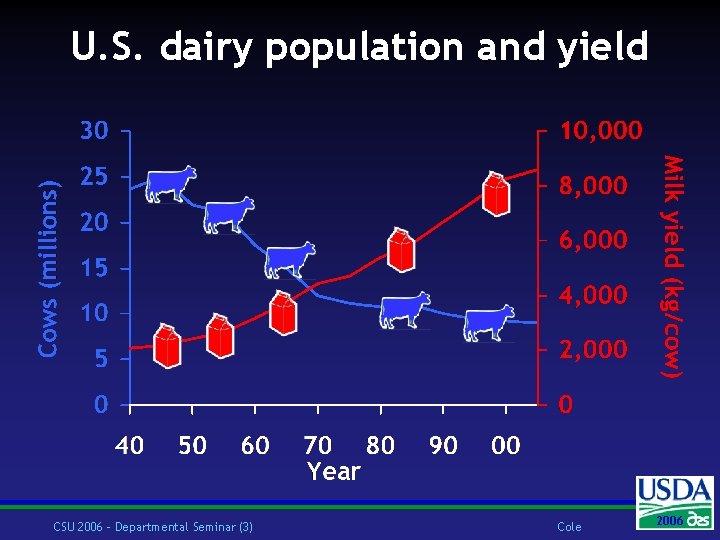
U. S. dairy population and yield CSU 2006 – Departmental Seminar (3) Cole 2006 2004

DHI statistics (2004) w 4. 1 million cows w 97% fat recorded w 93% protein recorded w 93% SCC recorded w 25, 000 herds w 164 cows/herd w 21, 250 lb (9640 kg)/cow w 3. 69% fat w 3. 09% (true) protein CSU 2006 – Departmental Seminar (4) Cole 2006 2004

U. S. progeny-test bulls (2000) w Major and marketing-only AI organizations plus breeder-proven w Breeds w. Ayrshire w. Brown Swiss w. Guernsey w. Holstein w. Jersey w. Milking Shorthorn CSU 2006 – Departmental Seminar (5) 10 bulls 53 bulls 15 bulls 1436 bulls 116 bulls 1 bull Cole 2006 2004
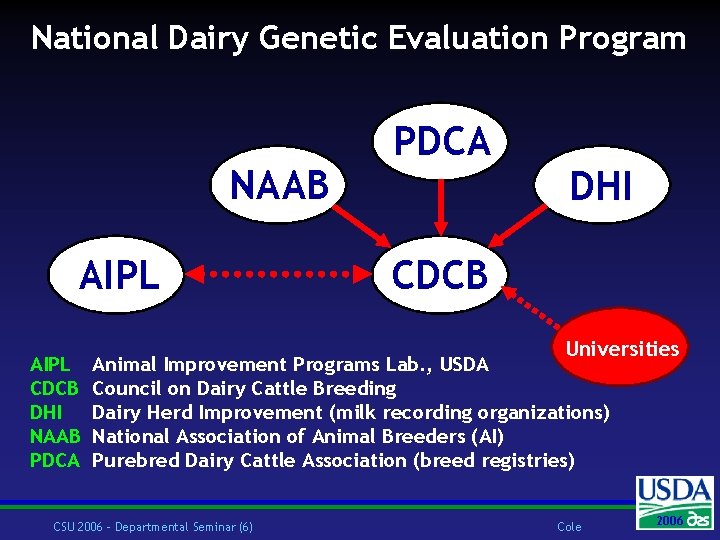
National Dairy Genetic Evaluation Program NAAB AIPL CDCB DHI NAAB PDCA DHI CDCB Universities Animal Improvement Programs Lab. , USDA Council on Dairy Cattle Breeding Dairy Herd Improvement (milk recording organizations) National Association of Animal Breeders (AI) Purebred Dairy Cattle Association (breed registries) CSU 2006 – Departmental Seminar (6) Cole 2006 2004

AIPL mission w Conduct research to discover, test, and implement improved genetic evaluation techniques for economically important traits of dairy cattle and goats w Genetically improve efficiency of dairy animals for yield and fitness CSU 2006 – Departmental Seminar (7) Cole 2006 2004

AIPL research objectives w Maintain a national database with animal identification, production, fitness, reproduction, and health traits to support research on dairy genetics and management w Provide data to others researchers submitting proposals compatible with industry needs CSU 2006 – Departmental Seminar (8) Cole 2006 2004

AIPL research objectives (cont. ) w Increase accuracy of genetic evaluations for traits through improved methodology and through inclusion and appropriate weighting of deviant data w Develop bioinformatic tools to automate data processing in support of quantitative trait locus detection, marker testing, and mapping methods CSU 2006 – Departmental Seminar (9) Cole 2006 2004

AIPL research objectives (cont. ) w Improve genetic rankings for overall economic merit by evaluating appropriate traits and by determining economic values of those traits in the index w Improved profit functions are derived from reviewing incomes and expenses associated with each trait available for selection CSU 2006 – Departmental Seminar (10) Cole 2006 2004

AIPL research objectives (cont. ) w Characterize dairy industry practices in milk recording, breed registry, and artificial-insemination to document status and changes in data collection and use and in observed and genetic trends in the population CSU 2006 – Departmental Seminar (11) Cole 2006 2004

Traits evaluated w Yield (milk, fat, protein volume; component percentages) w Type/conformation w Productive life/longevity w Somatic cell score/mastitis resistance w Fertility w w Daughter pregnancy rate (cow) Estimated relative conception rate (bull) w Dystocia and stillbirth (service sire, daughter) CSU 2006 – Departmental Seminar (12) Cole 2006 2004

Evaluation methods w Animal model (linear) w Yield (milk, fat, protein) w Type Heritability 25 – 40% 7 – 54% (Ayrshire, Brown Swiss, Guernsey, Jersey) w Productive life w SCS w Daughter pregnancy rate 8. 5% 12% 4% w Sire – maternal grandsire model (threshold) w Service sire calving ease w Daughter calving ease w Service sire stillbirth w Daughter stillbirth CSU 2006 – Departmental Seminar (13) 8. 6% 3. 0% 6. 5% Cole 2006 2004
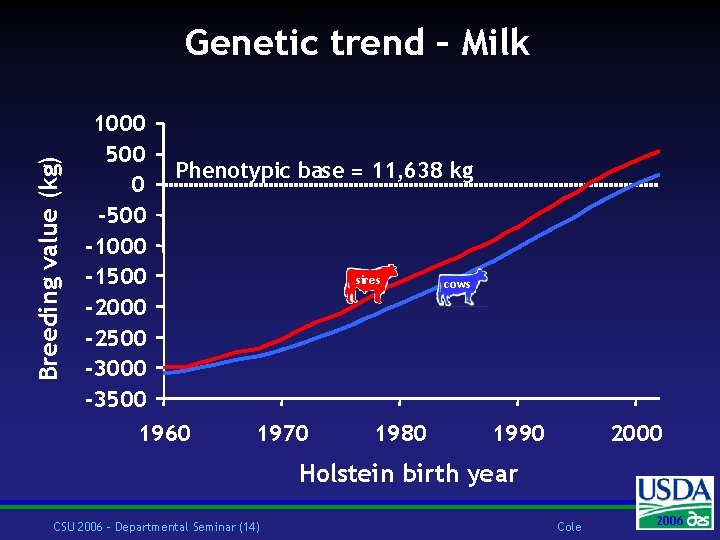
Breeding value (kg) Genetic trend – Milk 1000 500 0 -500 -1000 -1500 -2000 -2500 -3000 -3500 Phenotypic base = 11, 638 kg 1960 sires 1970 1980 cows 1990 2000 Holstein birth year CSU 2006 – Departmental Seminar (14) Cole 2006 2004
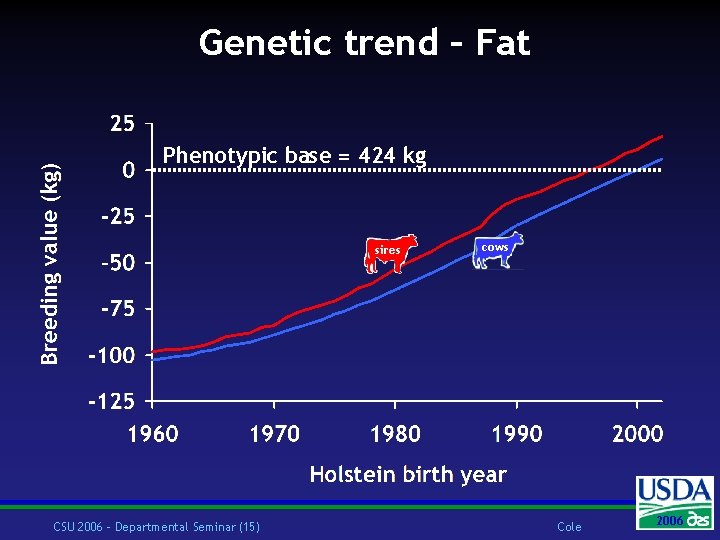
Genetic trend – Fat Phenotypic base = 424 kg sires CSU 2006 – Departmental Seminar (15) cows Cole 2006 2004
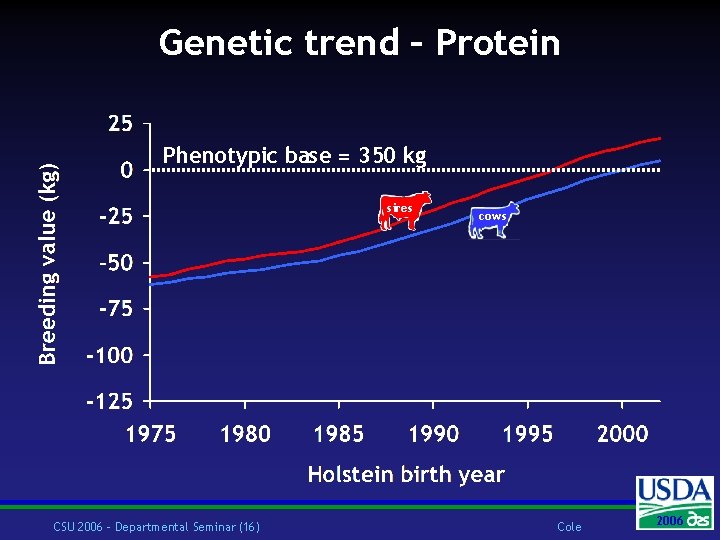
Genetic trend – Protein Phenotypic base = 350 kg sires CSU 2006 – Departmental Seminar (16) cows Cole 2006 2004
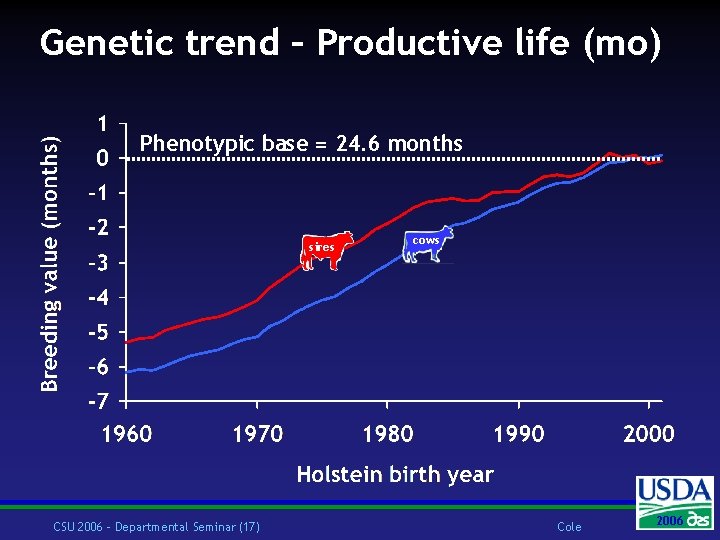
Genetic trend – Productive life (mo) Phenotypic base = 24. 6 months sires CSU 2006 – Departmental Seminar (17) cows Cole 2006 2004
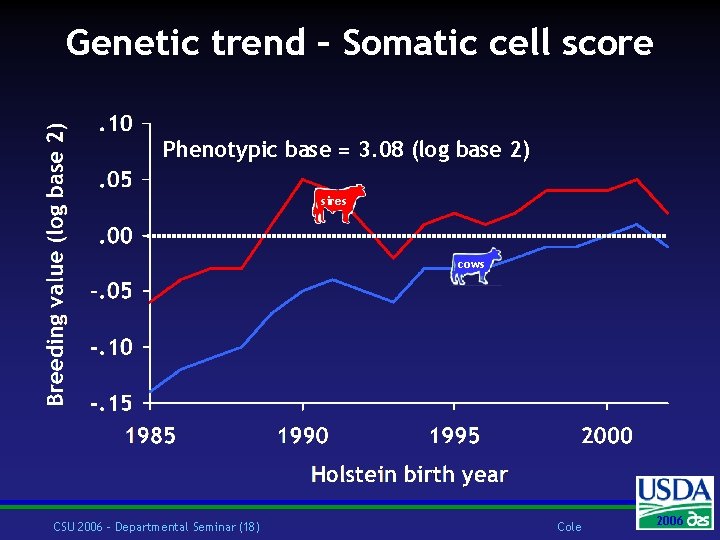
Genetic trend – Somatic cell score Phenotypic base = 3. 08 (log base 2) sires cows CSU 2006 – Departmental Seminar (18) Cole 2006 2004
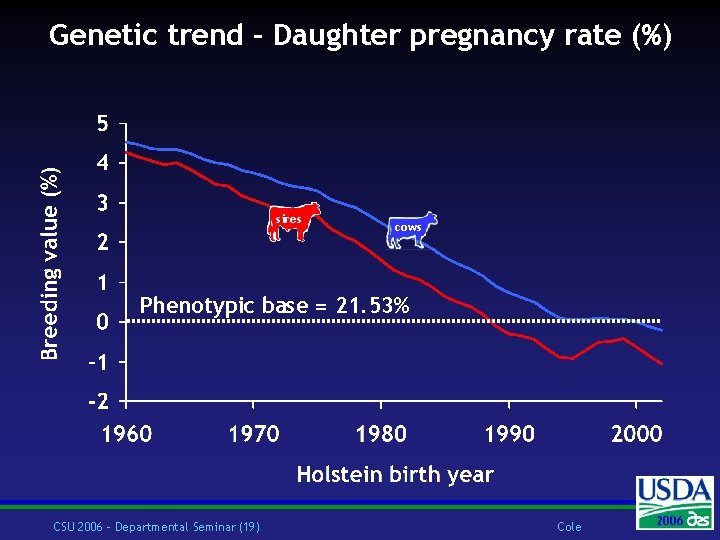
Genetic trend – Daughter pregnancy rate (%) sires cows Phenotypic base = 21. 53% CSU 2006 – Departmental Seminar (19) Cole 2006 2004
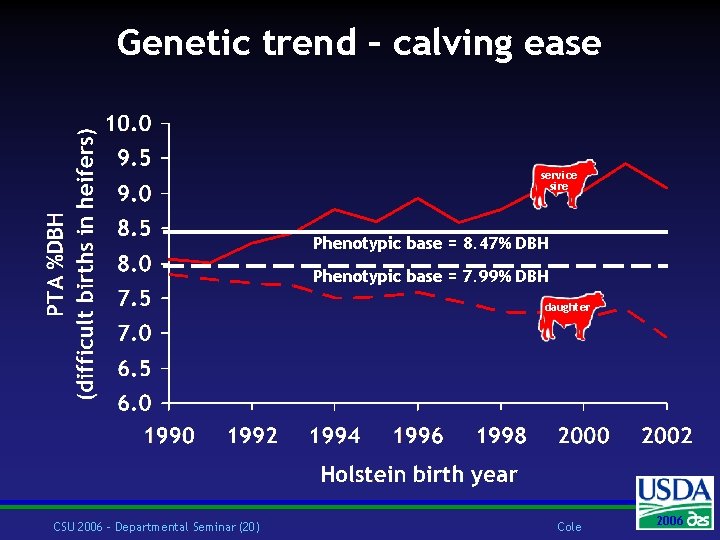
Genetic trend – calving ease service sire Phenotypic base = 8. 47% DBH Phenotypic base = 7. 99% DBH daughter CSU 2006 – Departmental Seminar (20) Cole 2006 2004
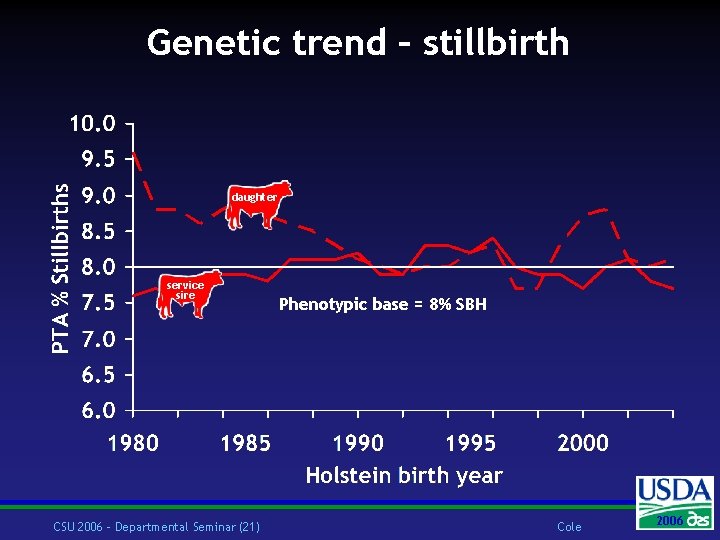
Genetic trend – stillbirth daughter service sire CSU 2006 – Departmental Seminar (21) Phenotypic base = 8% SBH Cole 2006 2004
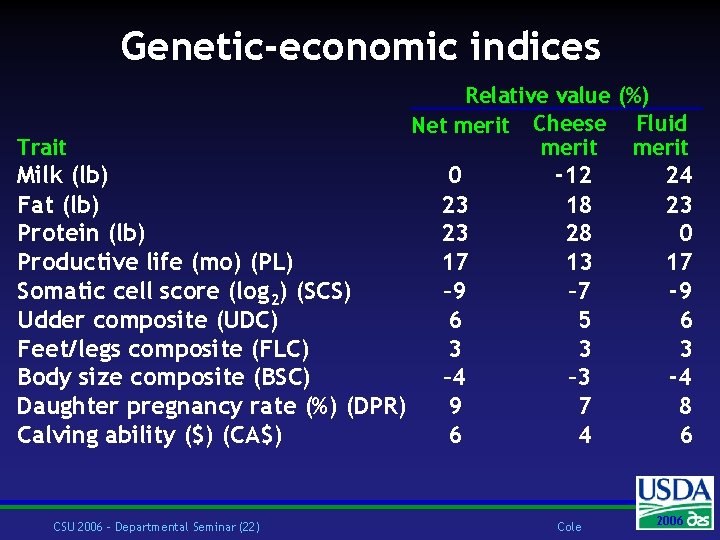
Genetic-economic indices Trait Milk (lb) Fat (lb) Protein (lb) Productive life (mo) (PL) Somatic cell score (log 2) (SCS) Udder composite (UDC) Feet/legs composite (FLC) Body size composite (BSC) Daughter pregnancy rate (%) (DPR) Calving ability ($) (CA$) CSU 2006 – Departmental Seminar (22) Relative value (%) Net merit Cheese Fluid merit 0 23 23 17 – 9 6 3 – 4 9 6 -12 18 28 13 – 7 5 3 – 3 7 4 Cole 24 23 0 17 -9 6 3 -4 8 6 2004
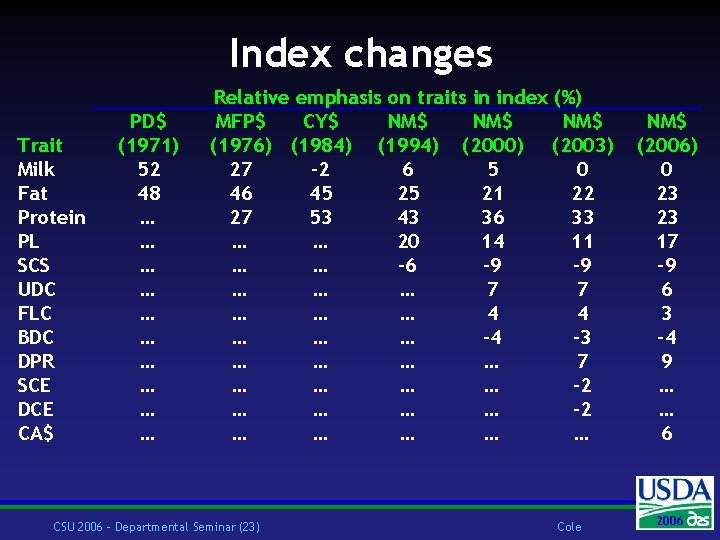
Index changes Trait Milk Fat Protein PL SCS UDC FLC BDC DPR SCE DCE CA$ PD$ (1971) 52 48 … … … … … Relative emphasis on traits in index (%) MFP$ CY$ NM$ NM$ (1976) (1984) (1994) (2000) (2003) 27 – 2 6 5 0 46 45 25 21 22 27 53 43 36 33 … … 20 14 11 … … – 6 – 9 … … … 7 7 … … … 4 4 … … … – 4 – 3 … … 7 … … … … – 2 … … … CSU 2006 – Departmental Seminar (23) Cole NM$ (2006) 0 23 23 17 – 9 6 3 – 4 9 … … 6 2004

Persistency CSU 2006 – Departmental Seminar (24) Cole 2006 2004

Introduction w At the same level of production cows with high persistency milk more at the end than the beginning of lactation w Best prediction of persistency is calculated as a function of traitspecific standard lactation curves and the linear regression of a cow’s test day deviations on days in milk CSU 2006 – Departmental Seminar (25) Cole 2006 2004

Best Prediction w Selection Index w Predict missing yields from measured yields w Condense daily into lactation yield and persistency w Only phenotypic covariances are needed w Mean and variance of herd assumed known w Reverse prediction w Daily yield predicted from lactation yield and persistency CSU 2006 – Departmental Seminar (26) Cole 2006 2004

Persistency Cole and Van. Raden 2006 JDS 89: 2722 -2728 w Definition w 305 daily yield deviations (DIM - DIMo) w Uncorrelated with yield by choosing DIMo were 161, 159, 166, and 155 for M, F, P, and SCS • DIM 0 have increased over time w Standardized estimate CSU 2006 – Departmental Seminar (27) Cole 2006 2004
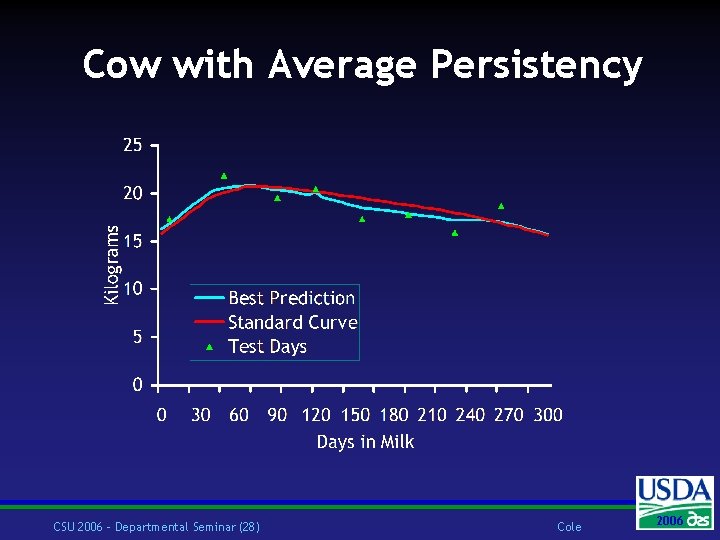
Cow with Average Persistency CSU 2006 – Departmental Seminar (28) Cole 2006 2004
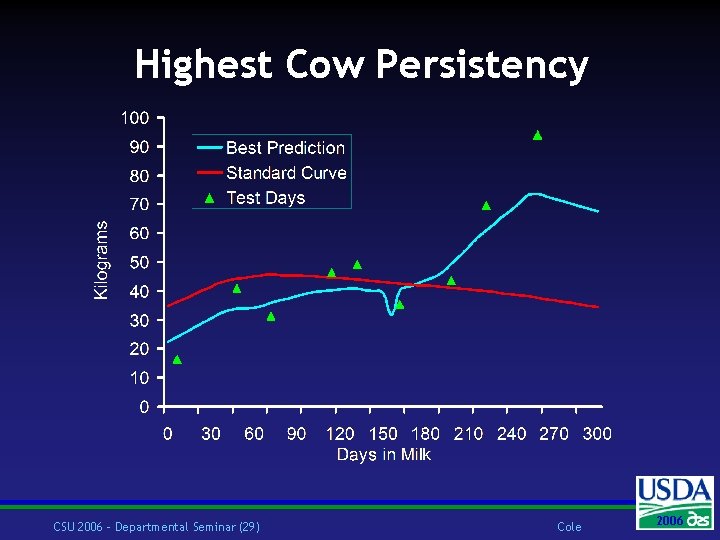
Highest Cow Persistency CSU 2006 – Departmental Seminar (29) Cole 2006 2004
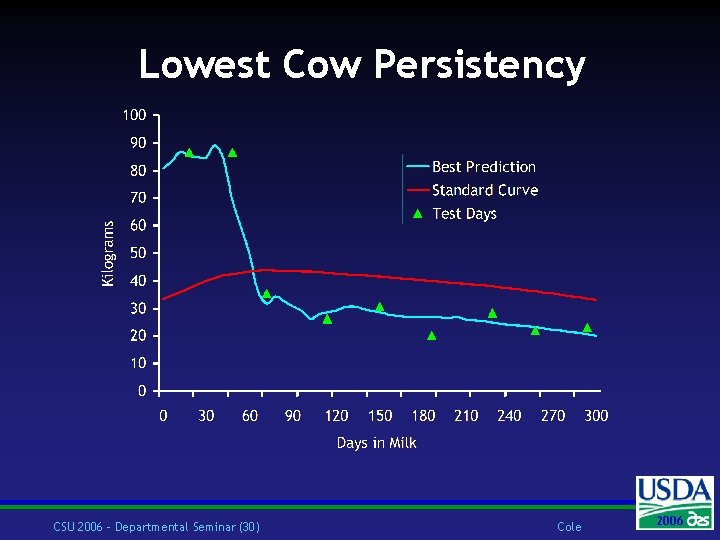
Lowest Cow Persistency CSU 2006 – Departmental Seminar (30) Cole 2006 2004

Model Repeatability animal model: yijkl = hysi + lacj + ak + pek + β(dojk) + eijkl yijkl hysi lac j ak pek dojk eijkl = = = = persistency of milk, fat, protein, or SCS fixed effect of herd-year-season of calving I fixed effect of lactation j random additive genetic effect of animal k random permanent environmental effect of animal k days open for lactation j of animal k random residual error CSU 2006 – Departmental Seminar (31) Cole 2006 2004
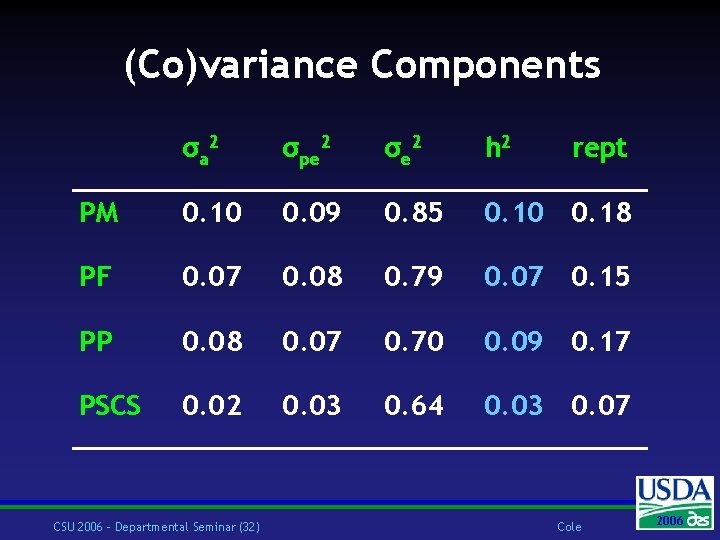
(Co)variance Components σa 2 σpe 2 σe 2 h 2 PM 0. 10 0. 09 0. 85 0. 10 0. 18 PF 0. 07 0. 08 0. 79 0. 07 0. 15 PP 0. 08 0. 07 0. 70 0. 09 0. 17 PSCS 0. 02 0. 03 0. 64 0. 03 0. 07 CSU 2006 – Departmental Seminar (32) rept Cole 2006 2004
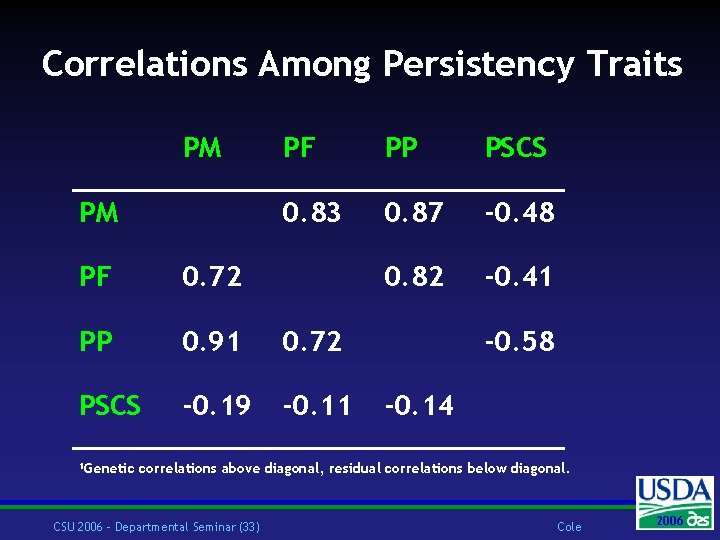
Correlations Among Persistency Traits PM PM PF PP PSCS 0. 83 0. 87 -0. 48 0. 82 -0. 41 PF 0. 72 PP 0. 91 0. 72 PSCS -0. 19 -0. 11 1 Genetic -0. 58 -0. 14 correlations above diagonal, residual correlations below diagonal. CSU 2006 – Departmental Seminar (33) Cole 2006 2004
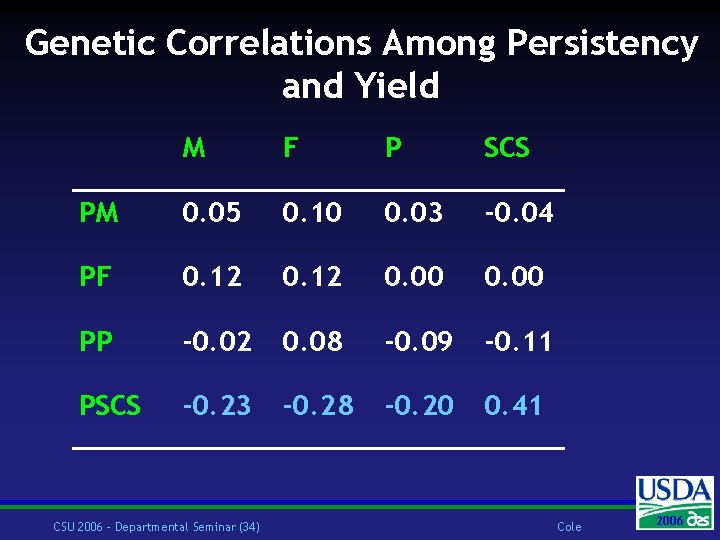
Genetic Correlations Among Persistency and Yield M F P SCS PM 0. 05 0. 10 0. 03 -0. 04 PF 0. 12 0. 00 PP -0. 02 0. 08 -0. 09 -0. 11 PSCS -0. 23 -0. 28 -0. 20 0. 41 CSU 2006 – Departmental Seminar (34) Cole 2006 2004

Factors Affecting Persistency w Parity: 1 st lactation cows tend to have flatter lactation curves than later lactation cows w Nutrition: underfeeding energy will reduce peak yield, leading to higher persistency w Stress: low persistency in cows under handling or heat stress w Diseases? w Breed differences? CSU 2006 – Departmental Seminar (35) Cole 2006 2004

Summary w Heritabilities and repeatabilities are low to moderate w Routine genetic evaluations for persistency are feasible w The shape of the lactation curve may be altered without affecting production CSU 2006 – Departmental Seminar (36) Cole 2006 2004

Diseases and Persistency Appuhamy, Cassell, and Cole 2006 w Other measures may improve disease resistance through indirect selection, e. g. productive life (PL), body condition scores, and persistency w Studies of the effect of diseases on milk yield is abundant in literature w Investigations of relationships between diseases and other traits are lacking (Muir et al. , 2004) CSU 2006 – Departmental Seminar (37) Cole 2006 2004

Objectives w Investigate the effect of common health disorders on persistency w Estimate phenotypic correlations among diseases and persistency w Measure breed effects (Holstein and Jersey) on these relationships CSU 2006 – Departmental Seminar (38) Cole 2006 2004

Materials and Methods Daily milk yield records of Holstein and Jersey cows at the Virginia Tech Dairy Complex from 07/18/2004 to 06/07/2006 Holstein First lactation (L 1) 41 Second lactation (L 2) 34 Third and later (L 3+) 40 Total Lactations 115 Total cows 93 CSU 2006 – Departmental Seminar (39) Jersey 10 08 15 33 33 Cole 2006 2004

Definition of Disease Variables Mastitis (MAST) : All causes of udder infections MAST 1 : in first 100 days (stage 1) MAST 2 : after 100 th DIM (stage 2) Post Partum Metabolic Diseases (METAB): Milk fever and/or ketosis Other diseases: LAME, DA, MET, PNEU, DIARR CSU 2006 – Departmental Seminar (40) Cole 2006 2004

Statistical Analysis where: Pijklm = Li + Yj +Dk + Ol + eijklm Yijklm = Lactation persistency of cow m Li = Effect of ith lactation (i = 1, 2, & 3) YSj = Effect of jth calving year-season ( j=1, 2, 3, 4, 5 & 6) Dk = Effect of kth status of the disease ( k =1 or 0) Ol = Effect of lth status of other diseases (l=1 or 0) eijklm = residual effect (Other diseases includes all diseases beside the disease of interest. ) CSU 2006 – Departmental Seminar (41) Cole 2006 2004
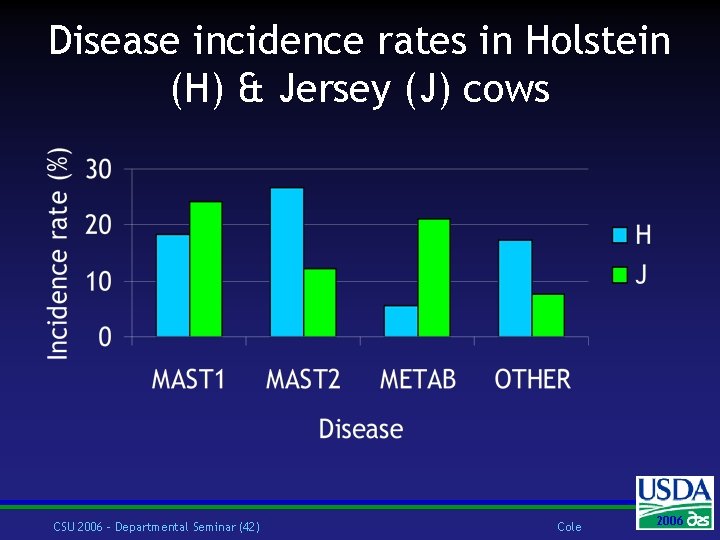
Disease incidence rates in Holstein (H) & Jersey (J) cows CSU 2006 – Departmental Seminar (42) Cole 2006 2004
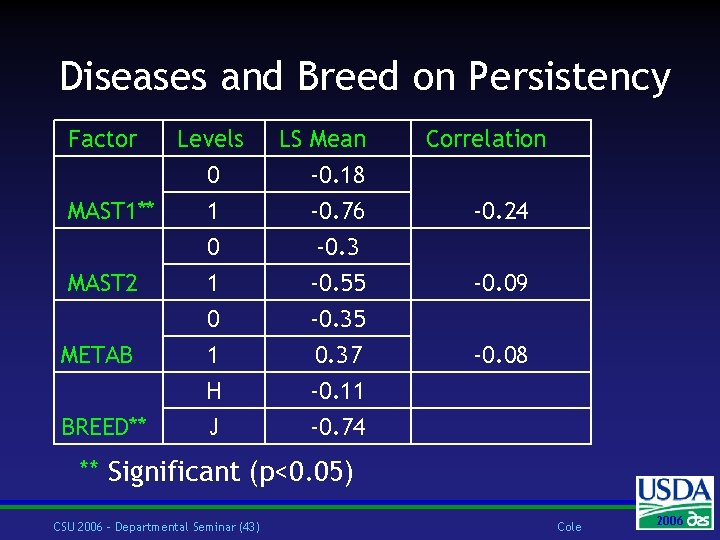
Diseases and Breed on Persistency Factor Levels 0 1 MAST 1** 0 MAST 2 METAB BREED** 1 0 1 H J LS Mean -0. 18 -0. 76 -0. 3 -0. 55 -0. 35 0. 37 -0. 11 -0. 74 Correlation -0. 24 -0. 09 -0. 08 ** Significant (p<0. 05) CSU 2006 – Departmental Seminar (43) Cole 2006 2004

Conclusions w Mastitis in early lactation has a significant, negative effect on persistency w Mastitis in late lactation and post partum metabolic diseases have nonsignificant, but negative, effects on persistency w Persistency differs significantly between Holstein and Jersey cows CSU 2006 – Departmental Seminar (44) Cole 2006 2004

All-Breeds Evaluation CSU 2006 – Departmental Seminar (45) Cole 2006 2004

Goals w Evaluate crossbred animals without biasing purebred evaluations w Accurately estimate breed differences w Compute national evaluations and examine changes w PTA of purebreds and crossbreds w Changes in reliability w Display results without confusion CSU 2006 – Departmental Seminar (46) Cole 2006 2004

Methods w All-breed animal model w Purebreds and crossbreds together w Unknown parents grouped by breed w Variance adjustments by breed w Age adjust to 36 months, not mature w 1988 software, good convergence w Within-breed-of-sire model examined but not used CSU 2006 – Departmental Seminar (47) Cole 2006 2004

Unknown Parent Groups w Groups formed based on w Birth year (flexible) w Breed (must have >10, 000 cows) w Path (dams of cows, sires of cows, parents of bulls) w Origin (domestic vs other countries) w Paths have >1000 in last 15 years w Groups each have >500 animals CSU 2006 – Departmental Seminar (48) Cole 2006 2004

Data w Numbers of cows of all breeds w 22. 6 w 16. 1 w 22. 5 w 19. 9 w 10. 5 million million for for for milk and fat protein productive life daughter pregnancy rate somatic cell score w Type evaluated in separate breed files w Calving ease joint HO, BS, and HO x BS w Goats in all-breed model since 1988 CSU 2006 – Departmental Seminar (49) Cole 2006 2004
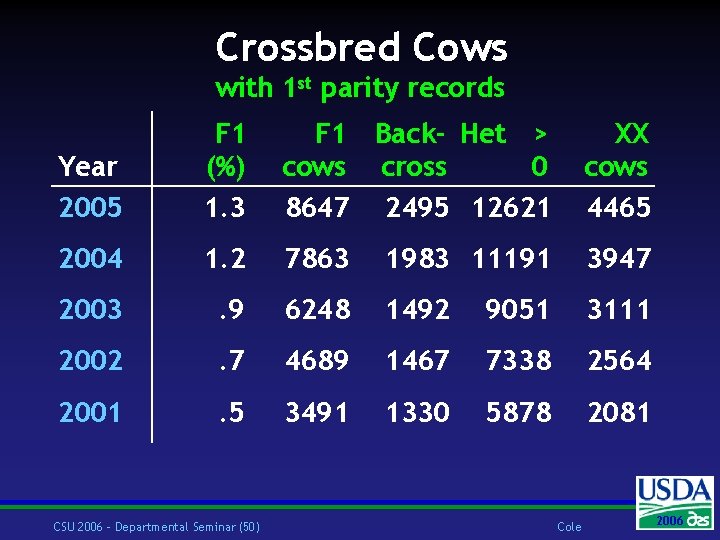
Crossbred Cows with 1 st parity records Year 2005 F 1 (%) 1. 3 F 1 Back- Het > cows cross 0 8647 2495 12621 XX cows 4465 2004 1. 2 7863 1983 11191 3947 2003 . 9 6248 1492 9051 3111 2002 . 7 4689 1467 7338 2564 2001 . 5 3491 1330 5878 2081 CSU 2006 – Departmental Seminar (50) Cole 2006 2004

Reliability w Crossbred cows w Will have PTA, most did not before w Accurate PTA from both parents w Purebred animals w Information from crossbred relatives w More contemporaries CSU 2006 – Departmental Seminar (51) Cole 2006 2004
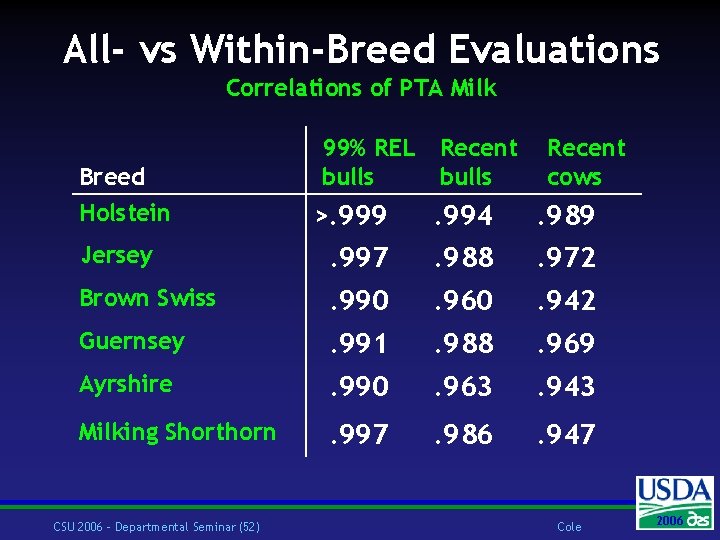
All- vs Within-Breed Evaluations Correlations of PTA Milk Breed Holstein 99% REL bulls Recent cows >. 999 . 994 . 989 Jersey . 997 . 988 . 972 Brown Swiss . 990 . 960 . 942 Guernsey . 991 . 988 . 969 Ayrshire . 990 . 963 . 943 Milking Shorthorn . 997 . 986 . 947 CSU 2006 – Departmental Seminar (52) Cole 2006 2004

Display of PTAs w Genetic base w Convert all-breed base back to within-breed -of-sire bases w Each animal gets just one PTA w PTAbrd = (PTAall – meanbrd) SDbrd/ SDall w Heterosis and inbreeding w Both effects removed in the animal model w Heterosis added to crossbred animal PTA w Expected Future Inbreeding (EFI) and merit differ with mate breed CSU 2006 – Departmental Seminar (53) Cole 2006 2004

Schedule w Interbull test run Feb. 1, 2006 w Trend validation w Convert all-breed PTA back to withinbreed bases w Scientific publication (JDS) w Implementation w Expected May 2007 CSU 2006 – Departmental Seminar (54) Cole 2006 2004

Conclusions w All breed model accounts for: w General heterosis w Unknown parent groups by breed w Heterogeneous variance by breed w PTA converted back to within breed bases, crossbreds to breed of sire w PTA changes more in breeds with fewer animals CSU 2006 – Departmental Seminar (55) Cole 2006 2004

Genomics CSU 2006 – Departmental Seminar (56) Cole 2006 2004

SNP Project Outcomes w Genome-wide selection w Parentage verification & traceability panels w Enhanced QTL mapping & gene discovery CSU 2006 – Departmental Seminar (57) Cole 2006 2004

Linkage disequilibrium (LD) w Non-random association of alleles at two or more loci, not necessarily on the same chromosome w Not the same as linkage, which describes the association of two or more loci on a chromosome with limited recombination between them CSU 2006 – Departmental Seminar (58) Cole 2006 2004
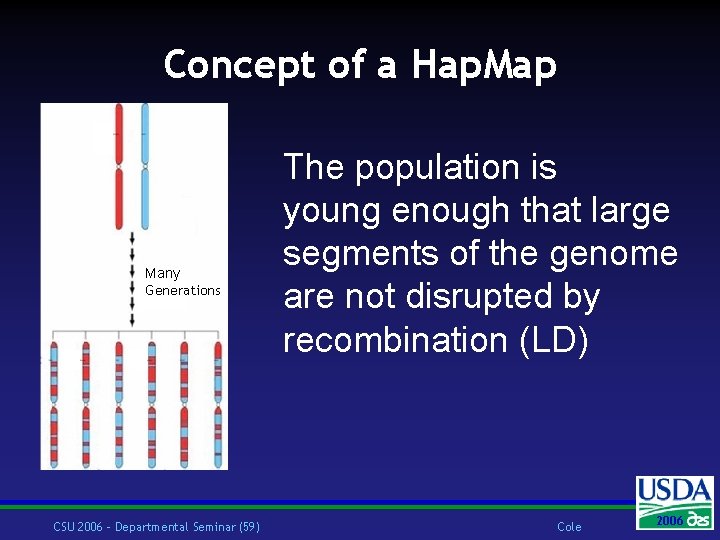
Concept of a Hap. Map Many Generations CSU 2006 – Departmental Seminar (59) The population is young enough that large segments of the genome are not disrupted by recombination (LD) Cole 2006 2004

Genome Selection w Animals are genotyped at birth w Genomic EBV calculated for many traits w Even those not typically recorded (e. g. semen quality) w Accuracy is predicted to be similar to progeny test evaluation CSU 2006 – Departmental Seminar (60) Cole 2006 2004

Advantages of Genome Selection w Generation intervals can be reduced w Costs of progeny testing can be decreased w More accurate selection among full sibs w Decreased risk in selection program CSU 2006 – Departmental Seminar (61) Cole 2006 2004

Low-cost parentage verification w SNP tests may make parentage validation cheap enough for widespread adoption w Develop a database and software to check parentage and suggest alternatives for invalid IDs w Determine rate of parentage errors in a sample of herds CSU 2006 – Departmental Seminar (62) Cole 2006 2004

In Conclusion CSU 2006 – Departmental Seminar (63) Cole 2006 2004

Ongoing Work w New traits w Stillbirth (HOL) w Milking speed (BSW) w Rear legs/rear view (BSW, GUE) w Bull fertility (transferred from DRMS) w Improved online tools w Fully buzzword-compliant w Web services for data delivery w Choice of scales CSU 2006 – Departmental Seminar (64) Cole 2006 2004

Senior research staff CSU 2006 – Departmental Seminar (65) Cole 2006 2004
- Slides: 65