CSE 5290 Algorithms for Bioinformatics Fall 2011 Suprakash

CSE 5290: Algorithms for Bioinformatics Fall 2011 Suprakash Datta datta@cse. yorku. ca Office: CSEB 3043 Phone: 416 -736 -2100 ext 77875 Course page: http: //www. cs. yorku. ca/course/5290 10/27/2021 CSE 5290, Fall 2011 1

Biological (genomic) data ATATTGAATTTTCAAAAATTCTTACTTTTTGGACGCAAAGAAGTTTAATAATCATATTACATGGCATTACCACCATATCCATATCT AATCTTATATGTTGTGGAAATGTAAAGAGCCCCATTATCTTAGCCTAAAAAAACCTTCTCTTTGGAACTTTCTAATACGCTTAACTGCTCATT GCTATATTGAAGTACGGATTAGAAGCCGCCGAGCGGGCGACAGCCCTCCGACGGAAGACTCTCCTCTGCGTCCTCGTCTTCACCGGTCGCGTT CCTGAAACGCAGATGTGCCTCGCGCCGCACTGCTCCGAACAATAAAGATTCTACAATACTCTTTTATGGTTATGAAGAGGAAAAATTGGCAGTAA CCTGGCCCCACAAACCTTCAAATTAACGAATCAAATTAACAACCATAGGATGAATGCGATTAGTTTTTTAGCCTTATTTCTGGGGTAATCAG CGAAGCGATGATTTTTGATCTATTAACAGATATATAAATGGAAGCTGCATAACCACTTTAACTAATACTTTCAACATTTTCAGTTTGTATTACTTCTT ATTCAAATGTCATAAAAGTATCAACAAAAAATTTAATATACCTCTATACTTTAACGTCAAGGAGAAAAAACTATAATGACTAAATCTCATTCAGAAGA AGTGATTGTACCTGAGTTCAACTAGCGCAAAGGAATTACCAAGACCATTGGCCGAAAAGTGCCCGAGCATAATTAAGAAATTTATAAGCGCTTAT GATGCTAAACCGGTTTGTTGCTAGATCGCCTGGTAGAGTCAATCTAATTGGTGAACATATTGATTATTGTGACTTCTCGGTTTTACCTTTAGCTAT TGATATGCTTTGCGCCGTCAAAGTTTTGAACGAGAAAAATCCATTACCTTAATAAATGCTGATCCCAAATTTGCTCAAAGGAATCGAT TTGCCGTTGGACGGTTCTTATGTCACAATTGATCCTTCTGTGTCGGACTGGTCTAATTACTTTAAATGTGGTCTCCATGTTGCACTCTTTTCTAAA GAAACTTGCACCGGAAAGGTTTGCCAGTGCTCCTCTGGCCGGGCTGCAAGTCTTCTGTGAGGGTGATGTACCATGGCAGTGATTGTCTTCTTCG GCCGCATTTGTGCCGTTGCTTTAGCTGTTGTTAAAGCGAATATGGGCCCTGGTTATCATATCCAAGCAAAATTTAATGCGTATTACGGTCGT TGCAGAACATTATGTTGGTGTTAACAATGGCGGTATGGATCAGGCTGCCTCTGTTTGGTGAGGAAGATCATGCTCTATACGTTGAGTTCAAACCG CAGTTGAAGGCTACTCCGTTTAAATTTCCGCAATTAAAAAACCATGAATAGCTTTGTTATTGCGAACACCCTTGTTGTATCTAACAAGTTTGAAACC GCCCCAACTATAATTTAAGAGTGGTAGAAGTCACCAGCTGCAAATGTTTTAGCTGCCACGTACGGTGTTGTTTTACTTTCTGGAAAAGAAG GATCGAGCACGAATAAAGGTAATCTAAGAGTTCATGAACGTTTATTATGCCAGATATCACAACATTTCCACACCCTGGAACGGCGATATTGAATC CGGCATCGAACGGTTAACAAAGGCTAGTACTAGTTGAAGAGTCTCTCGCCAATAAGAAACAGGGCTTTAGTGTTGACGATGTCGCACAATCCTT GAATTGTTCTCGCGAAATTCACAAGAGACTACTTAACAACATCTCCAGTGAGATTTCAAGTCTTAAAGCTATATCAGAGGGCTAAGCATGTGTATT CTGAATTTAAGAGTCTTGAAGGCTGTGAAATTAATGACTACAGCGAGCTTTACTGCCGACGAAGACTTTTTCAAGCAATTTGGTGCCTTGATGCG AGTCTCAAGCTTCTTGCGATAAACTTTACGAATGTTCTTGTCCAGAGATTGACAAAATTTGTTCCATTGCTTTGTCAAATGGATCATGGTTCCCGTT TGACCGGAGCTGGGGTGGTTGTACTGTTCACTTGGTTCCAGGGGGCCCAAATGGCAACATAGAAAAGGTAAGAAGCCCTTGCCAATGAG TTCTACAAGGTCAAGTACCCTAAGATCACTGATGCTGAGCTAGAAAATGCTATCATCGTCTCTAAACCAATTGGGCAGCTGTCTATATGAATTATA AGTATACTTCTTTTACTTTGTTCAGAACAACTTCTCATTTTTTTCTACTCATAACTAGCATCACAAAATACGCAATAATAACGAGTAGTAACAC TTTTATAGTTCATACATGCTTCAACTACTTAATAAATGATTGTATGATAGTTTTCAATGTAAGAGATTTCGATTATCCACAAACTTTAAAACACAGGG ACAAAATTCTTGATATGCTTTCAACCGCTGCGTTTTGGACCTATTCTTGACATGATATGACTACCATTTTGTTATTGTACGTGGGGCAGTTGACGT CTTATCATATGTCAAAGTCATTTGCGAAGCTTGGCAAGTTGCCAACTGACGAGATGCAGTAAAAAGAGATTGCCGTCTTGAAACTTTTTGTCCTTT TTTTTTTCCGGGGACTCTACGAACCCTTTGTCCTACTGATTAATTTTGTACTGAATTTGGACAATTCAGATTTTAGTAGACAAGCGCGAGGAGGAA AAGAAATGACAAAAATTCCGATGGACAAGAAGATAGGAAAAAAGCTTTCACCGATTTCCTAGACCGGAAAAAAGTCGTATGACATCAGAA TGAAATTTTCAAGTTAGACAAGGACAAAATCAGGACAAATTGTAAAGATATAATAAACTATTTGATTCAGCGCCAATTTGCCCTTTTCCATT AAATCTCTGTTCTCTCTTATATGATGATTAGGTATCATCTGTATAAAACTCCTTTCTTAATTTCACTCTAAAGCATCCCATAGAGAAGATCTTT CGGTTCGAAGACATTCCTACGCATAATAAGAATAGGAGGGAATAATGCCAGACAATCTATCATTAGCGGCTCTTCAAAAAGATTGAACTCT CGCCAACTTATGGAATCTTCCAATGAGACCTTTGCGCCAAATAATGTGGATTTGGAAAAAGTATAAGTCATCTCAGAGTAATATAACTACCGAAGT TTATGAGGCATCGAGCTTTGAAGAAAAAGTAAGCTCAGAAAAACCTCAATAGCTCATTCTGGAAGAAAATCTATTATGAATATGTGGTCGTTGACA AATCTTGGGTGTTTCTATTCTGGATTCATTTATGTACACAGGACTTGAAGCCCGTCGAAAAAGGCGGGTTTGGTCCTGGTACAATTAT TGTTACTTCTGGCTTGCTGAATGTTTCAATATCCACTTGGCAAATTGCAGCTACAGGTCTACAACTGGGTCTAAATTGGTGGCAGTGTTGGATAA CAATTTGGATTGGGTACGGTTTCGTGTGCTTTTGTTGTTTTGGCCTCTAGAGTTGGATCTGCTTATCATTTGTCATTCCCTATATCATCTAGAGCAT CATTCGGTATTTTCT 10/27/2021 CSE 5290, Fall 2011 2

Annotated data ATATTGAATTTTCAAAAATTCTTACTTTTTGGACGCAAAGAAGTTTAATAATCATATTACATGGCATTACCACCATATCCATATCTAAT CTTATATGTTGTGGAAATGTAAAGAGCCCCATTATCTTAGCCTAAAAAAACCTTCTCTTTGGAACTTTCTAATACGCTTAACTGCTCATTGCTAT ATTGAAGTACGGATTAGAAGCCGCCGAGCGGGCGACAGCCCTCCGACGGAAGACTCTCCTCTGCGTCCTCGTCTTCACCGGTCGCGTTCCTGAAA CGCAGATGTGCCTCGCGCCGCACTGCTCCGAACAATAAAGATTCTACAATACTCTTTTATGGTTATGAAGAGGAAAAATTGGCAGTAACCTGGCCC CACAAACCTTCAAATTAACGAATCAAATTAACAACCATAGGATGAATGCGATTAGTTTTTTAGCCTTATTTCTGGGGTAATCAGCGATG ATTTTTGATCTATTAACAGATATATAAATGGAAGCTGCATAACCACTTTAACTAATACTTTCAACATTTTCAGTTTGTATTACTTCTTATTCAAATGTCAT AAAAGTATCAACAAAAAATTTAATATACCTCTATACTTTAACGTCAAGGAGAAAAAACTATAATGACTAAATCTCATTCAGAAGAAGTGATTGTACCTG AGTTCACTAGCGCAAAGGAATTACCAAGACCATTGGCCGAAAAGTGCCGAGCATAATTAAGAAATTTATAAGCGCTTATGATGCTAAACCGAGTTGT ATGTATTTGGCCTTATGTAGCTCGCGCCCGTTCGAGATAAGGATGTTTCTAGAAATCCGTAAAGATATAGAGATGTACACACATCTACATTTGTAACT CTATTTATAGTTAGAAACTTGTCCTCGAGGTCTCTCTATAAACCTTTTTGTACGTCCATAAATGTGGAAATCTACCGATCTTTTTGTCTCCGTATATAG GGAAACAGCGTTTTCGCTATACCTGGGTACAAACAGAGTTTTGTAGCTCCACGTTTCGCTGTCTCGTTCCGTGGAGCCCTGGGGGTCCTTAGACAT ATACTCTTTTTACATAGTTGGATGGGGGTCTGTACCTAGTTAGCTCGAGACAGCGATAGAGAATTTTGTATACTTGTCCGTTTACGTGTACC CGGCGATCTGTCTATGGATAGCCACGTTTATGTCGTTTTGTAGGTCGTTGTATATCGATATATAGAGCGCGGATAATTAGGTCGACC GCGCTGTGGCTCTATCTCTAGTTATTTGTAGGTCGATGTGTAGATGTAATTCTAGCTGGACATCCATACCTGTGTTTGTAGGTATTTCCATAAA ACCACAGCGATGTTTGTAGAAAACGCGCGCTACCCCTACACCGCTATATACATAATATATCTCTGTACAAAGATGTATAGAGATAAAGACACAGTTC GAAACCTATCGACTTGGACAAACAGTTGTTTATTTTTAAGTCGCTCGACCGAACTAGTTACACCGAGATCGATTTGTTTCTCTATACACCTCTG TGGAGAAACAGAGCGAGAAGTAGATTTCGAGAAGCCACCGGGACAATTACAGAAAGCGGTAGATTTACAAAGAAGGAGACTTATCGATACAC ATAGAGGTATCGATAACGATGTATACCTACATCCAGCTCCATACCTAAAGGTAGAAAGACATGTGTCGACATGTTTACGTTTAGATATGGACCTAGAT ATGTCTACGGACACGAGACTTACACGTCTAGATAGTTGTGTATTTCTCTCTGGACAGATCTGTAAAGGTACGTCTACATGTCGATATCGGTCTGTCG GTAGATAAAAATCTATATTTAGACCGATAACTAGGTCTCGATGTCGTTCTCTAACGATGGACCTGTAGACCGAAAAAGAACTTTTTGTTTTCCACAAG TCTAGACTTTTTTGGGTCTAGACACTTGTGGAATTCGAGATAGGGCTCTCCCTCTGGCTCTATAGATCGACGGGTATCGCTATCGAGACGTGGGTC GAGATGGGTATCTCGCTATACATGGATTTCCAACCTTGTAGGTGTCTCTCGAAGGCGGTAGGGACACAAAATAGCTGTAGCTACAACTACGTATCG ATACATAAAGAGCTACAAATTGGGCAGCTGTCTATATGAATTATAAGTATACTTCTTTTACTTTGTTCAGAACAACTTCTCATTTTTTTCTACTCA TAACTAGCATCACAAAATACGCAATAATAACGAGTAGTAACACTTTTATAGTTCATACATGCTTCAACTACTTAATAAATGATTGTATGATAGTTTTCAA TGTAAGAGATTTCGATTATCCACAAACTTTAAAACACAGGGACAAAATTCTTGATATGCTTTCAACCGCTGCGTTTTGGACCTATTCTTGACATGATAT GACTACCATTTTGTTATTGTACGTGGGGCAGTTGACGTCTTATCATATGTCAAAGTCATTTGCGAAGCTTGGCAAGTTGCCAACTGACGAGATGCAG TAAAAAGAGATTGCCGTCTTGAAACTTTTTGTCCTTTTTCCGGGGACTCTACGAACCCTTTGTCCTACTGATTAATTTTGTACTGAATTTGGAC AATTCAGATTTTAGTAGACAAGCGCGAGGAGGAAAAGAAATGACAAAAATTCCGATGGACAAGAAGATAGGAAAAAAGCTTTCACCGATTTC CTAGACCGGAAAAAAGTCGTATGACATCAGAATGAAATTTTCAAGTTAGACAAGGACAAAATCAGGACAAATTGTAAAGATATAATAAACTATTTGAT TCAGCGCCAATTTGCCCTTTTCCATTAAATCTCTGTTCTCTCTTATATGATGATTAGGTATCATCTGTATAAAACTCCTTTCTTAATTTCA CTCTAAAGCATACCGCCAGGTACGTATACAGAAATACATGTATCTGTGGATATCCGTACATCGAGCCACATATCCCTTTAACTGGCGAAATAT ACTTATACCGAAAATTAGAGGGAACGCGGTATATGTACGACACAATGAAACTAGATTGCGTAATTTCTAGTGTAAACAAATATGGCTATCTAAA TGTCTCTAGGTACATCGAAAGTTACATATATTTAAATCGATAACTACGTAGATGGGTTTCTAGTTGTAGAGCGACAAATCTCGAAAGCTCTTTT TGGAGAGGTAGATATATAGTATATATCGCTGTCGAAGTATACAAATATCTACTTCGATAACCAACGGTATCGGTCTAGAAAAGTCTCGCCAGG TCCGTAAACAAAGAGGTACATAACGAGACCGGTGTTTCGGTATACACTTGTGGGTATCGAGACATGTTTGTGTGTAACTATATCCAAGG TCTTTGTGGACTTGTAGAGGTGTATCTGTCGATATTTACGTC 10/27/2021 CSE 5290, Fall 2011 3

What do we do with genomes? • Inter-species comparison • Intra-species comparison • Diseases from a genomic standpoint, drug/therapy design 10/27/2021 CSE 5290, Fall 2011 4

Importance of algorithms – Compare human vs. mouse (blocks of 1, 000 nucleotides) • 3, 000*3, 000 comparisons, each 1, 000*1, 000 operations (w/dynamic progr. ) • At 1 trillion operations per second, it would take 104 days – Search all regulatory motifs of length 20 in the human genome: • 426 years 10/27/2021 CSE 5290, Fall 2011 5

Clustering flow cytometry data • • 1 million vectors Each of length 25 (real numbers) Need quick output! Results should be biologically meaningful! 10/27/2021 CSE 5290, Fall 2011 6

R: introduction Why R? • Lots of available libraries (statistics, machine learning, …. . ) • Very good visualization capability • Free • Multiplatform • Easy to publish code • Biologists use it! 10/27/2021 CSE 5290, Fall 2011 7

R - contd • Grew out of a popular statistics package • Used extensively by statisticians and computational biologists • Lots of resources (see class web page) • Some similarities with Mat. Lab 10/27/2021 CSE 5290, Fall 2011 8

R – strengths and weaknesses Strengths • Allows very quick testing of ideas • Libraries available for most purposes • Allows integration with C code Weaknesses • Not as efficient as Mat. Lab on matrix operations • Not very good at handling large data sets 10/27/2021 CSE 5290, Fall 2011 9

Next: • Ch 3 of text: Some techniques that biologists use for data gathering Things to do… • Read Ch 1 and 2 on your own. • Get familiar with R 10/27/2021 CSE 5290, Fall 2011 10

Analyzing a Genome • How to analyze a genome in four easy steps. – Cut it • Use enzymes to cut the DNA in to small fragments. – Copy it • Copy it many times to make it easier to see and detect. – Read it • Use special chemical techniques to read the small fragments. – Assemble it • Take all the fragments and put them back together. This is hard!!! • Bioinformatics takes over – What can we learn from the sequenced DNA. – Compare interspecies and intraspecies. 10/27/2021 CSE 5290, Fall 2011 11

8. 1 Copying DNA 10/27/2021 CSE 5290, Fall 2011 12

Why we need so many copies • Biologists needed to find a way to read DNA codes. • How do you read base pairs that are angstroms in size? – It is not possible to directly look at it due to DNA’s small size. – Need to use chemical techniques to detect what you are looking for. – To read something so small, you need a lot of it, so that you can actually detect the chemistry. • Need a way to make many copies of the base pairs, and a method for reading the pairs. 10/27/2021 CSE 5290, Fall 2011 13

Polymerase Chain Reaction (PCR) • Polymerase Chain Reaction (PCR) – Used to massively replicate DNA sequences. • How it works: – Separate the two strands with low heat – Add some base pairs, primer sequences, and DNA Polymerase • Creates double stranded DNA from a single strand. • Primer sequences create a seed from which double stranded DNA grows. – Now you have two copies. – Repeat. Amount of DNA grows exponentially. • 1→ 2→ 4→ 8→ 16→ 32→ 64→ 128→ 256… 10/27/2021 CSE 5290, Fall 2011 14
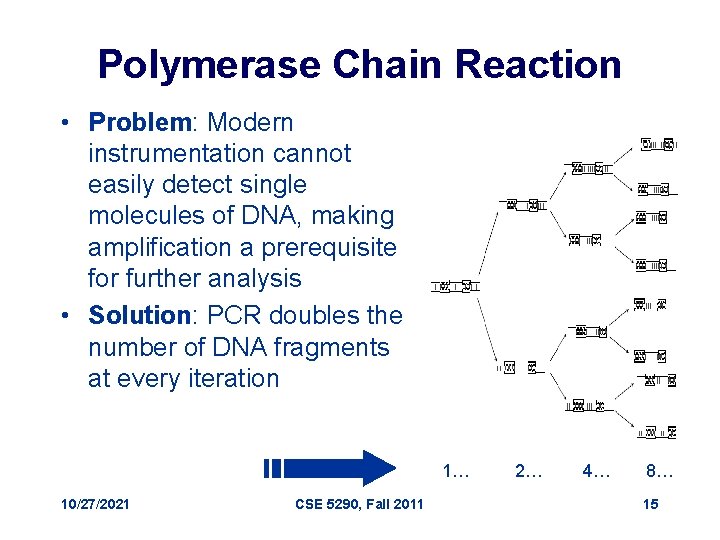
Polymerase Chain Reaction • Problem: Modern instrumentation cannot easily detect single molecules of DNA, making amplification a prerequisite for further analysis • Solution: PCR doubles the number of DNA fragments at every iteration 1… 10/27/2021 CSE 5290, Fall 2011 2… 4… 8… 15
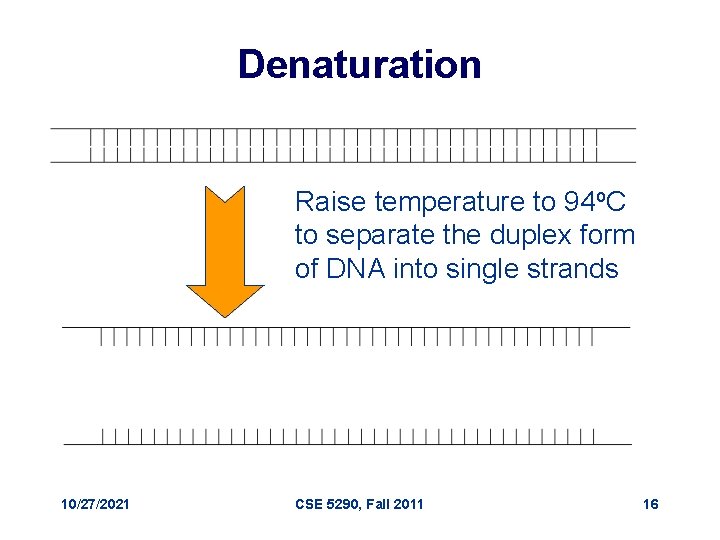
Denaturation Raise temperature to 94 o. C to separate the duplex form of DNA into single strands 10/27/2021 CSE 5290, Fall 2011 16

Design primers • To perform PCR, a 10 -20 bp sequence on either side of the sequence to be amplified must be known because DNA polymerase requires a primer to synthesize a new strand of DNA 10/27/2021 CSE 5290, Fall 2011 17
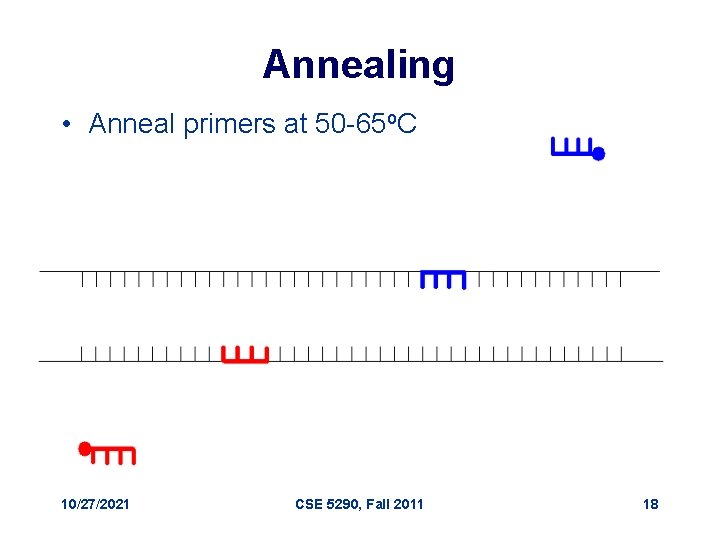
Annealing • Anneal primers at 50 -65 o. C 10/27/2021 CSE 5290, Fall 2011 18
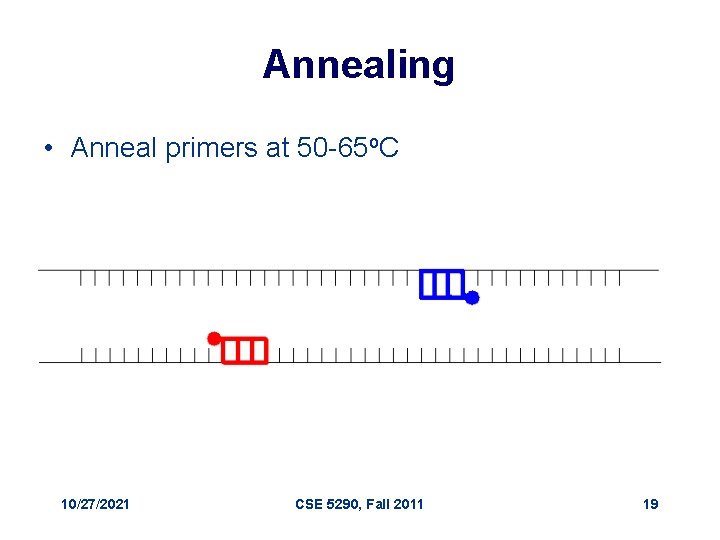
Annealing • Anneal primers at 50 -65 o. C 10/27/2021 CSE 5290, Fall 2011 19
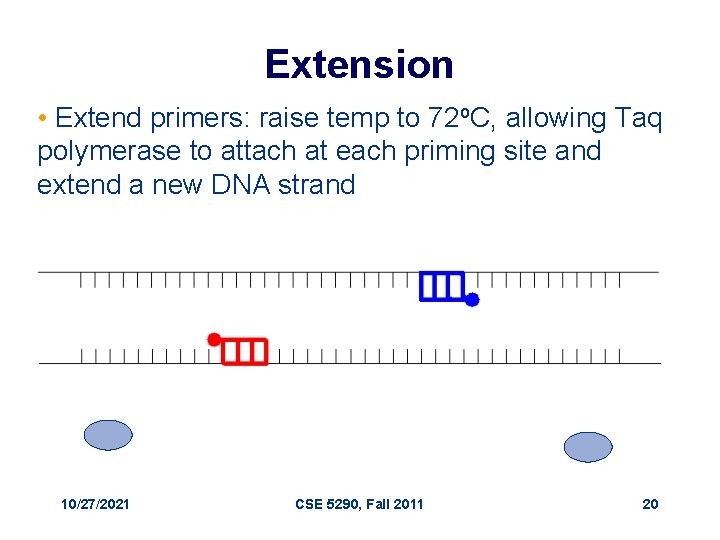
Extension • Extend primers: raise temp to 72 o. C, allowing Taq polymerase to attach at each priming site and extend a new DNA strand 10/27/2021 CSE 5290, Fall 2011 20
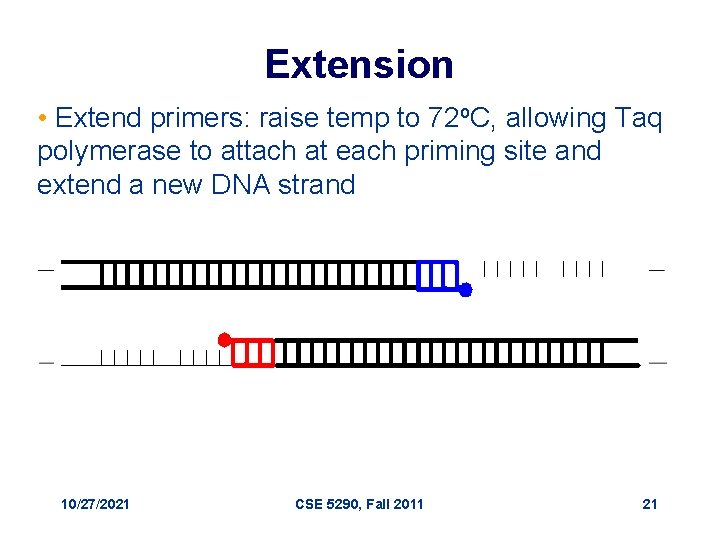
Extension • Extend primers: raise temp to 72 o. C, allowing Taq polymerase to attach at each priming site and extend a new DNA strand 10/27/2021 CSE 5290, Fall 2011 21
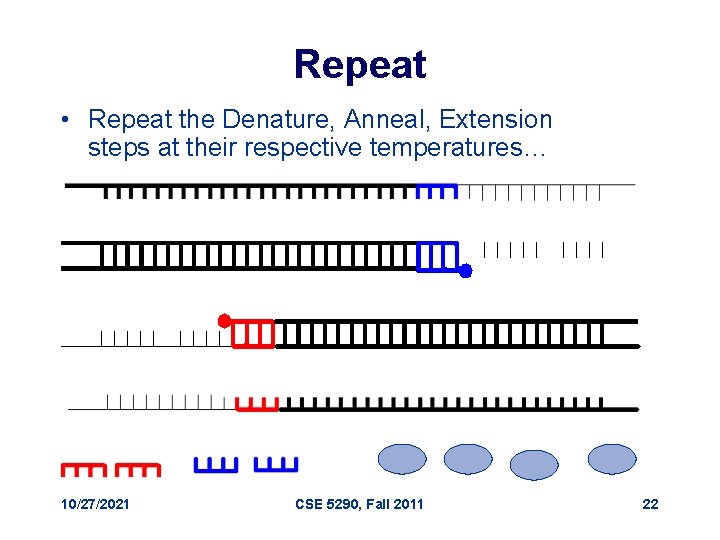
Repeat • Repeat the Denature, Anneal, Extension steps at their respective temperatures… 10/27/2021 CSE 5290, Fall 2011 22
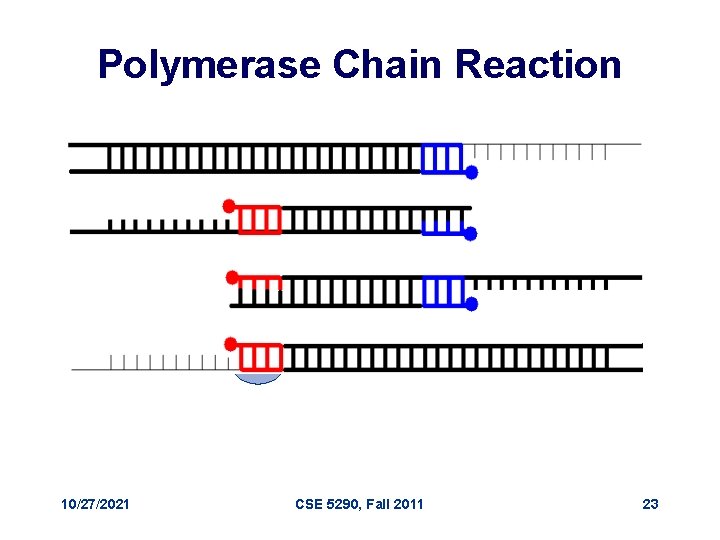
Polymerase Chain Reaction 10/27/2021 CSE 5290, Fall 2011 23
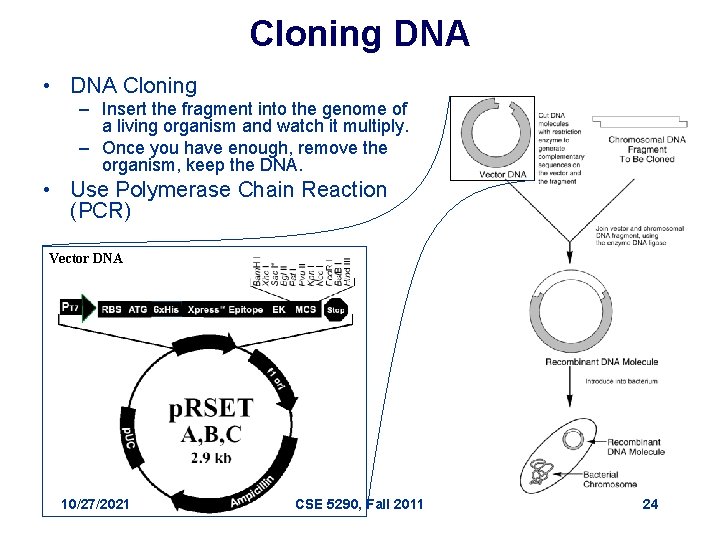
Cloning DNA • DNA Cloning – Insert the fragment into the genome of a living organism and watch it multiply. – Once you have enough, remove the organism, keep the DNA. • Use Polymerase Chain Reaction (PCR) Vector DNA 10/27/2021 CSE 5290, Fall 2011 24

8. 2 Cutting and Pasting DNA 10/27/2021 CSE 5290, Fall 2011 25

Restriction Enzymes • Discovered in the early 1970’s – Used as a defense mechanism by bacteria to break down the DNA of attacking viruses. – They cut the DNA into small fragments. • Can also be used to cut the DNA of organisms. – This allows the DNA sequence to be in a more manageable bite-size pieces. • It is then possible using standard purification techniques to single out certain fragments and duplicate them to macroscopic quantities. 10/27/2021 CSE 5290, Fall 2011 26
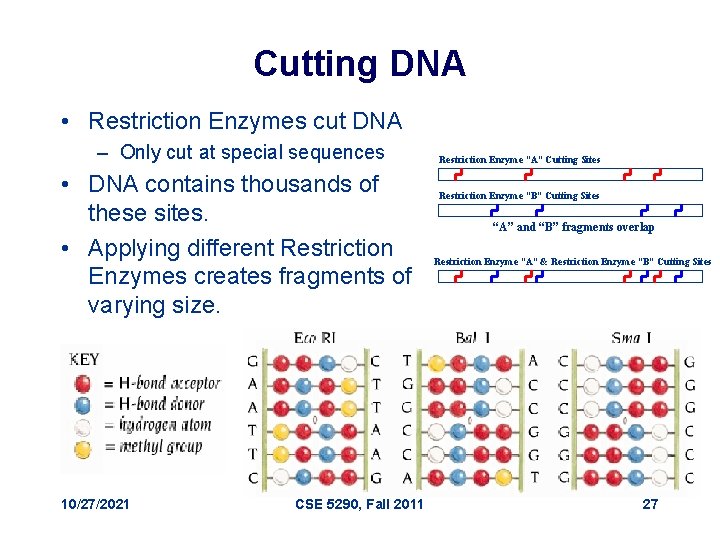
Cutting DNA • Restriction Enzymes cut DNA – Only cut at special sequences • DNA contains thousands of these sites. • Applying different Restriction Enzymes creates fragments of varying size. 10/27/2021 CSE 5290, Fall 2011 Restriction Enzyme “A” Cutting Sites Restriction Enzyme “B” Cutting Sites “A” and “B” fragments overlap Restriction Enzyme “A” & Restriction Enzyme “B” Cutting Sites 27
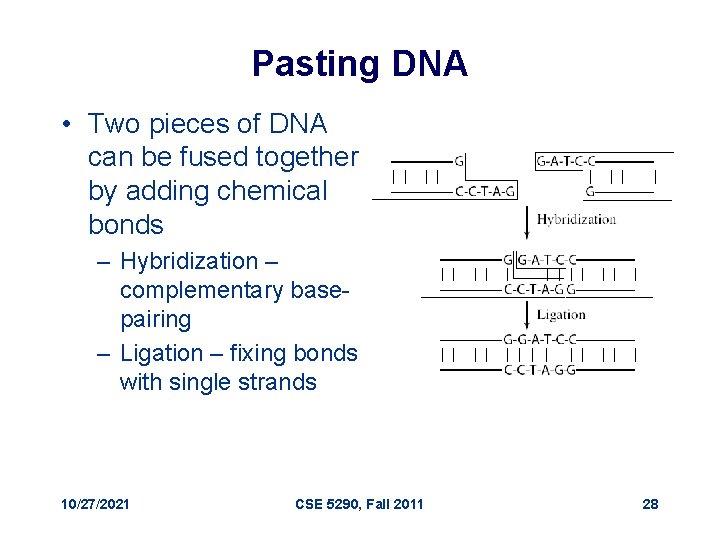
Pasting DNA • Two pieces of DNA can be fused together by adding chemical bonds – Hybridization – complementary basepairing – Ligation – fixing bonds with single strands 10/27/2021 CSE 5290, Fall 2011 28

8. 3 Measuring DNA Length 10/27/2021 CSE 5290, Fall 2011 29
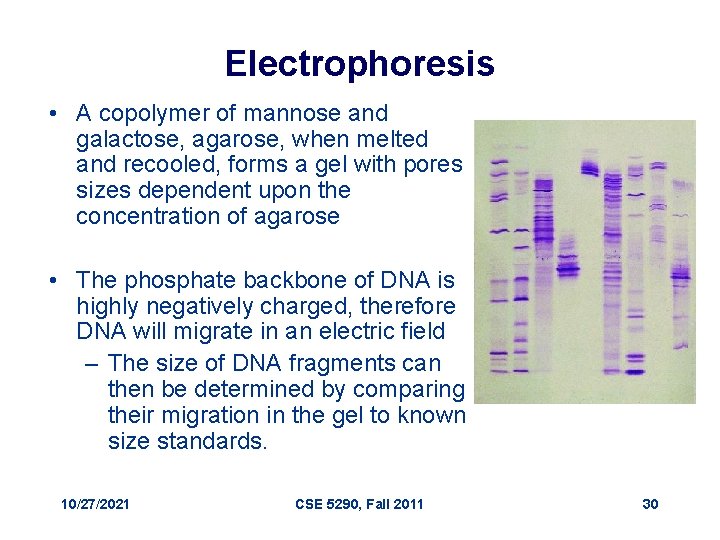
Electrophoresis • A copolymer of mannose and galactose, agarose, when melted and recooled, forms a gel with pores sizes dependent upon the concentration of agarose • The phosphate backbone of DNA is highly negatively charged, therefore DNA will migrate in an electric field – The size of DNA fragments can then be determined by comparing their migration in the gel to known size standards. 10/27/2021 CSE 5290, Fall 2011 30

Reading DNA • • Electrophoresis – Reading is done mostly by using this technique. This is based on separation of molecules by their size (and in 2 D gel by size and charge). – DNA or RNA molecules are charged in aqueous solution and move to a definite direction by the action of an electric field. – The DNA molecules are either labeled with radioisotopes or tagged with fluorescent dyes. In the latter, a laser beam can trace the dyes and send information to a computer. – Given a DNA molecule it is then possible to obtain all fragments from it that end in either A, or T, or G, or C and these can be sorted in a gel experiment. Another route to sequencing is direct sequencing using gene chips. 10/27/2021 CSE 5290, Fall 2011 31
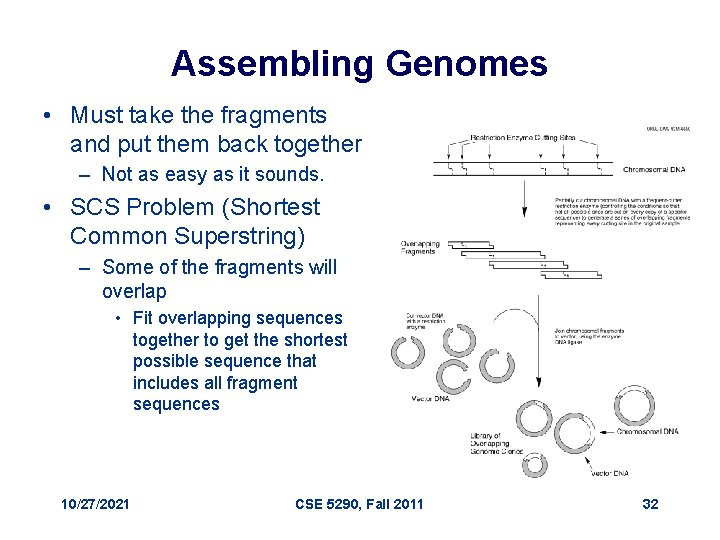
Assembling Genomes • Must take the fragments and put them back together – Not as easy as it sounds. • SCS Problem (Shortest Common Superstring) – Some of the fragments will overlap • Fit overlapping sequences together to get the shortest possible sequence that includes all fragment sequences 10/27/2021 CSE 5290, Fall 2011 32

Assembling Genomes • DNA fragments contain sequencing errors • Two complements of DNA – Need to take into account both directions of DNA • Repeat problem – 50% of human DNA is just repeats – If you have repeating DNA, how do you know where it goes? 10/27/2021 CSE 5290, Fall 2011 33

8. 4 Probing DNA 10/27/2021 CSE 5290, Fall 2011 34 34

DNA probes • Oligonucleotides: single-stranded DNA 20 -30 nucleotides long • Oligonucleotides used to find complementary DNA segments. • Made by working backwards---AA sequence----m. RNA--c. DNA. • Made with automated DNA synthesizers and tagged with a radioactive isotope. 10/27/2021 CSE 5290, Fall 2011 35 35

DNA Hybridization • Single-stranded DNA will naturally bind to complementary strands. • Hybridization is used to locate genes, regulate gene expression, and determine the degree of similarity between DNA from different sources. • Hybridization is also referred to as annealing or renaturation. 10/27/2021 CSE 5290, Fall 2011 36 36

Create a Hybridization Reaction 1. 2. Hybridization is binding two genetic sequences. The binding occurs because of the hydrogen bonds [pink] between base pairs. When using hybridization, DNA must first be denatured, usually by using use heat or chemical. 10/27/2021 CSE 5290, Fall 2011 T C A G T TAGGC T C TA ATCCGACAATGACGCC 37
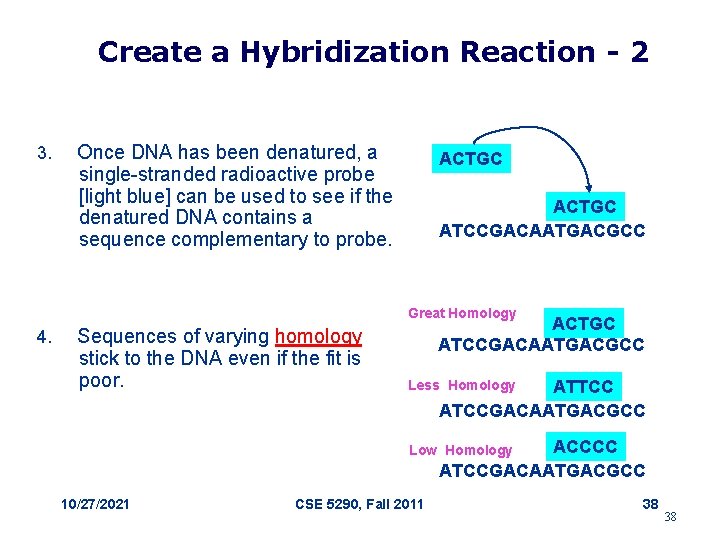
Create a Hybridization Reaction - 2 3. Once DNA has been denatured, a single-stranded radioactive probe [light blue] can be used to see if the denatured DNA contains a sequence complementary to probe. ACTGC ATCCGACAATGACGCC Great Homology 4. Sequences of varying homology stick to the DNA even if the fit is poor. ACTGC ATCCGACAATGACGCC ATTCC ATCCGACAATGACGCC Less Homology ACCCC ATCCGACAATGACGCC Low Homology 10/27/2021 CSE 5290, Fall 2011 38 38
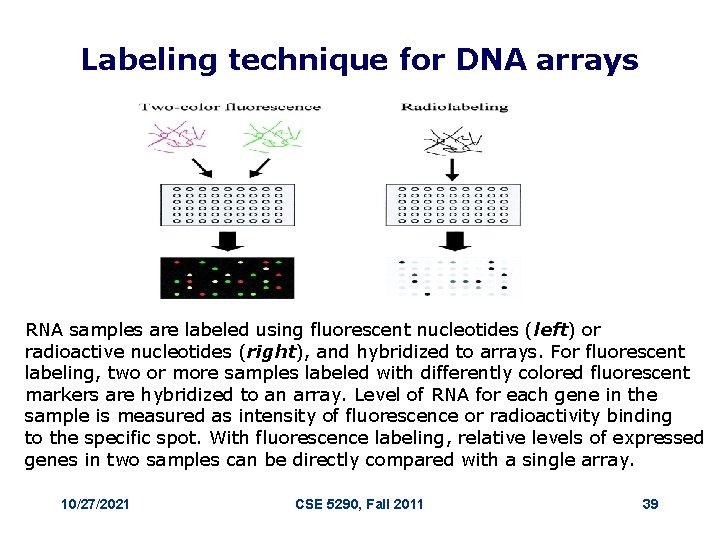
Labeling technique for DNA arrays RNA samples are labeled using fluorescent nucleotides (left) or radioactive nucleotides (right), and hybridized to arrays. For fluorescent labeling, two or more samples labeled with differently colored fluorescent markers are hybridized to an array. Level of RNA for each gene in the sample is measured as intensity of fluorescence or radioactivity binding to the specific spot. With fluorescence labeling, relative levels of expressed genes in two samples can be directly compared with a single array. 10/27/2021 CSE 5290, Fall 2011 39

DNA Arrays--Technical Foundations • An array works by exploiting the ability of a given m. RNA molecule to hybridize to the DNA template. • Using an array containing many DNA samples in an experiment, the expression levels of hundreds or thousands genes within a cell by measuring the amount of m. RNA bound to each site on the array. • With the aid of a computer, the amount of m. RNA bound to the spots on the microarray is precisely measured, generating a profile of gene expression in the cell. 10/27/2021 CSE 5290, Fall 2011 40

An experiment on a microarray In this schematic: GREEN represents Control DNA RED represents Sample DNA YELLOW represents a combination of Control and Sample DNA BLACK represents areas where neither the Control nor Sample DNA Each color in an array represents either healthy (control) or diseased (sample) tissue. The location and intensity of a color tell us whether the gene, or mutation, is present in the control and/or sample DNA. 10/27/2021 CSE 5290, Fall 2011 41
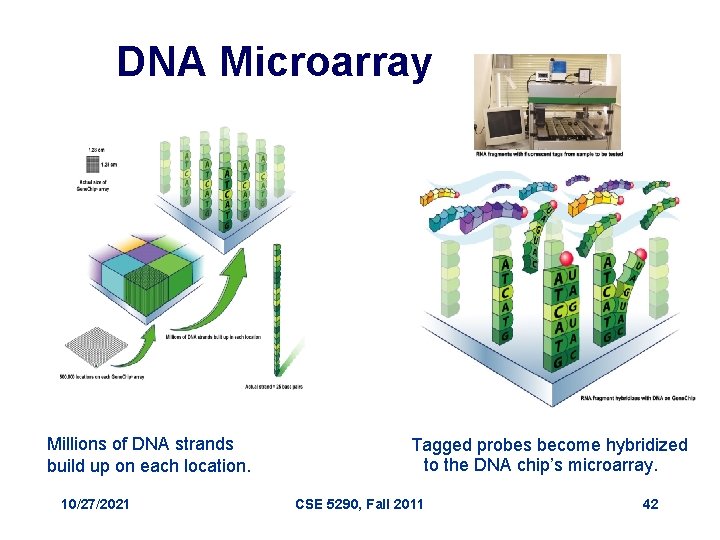
DNA Microarray Millions of DNA strands build up on each location. 10/27/2021 Tagged probes become hybridized to the DNA chip’s microarray. CSE 5290, Fall 2011 42
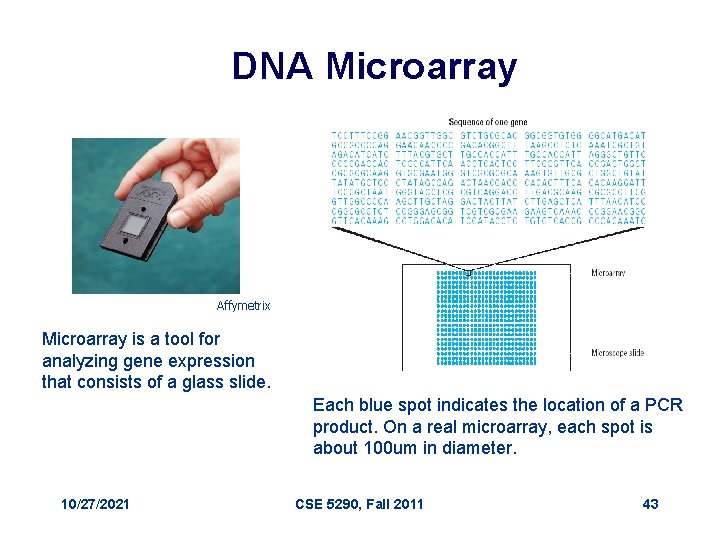
DNA Microarray Affymetrix Microarray is a tool for analyzing gene expression that consists of a glass slide. Each blue spot indicates the location of a PCR product. On a real microarray, each spot is about 100 um in diameter. 10/27/2021 CSE 5290, Fall 2011 43

Photolithography Light directed oligonucleotide synthesis. • A solid support is derivatized with a covalent linker molecule terminated with a photolabile protecting group. • Light is directed through a mask to deprotect and activate selected sites, and protected nucleotides couple to the activated sites. • The process is repeated, activating different set of sites and coupling different based allowing arbitrary DNA probes to be constructed at each site. • 10/27/2021 CSE 5290, Fall 2011 44
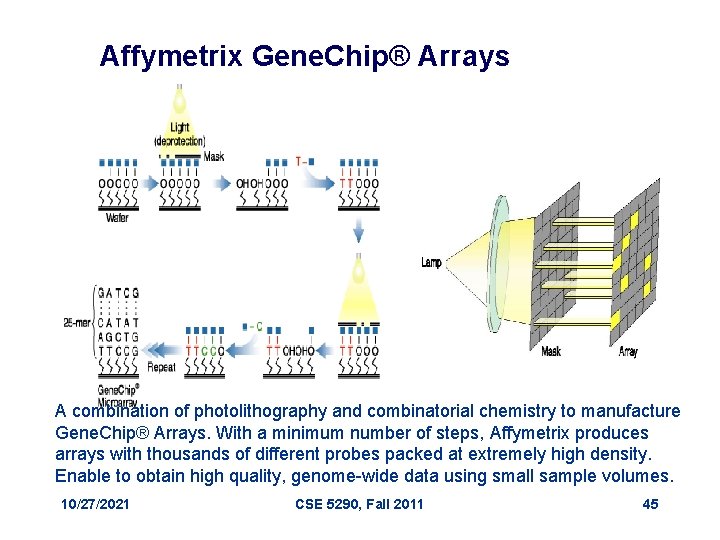
Affymetrix Gene. Chip® Arrays A combination of photolithography and combinatorial chemistry to manufacture Gene. Chip® Arrays. With a minimum number of steps, Affymetrix produces arrays with thousands of different probes packed at extremely high density. Enable to obtain high quality, genome-wide data using small sample volumes. 10/27/2021 CSE 5290, Fall 2011 45

Affymetrix Gene. Chip® Arrays Data from an experiment showing the expression of thousands of genes on a single Gene. Chip® probe array. 10/27/2021 CSE 5290, Fall 2011 46

Next • DNA Mapping and Brute Force Algorithms • How is DNA sequenced? 10/27/2021 CSE 5290, Fall 2011 47

Full Restriction Digest • Cutting DNA at each restriction site creates multiple restriction fragments: • Is it possible to reconstruct the order of the fragments from the sizes of the fragments {3, 5, 5, 9} ? 10/27/2021 CSE 5290, Fall 2011 48

Full Restriction Digest: Multiple Solutions • Alternative ordering of restriction fragments: vs 10/27/2021 CSE 5290, Fall 2011 49

Measuring Length of Restriction Fragments • Restriction enzymes break DNA into restriction fragments. • Gel electrophoresis is a process for separating DNA by size and measuring sizes of restriction fragments • Can separate DNA fragments that differ in length in only 1 nucleotide for fragments up to 500 nucleotides long 10/27/2021 CSE 5290, Fall 2011 50

Partial Restriction Digest • The sample of DNA is exposed to the restriction enzyme for only a limited amount of time to prevent it from being cut at all restriction sites • This experiment generates the set of all possible restriction fragments between every 2 (not necessarily consecutive) cuts • This set of fragment sizes is used to determine the positions of the restriction sites in the DNA sequence 10/27/2021 CSE 5290, Fall 2011 51

Partial Digest Example • Partial Digest results in the following 10 restriction fragments: 10/27/2021 CSE 5290, Fall 2011 52
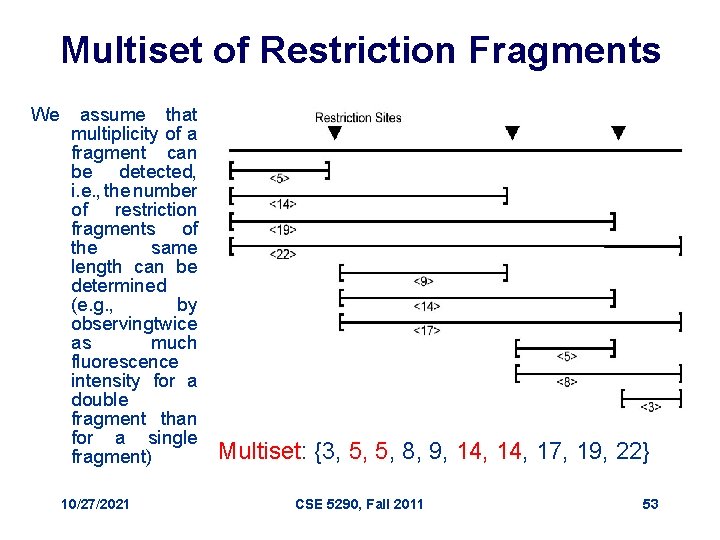
Multiset of Restriction Fragments We assume that multiplicity of a fragment can be detected, i. e. , the number of restriction fragments of the same length can be determined (e. g. , by observing twice as much fluorescence intensity for a double fragment than for a single fragment) 10/27/2021 Multiset: {3, 5, 5, 8, 9, 14, 17, 19, 22} CSE 5290, Fall 2011 53

Partial Digest Fundamentals X: the set of n integers representing the location of all cuts in the restriction map, including the start and end n: the total number of cuts DX: the multiset of integers representing lengths of each of the n. C 2 fragments produced from a partial digest 10/27/2021 CSE 5290, Fall 2011 54

One More Partial Digest Example X 0 2 4 0 2 2 4 4 2 7 7 5 3 7 10 10 10 8 6 3 Representation of DX = {2, 2, 3, 3, 4, 5, 6, 7, 8, 10} as a two dimensional table, with elements of X = {0, 2, 4, 7, 10} along both the top and left side. The elements at (i, j) in the table is xj – xi for 1 ≤ i < j ≤ n. 10/27/2021 CSE 5290, Fall 2011 55

Partial Digest Problem: Formulation Goal: Given all pairwise distances between points on a line, reconstruct the positions of those points • Input: The multiset of pairwise distances L, containing n(n-1)/2 integers • Output: A set X, of n integers, such that DX = L 10/27/2021 CSE 5290, Fall 2011 56

Partial Digest: Multiple Solutions • It is not always possible to uniquely reconstruct a set X based only on DX. • For example, the set X = {0, 2, 5} and (X + 10) = {10, 12, 15} both produce DX={2, 3, 5} as their partial digest set. • The sets {0, 1, 2, 5, 7, 9, 12} and {0, 1, 5, 7, 8, 10, 12} present a less trivial example of non-uniqueness. They both digest into: {1, 1, 2, 2, 2, 3, 3, 4, 4, 5, 5, 5, 6, 7, 7, 7, 8, 9, 10, 11, 12} 10/27/2021 CSE 5290, Fall 2011 57
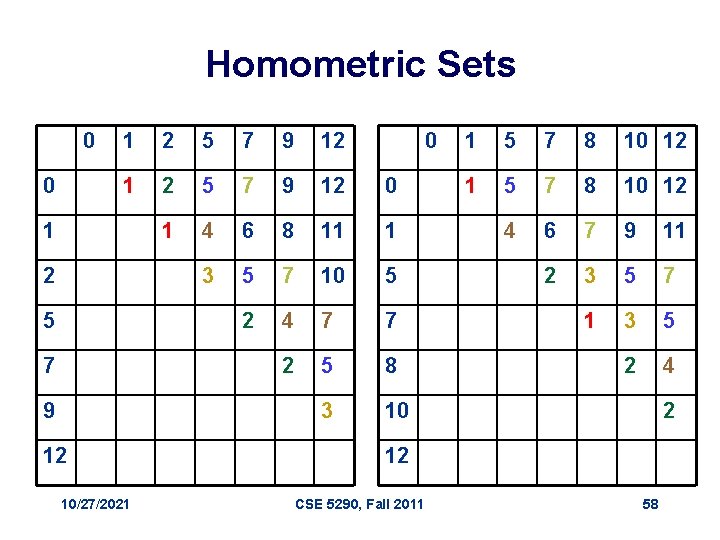
Homometric Sets 0 0 1 2 5 7 9 12 0 1 4 6 8 11 1 3 5 7 10 5 2 4 7 7 2 5 8 3 10 1 2 5 7 9 12 10/27/2021 0 1 5 7 8 10 12 4 6 7 9 11 2 3 5 7 1 3 5 2 4 2 12 CSE 5290, Fall 2011 58

Brute Force Algorithms • Also known as exhaustive search algorithms; examine every possible variant to find a solution • Efficient in rare cases; usually impractical 10/27/2021 CSE 5290, Fall 2011 59

Partial Digest: Brute Force 1. Find the restriction fragment of maximum length M. M is the length of the DNA sequence. 2. For every possible set X={0, x 2, … , xn-1, M} compute the corresponding DX 5. If DX is equal to the experimental partial digest L, then X is the correct restriction map 10/27/2021 CSE 5290, Fall 2011 60

Brute. Force. PDP 1. Brute. Force. PDP(L, n): 2. M <- maximum element in L 3. for every set of n – 2 integers 0 < x 2 < … xn-1 < M 4. X <- {0, x 2, …, xn-1, M} 5. Form DX from X 6. if DX = L 7. return X 8. output “no solution” 10/27/2021 CSE 5290, Fall 2011 61

Efficiency of Brute. Force. PDP • Brute. Force. PDP takes O(M n-2) time since it must examine all possible sets of positions. • One way to improve the algorithm is to limit the values of xi to only those values which occur in L. 10/27/2021 CSE 5290, Fall 2011 62

Another. Brute. Force. PDP 1. 2. 3. 4. 5. 6. 7. 8. Another. Brute. Force. PDP(L, n) M <- maximum element in L for every set of n – 2 integers 0 < x 2 < … xn-1 < M X <- { 0, x 2, …, xn-1, M } Form DX from X if DX = L return X output “no solution” 10/27/2021 CSE 5290, Fall 2011 63

Another. Brute. Force. PDP 1. 2. 3. 4. 5. 6. 7. 8. Another. Brute. Force. PDP(L, n) M <- maximum element in L for every set of n – 2 integers 0 < x 2 < … xn-1 < M from L X <- { 0, x 2, …, xn-1, M } Form DX from X if DX = L return X output “no solution” 10/27/2021 CSE 5290, Fall 2011 64

Efficiency of Another. Brute. Force. PDP • It’s more efficient, but still slow • If L = {2, 998, 1000} (n = 3, M = 1000), Brute. Force. PDP will be extremely slow, but Another. Brute. Force. PDP will be quite fast • Fewer sets are examined, but runtime is still exponential: O(n 2 n-4) 10/27/2021 CSE 5290, Fall 2011 65

Defining D(y, X) • Before describing Partial. Digest, first define D(y, X) as the multiset of all distances between point y and all other points in the set X D(y, X) = {|y – x 1|, |y – x 2|, …, |y – xn|} for X = {x 1, x 2, …, xn} 10/27/2021 CSE 5290, Fall 2011 66

Partial. Digest Algorithm Partial. Digest(L): width <- Maximum element in L DELETE(width, L) X <- {0, width} PLACE(L, X) 10/27/2021 CSE 5290, Fall 2011 67

Partial. Digest Algorithm (cont’d) 1. PLACE(L, X) 2. if L is empty 3. output X 4. return 5. y <- maximum element in L 6. Delete(y, L) 7. if D(y, X ) Í L 8. Add y to X and remove lengths D(y, X) from L 9. PLACE(L, X ) 10. Remove y from X and add lengths D(y, X) to L 11. if D(width-y, X ) Í L 12. Add width-y to X and remove lengths D(width-y, X) from L 13. PLACE(L, X ) 14. Remove width-y from X and add lengths D(width-y, X ) to L 15. return 10/27/2021 CSE 5290, Fall 2011 68

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X={0} 10/27/2021 CSE 5290, Fall 2011 69

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X={0} Remove 10 from L and insert it into X. We know this must be the length of the DNA sequence because it is the largest fragment. 10/27/2021 CSE 5290, Fall 2011 70

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 10 } 10/27/2021 CSE 5290, Fall 2011 71

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 10 } Take 8 from L and make y = 2 or 8. But since the two cases are symmetric, we can assume y = 2. 10/27/2021 CSE 5290, Fall 2011 72

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 10 } We find that the distances from y=2 to other elements in X are D(y, X) = {8, 2}, so we remove {8, 2} from L and add 2 to X. 10/27/2021 CSE 5290, Fall 2011 73

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 2, 10 } 10/27/2021 CSE 5290, Fall 2011 74

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 2, 10 } Take 7 from L and make y = 7 or y = 10 – 7 = 3. We will explore y = 7 first, so D(y, X ) = {7, 5, 3}. 10/27/2021 CSE 5290, Fall 2011 75

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 2, 10 } For y = 7 first, D(y, X ) = {7, 5, 3}. Therefore we remove {7, 5 , 3} from L and add 7 to X. D(y, X) = {7, 5, 3} = {½ 7 – 0½, ½ 7 – 2½, ½ 7 – 10½} 10/27/2021 CSE 5290, Fall 2011 76

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 2, 7, 10 } 10/27/2021 CSE 5290, Fall 2011 77

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 2, 7, 10 } Take 6 from L and make y = 6. Unfortunately D(y, X) = {6, 4, 1 , 4}, which is not a subset of L. Therefore we won’t explore this branch. 6 10/27/2021 CSE 5290, Fall 2011 78

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 2, 7, 10 } This time make y = 4. D(y, X) = {4, 2, 3 , 6}, which is a subset of L so we will explore this branch. We remove {4, 2, 3 , 6} from L and add 4 to X. 10/27/2021 CSE 5290, Fall 2011 79

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 2, 4, 7, 10 } 10/27/2021 CSE 5290, Fall 2011 80

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 2, 4, 7, 10 } L is now empty, so we have a solution, which is X. 10/27/2021 CSE 5290, Fall 2011 81

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 2, 7, 10 } To find other solutions, we backtrack. 10/27/2021 CSE 5290, Fall 2011 82

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 2, 10 } More backtrack. 10/27/2021 CSE 5290, Fall 2011 83

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 2, 10 } This time we will explore y = 3. D(y, X) = {3, 1, 7}, which is not a subset of L, so we won’t explore this branch. 10/27/2021 CSE 5290, Fall 2011 84

An Example L = { 2, 2, 3, 3, 4, 5, 6, 7, 8, 10 } X = { 0, 10 } We backtracked back to the root. Therefore we have found all the solutions. 10/27/2021 CSE 5290, Fall 2011 85

Analyzing Partial. Digest Algorithm • Still exponential in worst case, but is very fast on average • Informally, let T(n) be time Partial. Digest takes to place n cuts – No branching case: T(n) < T(n-1) + O(n) • Quadratic – Branching case: T(n) < 2 T(n-1) + O(n) • Exponential 10/27/2021 CSE 5290, Fall 2011 86

Double Digest Mapping • Double Digest is yet another experimentally method to construct restriction maps – Use two restriction enzymes; three full digests: • One with only first enzyme • One with only second enzyme • One with both enzymes • Computationally, Double Digest problem is more complex than Partial Digest problem 10/27/2021 CSE 5290, Fall 2011 87

Next: Finding Regulatory Motifs in DNA sequences 10/27/2021 CSE 5290, Fall 2011 88

Combinatorial Gene Regulation • A microarray experiment showed that when gene X is knocked out, 20 other genes are not expressed – How can one gene have such drastic effects? 10/27/2021 CSE 5290, Fall 2011 89

Regulatory Proteins • Gene X encodes regulatory protein, a. k. a. a transcription factor (TF) • The 20 unexpressed genes rely on gene X’s TF to induce transcription • A single TF may regulate multiple genes 10/27/2021 CSE 5290, Fall 2011 90

Regulatory Regions • Every gene contains a regulatory region (RR) typically stretching 100 -1000 bp upstream of the transcriptional start site • Located within the RR are the Transcription Factor Binding Sites (TFBS), also known as motifs, specific for a given transcription factor • TFs influence gene expression by binding to a specific location in the respective gene’s regulatory region - TFBS 10/27/2021 CSE 5290, Fall 2011 91

Transcription Factor Binding Sites • A TFBS can be located anywhere within the Regulatory Region. • TFBS may vary slightly across different regulatory regions since non-essential bases could mutate 10/27/2021 CSE 5290, Fall 2011 92
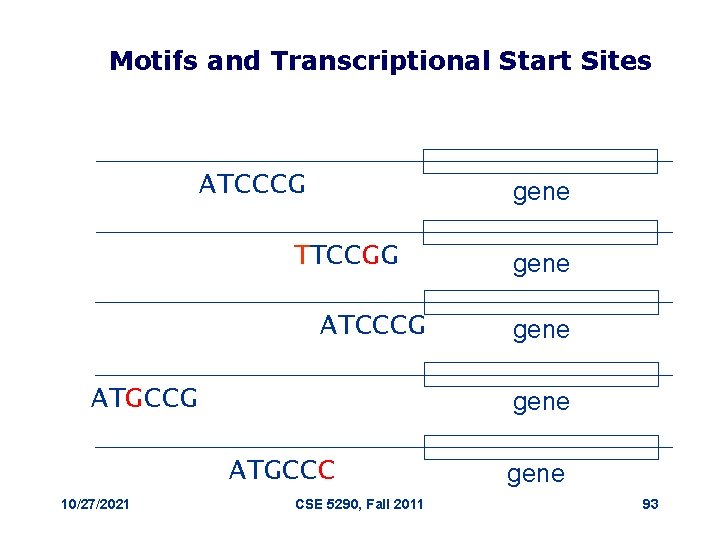
Motifs and Transcriptional Start Sites ATCCCG gene TTCCGG ATCCCG ATGCCG gene ATGCCC 10/27/2021 gene CSE 5290, Fall 2011 gene 93
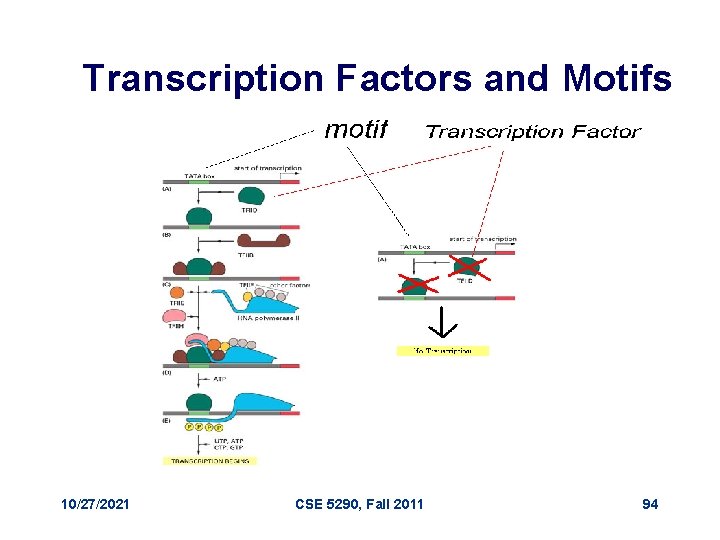
Transcription Factors and Motifs 10/27/2021 CSE 5290, Fall 2011 94

Motif Logo • Motifs can mutate on non important bases • The five motifs in five different genes have mutations in position 3 and 5 • Representations called motif logos illustrate the conserved and variable regions of a motif 10/27/2021 CSE 5290, Fall 2011 TGGGGGA TGAGAGA TGAGGGA 95

Identifying Motifs • Genes are turned on or off by regulatory proteins • These proteins bind to upstream regulatory regions of genes to either attract or block an RNA polymerase • Regulatory protein (TF) binds to a short DNA sequence called a motif (TFBS) • So finding the same motif in multiple genes’ regulatory regions suggests a regulatory relationship amongst those genes 10/27/2021 CSE 5290, Fall 2011 96

Identifying Motifs: Complications • We do not know the motif sequence • We do not know where it is located relative to the genes start • Motifs can differ slightly from one gene to the next • How to discern it from “random” motifs? 10/27/2021 CSE 5290, Fall 2011 97

Motifs: summary • Short strings • Transcription factors bind to motifs • Regulate downstream genes Challenge: must find patterns without knowing them 10/27/2021 CSE 5290, Fall 2011 98

How can we do this? • Statistics? • Brute force search 10/27/2021 CSE 5290, Fall 2011 99

Statistical approaches • Deciphering encoded English text. 10/27/2021 CSE 5290, Fall 2011 100

Motif Finding and The Gold Bug Problem: Similarities – Nucleotides in motifs encode for a message in the “genetic” language. Symbols in “The Gold Bug” encode for a message in English – In order to solve the problem, we analyze the frequencies of patterns in DNA/Gold Bug message. – Knowledge of established regulatory motifs makes the Motif Finding problem simpler. Knowledge of the words in the English dictionary helps to solve the Gold Bug problem. 10/27/2021 CSE 5290, Fall 2011 101

Similarities (cont’d) • Motif Finding: – In order to solve the problem, we analyze the frequencies of patterns in the nucleotide sequences • Gold Bug Problem: – In order to solve the problem, we analyze the frequencies of patterns in the text written in English 10/27/2021 CSE 5290, Fall 2011 102

Similarities (cont’d) • Motif Finding: – Knowledge of established motifs reduces the complexity of the problem • Gold Bug Problem: – Knowledge of the words in the dictionary is highly desirable 10/27/2021 CSE 5290, Fall 2011 103

Motif Finding and The Gold Bug Problem: Differences Motif Finding is harder than Gold Bug problem: – We don’t have the complete dictionary of motifs – The “genetic” language does not have a standard “grammar” – Only a small fraction of nucleotide sequences encode for motifs; the size of data is enormous 10/27/2021 CSE 5290, Fall 2011 104

Challenge Problem – Find a motif in a sample of - 20 “random” sequences (e. g. 600 nt long) - each sequence containing an implanted pattern of length 15, - each pattern appearing with 4 mismatches as (15, 4)-motif. 10/27/2021 CSE 5290, Fall 2011 105

Exhaustive searches 10/27/2021 CSE 5290, Fall 2011 106

The Motif Finding Problem • Given a random sample of DNA sequences: cctgatagacgctatctggctatccacgtaggtcctctgtgcgaatctatgcgtttccaaccat agtactggtgtacatttgatacgtacaccggcaacctgaaacgctcagaaccagaagtgc aaacgtgcaccctcttcgtggctctggccaacgagggctgatgtataagacgaaaatttt agcctccgatgtaagtcatagctgtaactattacctgccacccctattacatcttacgtataca ctgttatacaacgcgtcatggcggggtatgcgttttggtcgtcgtacgctcgatcgttaacgtc • Find the pattern that is implanted in each of the individual sequences, namely, the motif 10/27/2021 CSE 5290, Fall 2011 107

The Motif Finding Problem (cont’d) • Additional information: – The hidden sequence is of length 8 – The pattern is not exactly the same in each array because random point mutations may occur in the sequences 10/27/2021 CSE 5290, Fall 2011 108

The Motif Finding Problem (cont’d) • The patterns revealed with no mutations: cctgatagacgctatctggctatccacgtaggtcctctgtgcgaatctatgcgtttccaaccat agtactggtgtacatttgatacgtacaccggcaacctgaaacgctcagaaccagaagtgc aaacgtgcaccctcttcgtggctctggccaacgagggctgatgtataagacgaaaatttt agcctccgatgtaagtcatagctgtaactattacctgccacccctattacatctt acgtataca ctgttatacaacgcgtcatggcggggtatgcgttttggtcgtcgtacgctcgatcgtta acgtc acgt Consensus String 10/27/2021 CSE 5290, Fall 2011 109

The Motif Finding Problem (cont’d) • The patterns with 2 point mutations: cctgatagacgctatctggctatcca. Ggtac. Ttaggtcctctgtgcgaatctatgcgtttcc aaccat agtactggtgtacatttgat. Cc. Atacgtacaccggcaacctgaaacgctcagaaccag aagtgc aaacgt. TAgtgcaccctcttcgtggctctggccaacgagggctgatgtataagacgaa aatttt agcctccgatgtaagtcatagctgtaactattacctgccacccctattacatcttacgt. Cc. A tataca ctgttatacaacgcgtcatggcggggtatgcgttttggtcgtcgtacgctcgatcgtta. Ccg tacg. Gc 10/27/2021 CSE 5290, Fall 2011 110

The Motif Finding Problem (cont’d) • The patterns with 2 point mutations: cctgatagacgctatctggctatcca. Ggtac. Ttaggtcctctgtgcgaatctatgcgtttcc aaccat agtactggtgtacatttgat. Cc. Atacgtacaccggcaacctgaaacgctcagaaccag aagtgc aaacgt. TAgtgcaccctcttcgtggctctggccaacgagggctgatgtataagacgaa aatttt agcctccgatgtaagtcatagctgtaactattacctgccacccctattacatcttacgt. Cc. A tataca ctgttatacaacgcgtcatggcggggtatgcgttttggtcgtcgtacgctcgatcgtta. Ccg tacg. Gc Can we still find the motif, now that we have 2 mutations? 10/27/2021 CSE 5290, Fall 2011 111

Defining Motifs • To define a motif, lets say we know where the motif starts in the sequence • The motif start positions in their sequences can be represented as s = (s 1, s 2, s 3, …, st) 10/27/2021 CSE 5290, Fall 2011 112
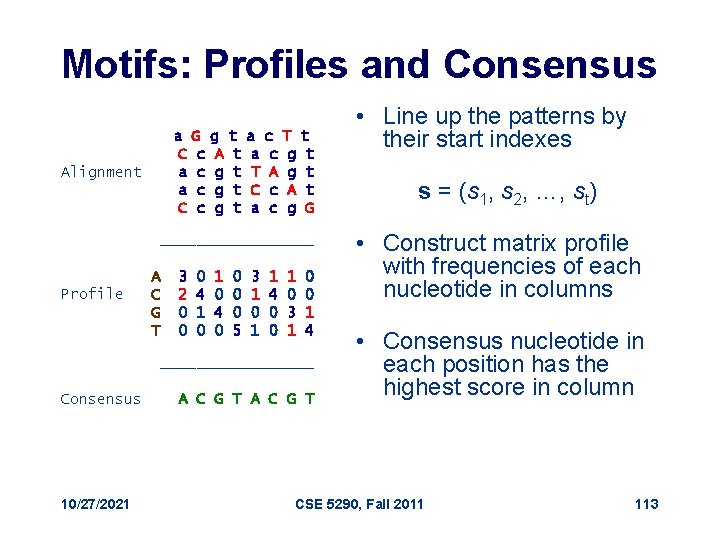
Motifs: Profiles and Consensus a G g t a c T t C c A t a c g t T A g t a c g t C c A t C c g t a c g G Alignment _________ Profile A C G T 3 2 0 0 0 4 1 0 4 0 0 5 3 1 0 1 1 4 0 0 1 0 3 1 0 0 1 4 _________ Consensus 10/27/2021 A C G T • Line up the patterns by their start indexes s = (s 1, s 2, …, st) • Construct matrix profile with frequencies of each nucleotide in columns • Consensus nucleotide in each position has the highest score in column CSE 5290, Fall 2011 113

Consensus • Think of consensus as an “ancestor” motif, from which mutated motifs emerged • The distance between a real motif and the consensus sequence is generally less than that for two real motifs 10/27/2021 CSE 5290, Fall 2011 114
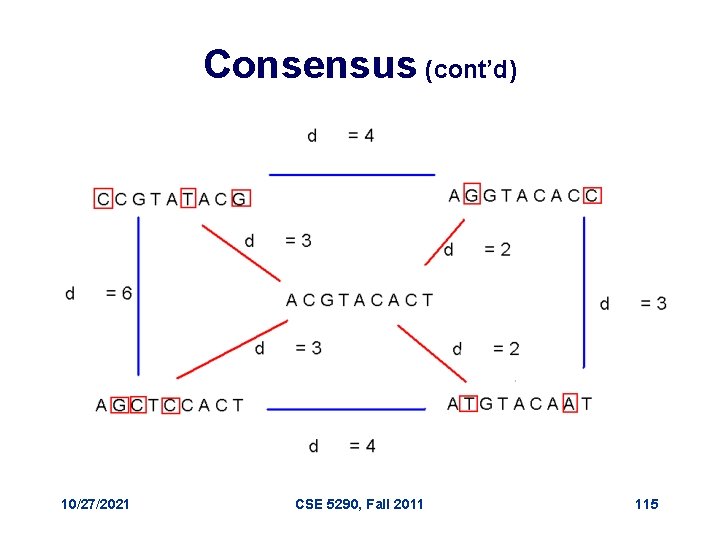
Consensus (cont’d) 10/27/2021 CSE 5290, Fall 2011 115

Evaluating Motifs • We have a guess about the consensus sequence, but how “good” is this consensus? • Need to introduce a scoring function to compare different guesses and choose the “best” one. 10/27/2021 CSE 5290, Fall 2011 116

Defining Some Terms • t - number of sample DNA sequences • n - length of each DNA sequence • DNA - sample of DNA sequences (t x n array) • l - length of the motif (l-mer) • si - starting position of an l-mer in sequence i • s=(s 1, s 2, … st) - array of motif’s starting positions 10/27/2021 CSE 5290, Fall 2011 117
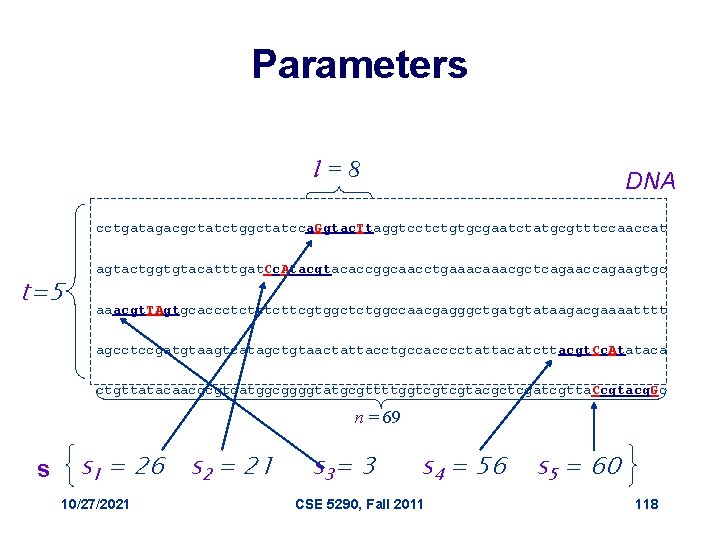
Parameters l=8 DNA cctgatagacgctatctggctatcca. Ggtac. Ttaggtcctctgtgcgaatctatgcgtttccaaccat t=5 agtactggtgtacatttgat. Cc. Atacgtacaccggcaacctgaaacgctcagaaccagaagtgc aaacgt. TAgtgcaccctcttcgtggctctggccaacgagggctgatgtataagacgaaaatttt agcctccgatgtaagtcatagctgtaactattacctgccacccctattacatcttacgt. Cc. Atataca ctgttatacaacgcgtcatggcggggtatgcgttttggtcgtcgtacgctcgatcgtta. Ccgtacg. Gc n = 69 s s 1 = 26 s 2 = 21 10/27/2021 s 3= 3 s 4 = 56 CSE 5290, Fall 2011 s 5 = 60 118

Scoring Motifs l • Given s = (s 1, … st) and DNA: Score(s, DNA) = a G g t a c T t C c A t a c g t T A g tt a c g t C c A t C c g t a c g G _________ A C G T Consensus 3 0 1 0 3 1 1 0 2 4 0 0 1 4 0 0 0 3 1 0 0 0 5 1 0 1 4 _________ a c g t Score 3+4+4+5+3+4=30 10/27/2021 CSE 5290, Fall 2011 119

The Motif Finding Problem • If starting positions s=(s 1, s 2, … st) are given, finding consensus is easy even with mutations in the sequences because we can simply construct the profile to find the motif (consensus) • But… the starting positions s are usually not given. How can we find the “best” profile matrix? 10/27/2021 CSE 5290, Fall 2011 120

The Motif Finding Problem: Formulation • Goal: Given a set of DNA sequences, find a set of l-mers, one from each sequence, that maximizes the consensus score • Input: A t x n matrix of DNA, and l, the length of the pattern to find • Output: An array of t starting positions s = (s 1, s 2, … st) maximizing Score(s, DNA) 10/27/2021 CSE 5290, Fall 2011 121

The Motif Finding Problem: Brute Force Solution – Compute the scores for each possible combination of starting positions s – The best score will determine the best profile and the consensus pattern in DNA – The goal is to maximize Score(s, DNA) by varying the starting positions si, where: si = [1, …, n-l+1] i = [1, …, t] 10/27/2021 CSE 5290, Fall 2011 122

Brute. Force. Motif. Search 1. Brute. Force. Motif. Search(DNA, t, n, l) 2. best. Score 0 3. for each s=(s 1, s 2 , . . . , st) from (1, 1. . . 1) to (n-l+1, . . . , n-l+1) 4. if (Score(s, DNA) > best. Score) 5. best. Score score(s, DNA) 6. best. Motif (s 1, s 2 , . . . , st) 7. return best. Motif 10/27/2021 CSE 5290, Fall 2011 123

Running Time of Brute. Force. Motif. Search • Varying (n - l + 1) positions in each of t sequences, we’re looking at (n - l + 1)t sets of starting positions • For each set of starting positions, the scoring function makes l operations, so complexity is l (n – l + 1)t = O(l nt) • That means that for t = 8, n = 1000, l = 10 we must perform approximately 1020 computations – it will take billions of years 10/27/2021 CSE 5290, Fall 2011 124

The Median String Problem • Given a set of t DNA sequences find a pattern that appears in all t sequences with the minimum number of mutations • This pattern will be the motif 10/27/2021 CSE 5290, Fall 2011 125

Hamming Distance • Hamming distance: – d. H(v, w) is the number of nucleotide pairs that do not match when v and w are aligned. For example: d. H(AAAAAA, ACAAAC) = 2 10/27/2021 CSE 5290, Fall 2011 126

Total Distance: Definition (Intuition: let v be a (candidate) motif) – For each DNA sequence i, compute all d. H(v, x), where x is an l-mer with starting position si (1 < si < n – l + 1) – Find minimum of d. H(v, x) among all l-mers in sequence i – Total. Distance(v, DNA, s) is the sum of the minimum Hamming distances for each DNA sequence i – Total. Distance(v, DNA) = mins d. H(v, s), where s is the set of starting positions s 1, s 2, … st 10/27/2021 CSE 5290, Fall 2011 127

The Median String Problem: Formulation • Goal: Given a set of DNA sequences, find a median string • Input: A t x n matrix DNA, and l, the length of the pattern to find • Output: A string v of l nucleotides that minimizes Total. Distance(v, DNA) over all strings of that length 10/27/2021 CSE 5290, Fall 2011 128

Median String Search Algorithm 1. Median. String. Search (DNA, t, n, l) 2. best. Word AAA…A 3. best. Distance ∞ 4. for each l-mer s from AAA…A to TTT…T 5. 6. 7. 8. if Total. Distance(s, DNA) < best. Distance Total. Distance(s, DNA) best. Word s return best. Word 10/27/2021 CSE 5290, Fall 2011 129

Motif Finding Problem == Median String Problem • The Motif Finding is a maximization problem while Median String is a minimization problem • However, the Motif Finding problem and Median String problem are computationally equivalent • Need to show that minimizing Total. Distance is equivalent to maximizing Score 10/27/2021 CSE 5290, Fall 2011 130

We are looking for the same thing l a G g t a c T t C c A t a c g t T A g t a c g t C c A t C c g t a c g G _________ Alignment Profile A C G T 3 0 1 0 3 1 1 0 2 4 0 0 1 4 0 0 0 3 1 0 0 0 5 1 0 1 4 _________ Consensus a c g t Score 3+4+4+5+3+4 • At any column i Scorei + Total. Distancei = t t • Because there are l columns Score + Total. Distance = l * t • Rearranging: Score = l * t - Total. Distance • l * t is constant the minimization of the right side is equivalent to the maximization of the left side Total. Distance 2+1+1+0+2+1 Sum 10/27/2021 5 5 5 5 CSE 5290, Fall 2011 131

Motif Finding Problem vs. Median String Problem • Why bother reformulating the Motif Finding problem into the Median String problem? – The Motif Finding Problem needs to examine all the combinations for s. That is (n - l + 1)t combinations!!! – The Median String Problem needs to examine all 4 l combinations for v. This number is relatively smaller 10/27/2021 CSE 5290, Fall 2011 132

Motif Finding: Improving the Running Time Recall the Brute. Force. Motif. Search: 1. 2. 3. 4. 5. 6. 7. Brute. Force. Motif. Search(DNA, t, n, l) best. Score 0 for each s=(s 1, s 2 , . . . , st) from (1, 1. . . 1) to (n-l+1, . . . , nl+1) if (Score(s, DNA) > best. Score) best. Score(s, DNA) best. Motif (s 1, s 2 , . . . , st) return best. Motif 10/27/2021 CSE 5290, Fall 2011 133

Structuring the Search • How can we perform the line for each s=(s 1, s 2 , . . . , st) from (1, 1. . . 1) to (n-l+1, . . . , nl+1) ? • We need a method for efficiently structuring and navigating the many possible motifs • This is not very different than exploring all tdigit numbers 10/27/2021 CSE 5290, Fall 2011 134

Median String: Improving the Running Time 1. Median. String. Search (DNA, t, n, l) 2. best. Word AAA…A 3. best. Distance ∞ 4. for each l-mer s from AAA…A to TTT…T if Total. Distance(s, DNA) < best. Distance 5. best. Distance Total. Distance(s, DNA) 6. best. Word s 7. return best. Word 10/27/2021 CSE 5290, Fall 2011 135

Structuring the Search – For the Median String Problem we need to consider all 4 l possible l-mers: l aa… aa aa… ac aa… ag aa… at. . tt… tt How to organize this search? 10/27/2021 CSE 5290, Fall 2011 136

Alternative Representation of the Search Space • Let A = 1, C = 2, G = 3, T = 4 • Then the sequences from AA…A to TT…T become: l 11… 11 11… 12 11… 13 11… 14. . 44… 44 • Notice that the sequences above simply list all numbers as if we were counting on base 4 without using 0 as a digit 10/27/2021 CSE 5290, Fall 2011 137

Linked List • Suppose l = 2 Start aa ac ag at ca cc cg ct ga gc gg gt ta tc tg tt • Need to visit all the predecessors of a sequence before visiting the sequence itself 10/27/2021 CSE 5290, Fall 2011 138

Linked List (cont’d) • Linked list is not the most efficient data structure for motif finding • Let’s try grouping the sequences by their prefixes aa ac ag 10/27/2021 at ca cc cg ct ga gc CSE 5290, Fall 2011 gg gt ta tc tg tt 139
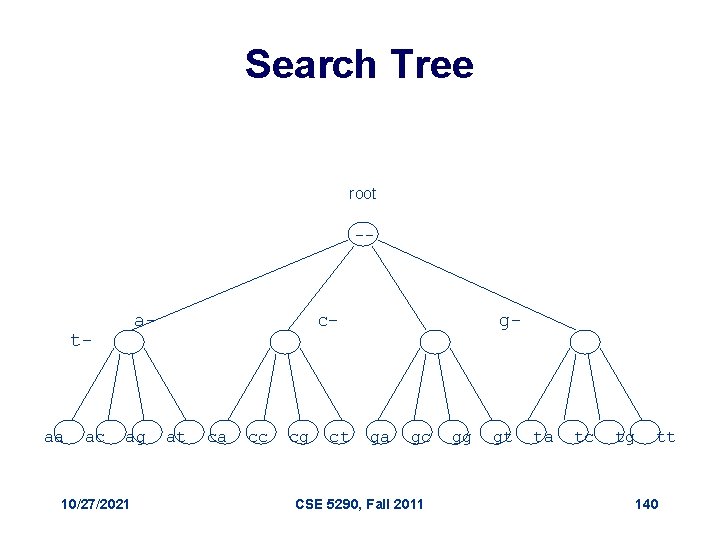
Search Tree root -- a- t- aa ac ag 10/27/2021 c- at ca cc cg ct g- ga gc CSE 5290, Fall 2011 gg gt ta tc tg tt 140

Analyzing Search Trees • Characteristics of the search trees: – The sequences are contained in its leaves – The parent of a node is the prefix of its children • How can we move through the tree? 10/27/2021 CSE 5290, Fall 2011 141

Visit the Next Leaf Given a current leaf a , we need to compute the “next” leaf: 1. Next. Leaf( a, L, k ) 2. for i L to 1 3. if ai < k 4. ai + 1 5. return a 6. ai 1 7. return a 10/27/2021 // a : the array of digits // L: length of the array // k : max digit value CSE 5290, Fall 2011 142

Next. Leaf (cont’d) • The algorithm is common addition in radix k: • Increment the least significant digit • “Carry the one” to the next digit position when the digit is at maximal value 10/27/2021 CSE 5290, Fall 2011 143
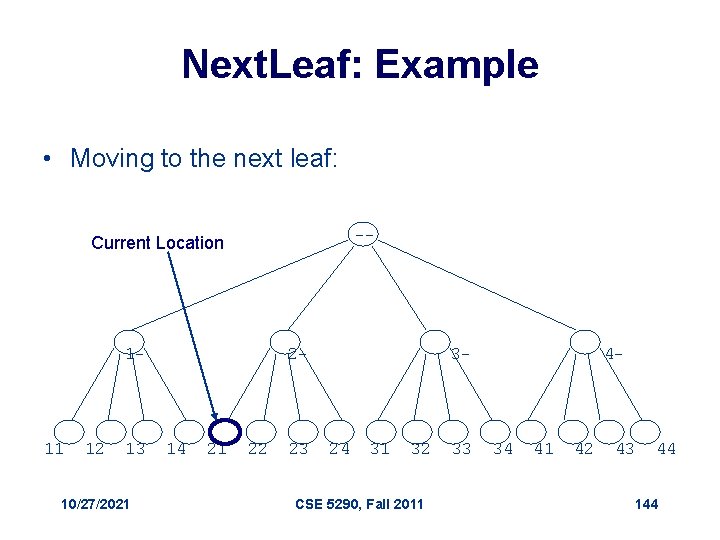
Next. Leaf: Example • Moving to the next leaf: -- Current Location 1 - 11 12 13 10/27/2021 2 - 14 21 22 23 3 - 24 31 32 CSE 5290, Fall 2011 33 4 - 34 41 42 43 44 144
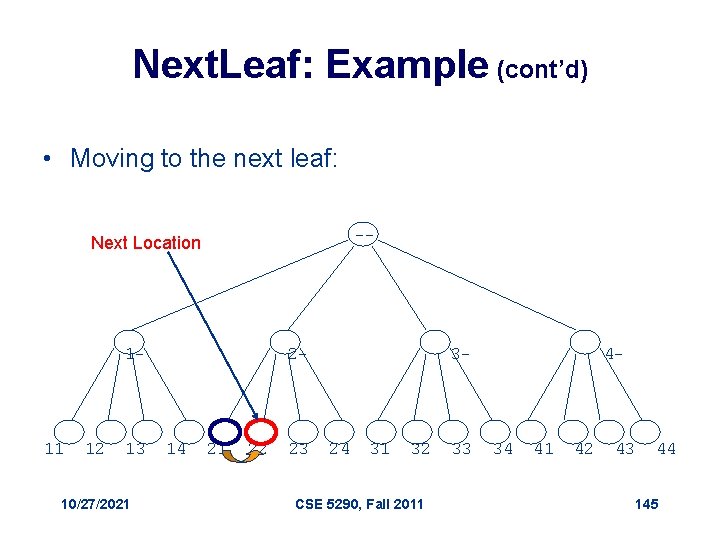
Next. Leaf: Example (cont’d) • Moving to the next leaf: -- Next Location 1 - 11 12 13 10/27/2021 2 - 14 21 22 23 3 - 24 31 32 CSE 5290, Fall 2011 33 4 - 34 41 42 43 44 145

Visit All Leaves • 1. 2. 3. 4. 5. Printing all permutations in ascending order: 6. 7. 10/27/2021 All. Leaves(L, k) // L: length of the sequence a (1, . . . , 1) // k : max digit value while forever // a : array of digits output a a Next. Leaf(a, L, k) if a = (1, . . . , 1) return CSE 5290, Fall 2011 146

Tree Search • So we can search leaves • How about searching all vertices of the tree? • We can do this with a depth first search 10/27/2021 CSE 5290, Fall 2011 147

Visit the Next Vertex 1. Next. Vertex(a, i, L, k) 2. if i < L 3. a i+ 1 1 4. return ( a, i+1) 5. else 6. for j l to 1 7. if aj < k 8. aj + 1 9. return( a, j ) 10. return(a, 0) 10/27/2021 // a : the array of digits // i : prefix length // L: max length // k : max digit value CSE 5290, Fall 2011 148
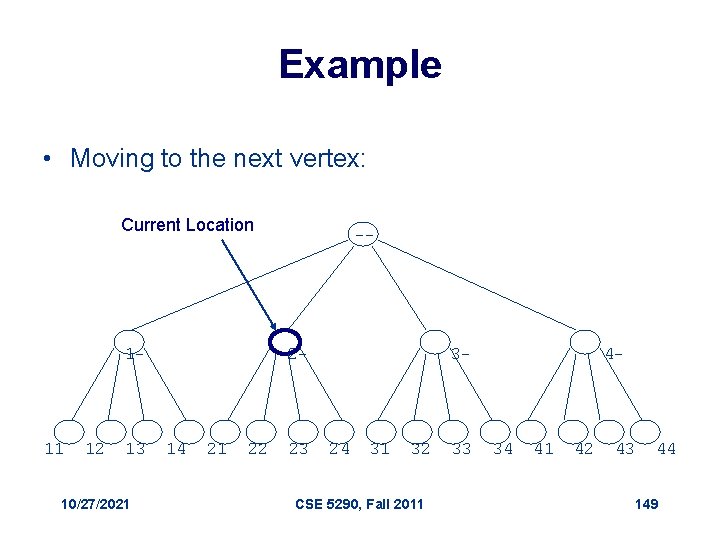
Example • Moving to the next vertex: Current Location 1 - 11 12 13 10/27/2021 -- 2 - 14 21 22 23 3 - 24 31 32 CSE 5290, Fall 2011 33 4 - 34 41 42 43 44 149
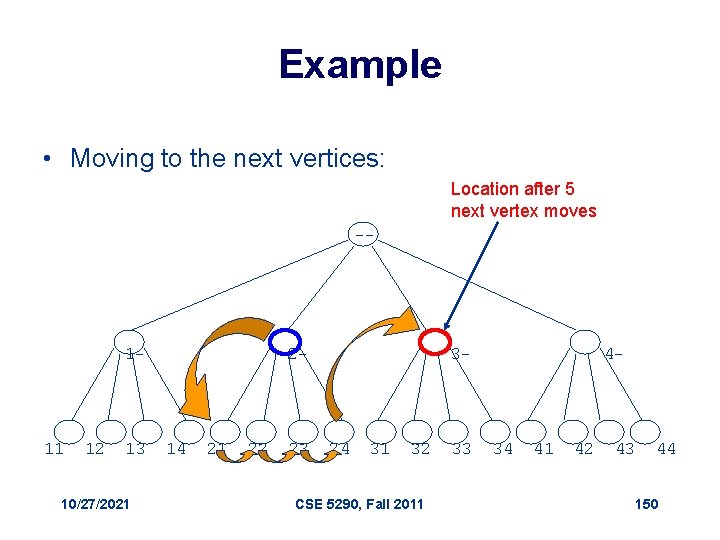
Example • Moving to the next vertices: Location after 5 next vertex moves -- 1 - 11 12 13 10/27/2021 2 - 14 21 22 23 3 - 24 31 32 CSE 5290, Fall 2011 33 4 - 34 41 42 43 44 150

Bypass Move • Given a prefix (internal vertex), find next vertex after skipping all its children 1. 2. 3. 4. 5. 6. Bypass(a, i, L, k) // a: array of digits for j i to 1 // i : prefix length if aj < k // L: maximum length aj +1 // k : max digit value return(a, j) return(a, 0) 10/27/2021 CSE 5290, Fall 2011 151
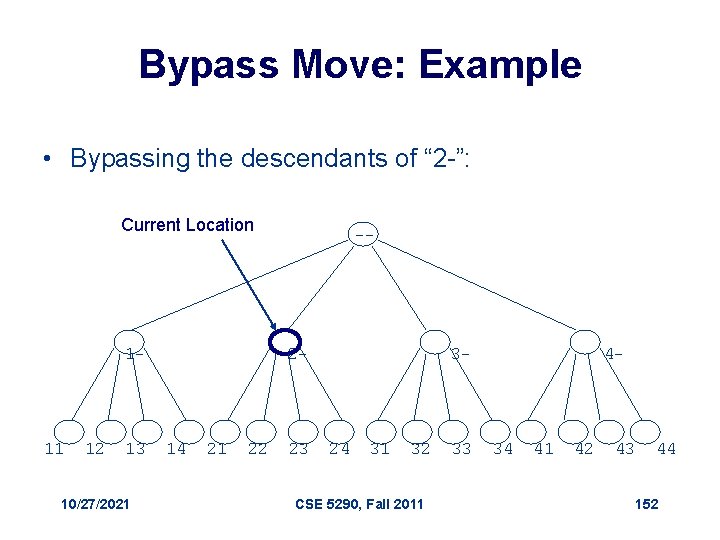
Bypass Move: Example • Bypassing the descendants of “ 2 -”: Current Location 1 - 11 12 13 10/27/2021 -- 2 - 14 21 22 23 3 - 24 31 32 CSE 5290, Fall 2011 33 4 - 34 41 42 43 44 152
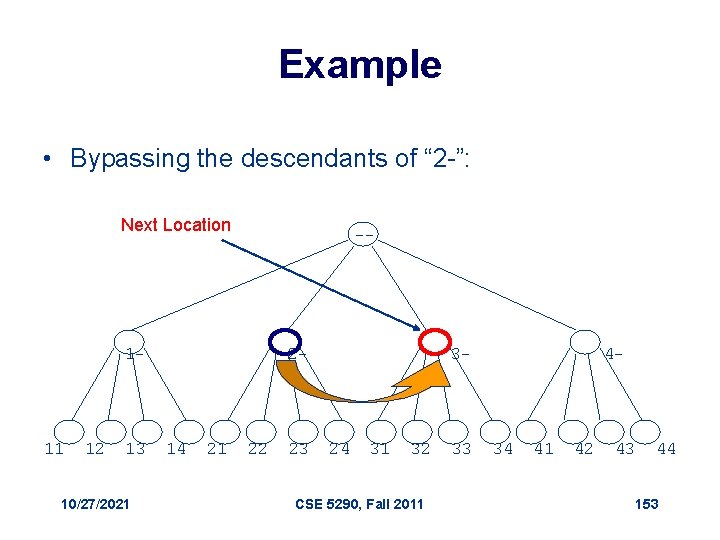
Example • Bypassing the descendants of “ 2 -”: Next Location -- 1 - 11 12 13 10/27/2021 2 - 14 21 22 23 3 - 24 31 32 CSE 5290, Fall 2011 33 4 - 34 41 42 43 44 153

Revisiting Brute Force Search • Now that we have a method for navigating the tree, lets look again at Brute. Force. Motif. Search 10/27/2021 CSE 5290, Fall 2011 154

Brute Force Search Again 1. 2. 3. 4. 5. 6. 7. 8. 9. Brute. Force. Motif. Search. Again(DNA, t, n, l) s (1, 1, …, 1) best. Score(s, DNA) while forever s Next. Leaf (s, t, n- l +1) if (Score(s, DNA) > best. Score) best. Score(s, DNA) best. Motif (s 1, s 2 , . . . , st) return best. Motif 10/27/2021 CSE 5290, Fall 2011 155

Can We Do Better? • Sets of s=(s 1, s 2, …, st) may have a weak profile for the first i positions (s 1, s 2, …, si) • Every row of alignment may add at most l to Score • Optimism: if all subsequent (t-i) positions (si+1, …st) add (t – i ) * l to Score(s, i, DNA) • If Score(s, i, DNA) + (t – i ) * l < Best. Score, it makes no sense to search in vertices of the current subtree – Use By. Pass() 10/27/2021 CSE 5290, Fall 2011 156

Branch and Bound Algorithm for Motif Search • Since each level of the tree goes deeper into search, discarding a prefix discards all following branches • This saves us from looking at (n – l + 1)t-i leaves – Use Next. Vertex() and By. Pass() to navigate the tree 10/27/2021 CSE 5290, Fall 2011 157

Pseudocode for Branch and Bound Motif Search 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. 17. Branch. And. Bound. Motif. Search(DNA, t, n, l) s (1, …, 1) best. Score 0 i 1 while i > 0 if i < t optimistic. Score(s, i, DNA) +(t – i ) * l if optimistic. Score < best. Score (s, i) Bypass(s, i, n-l +1) else (s, i) Next. Vertex(s, i, n-l +1) else if Score(s, DNA) > best. Score(s) best. Motif (s 1, s 2, s 3, …, st) (s, i) Next. Vertex(s, i, t, n-l + 1) return best. Motif 10/27/2021 CSE 5290, Fall 2011 158

Median String Search Improvements • Recall the computational differences between motif search and median string search – The Motif Finding Problem needs to examine all (n-l +1)t combinations for s. – The Median String Problem needs to examine 4 l combinations of v. This number is relatively small • We want to use median string algorithm with the Branch and Bound improvement. 10/27/2021 CSE 5290, Fall 2011 159

Branch and Bound Applied to Median String Search • Note that if the total distance for a prefix is greater than that for the best word so far: Total. Distance (prefix, DNA) > Best. Distance there is no use exploring the remaining part of the word • We can eliminate that branch and BYPASS exploring that branch further 10/27/2021 CSE 5290, Fall 2011 160

Bounded Median String Search 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. 17. 18. 19. Branch. And. Bound. Median. String. Search(DNA, t, n, l ) s (1, …, 1) best. Distance ∞ i 1 while i > 0 if i < l prefix string corresponding to the first i nucleotides of s optimistic. Distance Total. Distance(prefix, DNA) if optimistic. Distance > best. Distance (s, i ) Bypass(s, i, l, 4) else (s, i ) Next. Vertex(s, i, l, 4) else word nucleotide string corresponding to s if Total. Distance(s, DNA) < best. Distance Total. Distance(word, DNA) best. Word word (s, i ) Next. Vertex(s, i, l, 4) return best. Word 10/27/2021 CSE 5290, Fall 2011 161

Improving the Bounds • Given an l-mer w, divided into two parts at point i – u : prefix w 1, …, wi, – v : suffix wi+1, . . . , wl • Find minimum distance for u in a sequence • No instances of u in the sequence have distance less than the minimum distance • Note this doesn’t tell us anything about whether u is part of any motif. We only get a minimum distance for prefix u 10/27/2021 CSE 5290, Fall 2011 162

Improving the Bounds (cont’d) • Repeating the process for the suffix v gives us a minimum distance for v • Since u and v are two substrings of w, and included in motif w, we can assume that the minimum distance of u plus minimum distance of v can only be less than the minimum distance for w 10/27/2021 CSE 5290, Fall 2011 163

Better Bounds 10/27/2021 CSE 5290, Fall 2011 164

Better Bounds (cont’d) • If d(prefix) + d(suffix) > best. Distance: – Motif w (prefix. suffix) cannot give a better (lower) score than d(prefix) + d(suffix) – In this case, we can By. Pass() 10/27/2021 CSE 5290, Fall 2011 165

More on the Motif Problem • Exhaustive Search and Median String are both exact algorithms • They always find the optimal solution, though they may be too slow to perform practical tasks • Many algorithms sacrifice optimal solution for speed • Examples: Local search, Stochastic sampling… 10/27/2021 CSE 5290, Fall 2011 166
- Slides: 166