CSE 280 b Population Genetics Vineet BafnaPavel Pevzner

CSE 280 b: Population Genetics Vineet Bafna/Pavel Pevzner March 2006 www. cse. ucsd. edu/classes/sp 06/cse 280 b Vineet Bafna

Population Sub-structure March 2006 Vineet Bafna

Population sub-structure can increase LD • • • Consider two populations that were isolated and evolving independently. They might have different allele frequencies in some regions. Pick two regions that are far apart (LD is very low, close to 0) March 2006 Vineet Bafna 0 0 0 1 0 0 0 . . . . 1 1 0 1 1 1 . . . . 0 0 0 1 0 0 0 Pop. A p 1=0. 1 q 1=0. 9 P 11=0. 1 D=0. 01 Pop. B p 1=0. 9 q 1=0. 1 P 11=0. 1 D=0. 01

Recent ad-mixing of population • • If the populations came together recently (Ex: African and European population), artificial LD might be created. D = 0. 15 (instead of 0. 01), increases 10 -fold This spurious LD might lead to false associations Other genetic events can cause LD to arise, and one needs to be careful March 2006 Vineet Bafna 0 0 0 1 0 0 0 . . . . 1 1 0 1 1 1 . . . . 0 0 0 1 0 0 0 Pop. A+B p 1=0. 5 q 1=0. 5 P 11=0. 1 D=0. 1 -0. 25=0. 15

Determining population sub-structure • • Given a mix of people, can you sub-divide them into ethnic populations. Turn the ‘problem’ of spurious LD into a clue. – – – Find markers that are too far apart to show LD If they do show LD (correlation), that shows the existence of multiple populations. Sub-divide them into populations so that LD disappears. March 2006 Vineet Bafna

Determining Population sub-structure • • • Same example as before: The two markers are too similar to show any LD, yet they do show LD. However, if you split them so that all 0. . 1 are in one population and all 1. . 0 are in another, LD disappears March 2006 Vineet Bafna 0 0 0 1 0 0 0 . . . . 1 1 0 1 1 1 . . . . 0 0 0 1 0 0 0

Iterative algorithm for population substructure • • Define N = number of individuals (each has a single chromosome) k = number of sub-populations. Z {1. . k}N is a vector giving the sub-population. – • • Zi=k’ => individual i is assigned to population k’ Xi, j = allelic value for individual i in position j Pk, j, l = frequency of allele l at position j in population k March 2006 Vineet Bafna

Example • • • Ex: consider the following assignment P 1, 1, 0 = 0. 9 P 2, 1, 0 = 0. 1 March 2006 Vineet Bafna 1 1 1 1 1 0 0 0 . . 1 1 0 1 1 1 1 2 2 2 2 2 1 1 0 1 1 1 1 . . 0 0 0 1 0 0 0

Goal • • X is known. P, Z are unknown. The goal is to estimate Pr(P, Z|X) Various learning techniques can be employed. – – – • max. P, Z Pr(X|P, Z) (Max likelihood estimate) max. P, Z Pr(X|P, Z) Pr(P, Z) (MAP) Sample P, Z from Pr(P, Z|X) Here a Bayesian (MCMC) scheme is employed to sample from Pr(P, Z|X). We will only consider a simplified version March 2006 Vineet Bafna

Algorithm: Structure • Iteratively estimate – • • (Z(0), P(0)), (Z(1), P(1)), . . , (Z(m), P(m)) After ‘convergence’, Z(m) is the answer. Iteration – – Guess Z(0) For m = 1, 2, . . • • • Sample P(m) from Pr(P | X, Z(m-1)) Sample Z(m) from Pr(Z | X, P(m)) How is this sampling done? March 2006 Vineet Bafna

Example • • • Choose Z at random, so each individual is assigned to be in one of 2 populations. See example. Now, we need to sample P(1) from Pr(P | X, Z(0)) Simply count Nk, j, l = number of people in pouplation k which have allele l in position j pk, j, l = Nk, j, l / N March 2006 Vineet Bafna 1 2 2 1 1 2 1 2 0 0 0 1 0 0 0 . . 1 1 0 1 1 1 1 2 2 1 1 1 0 1 1 1 1 . . 0 0 0 1 0 0 0

Example • • Nk, j, l = number of people in population k which have allele l in position j pk, j, l = Nk, j, l / Nk, j, * N 1, 1, 0 = 4 N 1, 1, 1 = 6 p 1, 1, 0 = 4/10 p 1, 2, 0 = 4/10 Thus, we can sample P(m) March 2006 Vineet Bafna 1 2 2 1 1 2 1 2 0 0 0 1 0 0 0 . . 1 1 0 1 1 1 1 2 2 1 1 1 0 1 1 1 1 . . 0 0 0 1 0 0 0
![Sampling Z • • • Pr[Z 1 = 1] = Pr[” 01” belongs to Sampling Z • • • Pr[Z 1 = 1] = Pr[” 01” belongs to](http://slidetodoc.com/presentation_image_h2/1d2d044f9367d9c442dccc4be195aee3/image-13.jpg)
Sampling Z • • • Pr[Z 1 = 1] = Pr[” 01” belongs to population 1]? We know that each position should be in linkage equilibrium and independent. Pr[” 01” |Population 1] = p 1, 1, 0 * p 1, 2, 1 =(4/10)*(6/10)=(0. 24) Pr[” 01” |Population 2] = p 2, 1, 0 * p 2, 2, 1 = (6/10)*(4/10)=0. 24 Pr [Z 1 = 1] = 0. 24/(0. 24+0. 24) = 0. 5 March 2006 Vineet Bafna Assuming, HWE, and LE

Sampling • • Suppose, during the iteration, there is a bias. Then, in the next step of sampling Z, we will do the right thing Pr[“ 01”| pop. 1] = p 1, 1, 0 * p 1, 2, 1 = 0. 7*0. 7 = 0. 49 Pr[“ 01”| pop. 2] = p 2, 1, 0 * p 2, 2, 1 =0. 3*0. 3 = 0. 09 Pr[Z 1 = 1] = 0. 49/(0. 49+0. 09) = 0. 85 Pr[Z 6 = 1] = 0. 49/(0. 49+0. 09) = 0. 85 Eventually all “ 01” will become 1 population, and all “ 10” will become a second population March 2006 Vineet Bafna 1 1 1 2 1 2 1 1 0 0 0 . . 1 1 0 1 1 1 1 2 2 2 1 1 1 0 1 1 1 1 . . 0 0 0 1 0 0 0

Allowing for admixture • • Define qi, k as the fraction of individual i that originated from population k. Iteration – – Guess Z(0) For m = 1, 2, . . • • March 2006 Sample P(m), Q(m) from Pr(P, Q | X, Z(m-1)) Sample Z(m) from Pr(Z | X, P(m), Q(m)) Vineet Bafna
Estimating Z (admixture case) • Instead of estimating Pr(Z(i)=k|X, P, Q), (origin of individual i is k), we estimate Pr(Z(i, j, l)=k|X, P, Q) i, 1 i, 2 j March 2006 Vineet Bafna
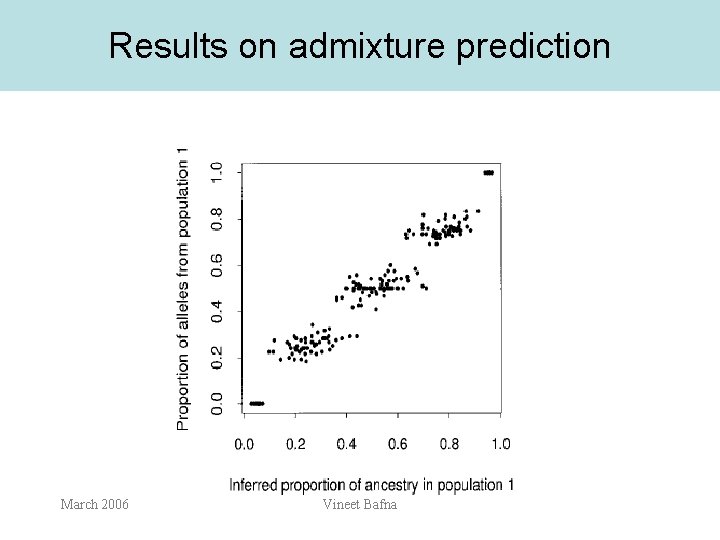
Results on admixture prediction March 2006 Vineet Bafna

Results: Thrush data • • For each individual, q(i) is plotted as the distance to the opposite side of the triangle. The assignment is reliable, and there is evidence of admixture. March 2006 Vineet Bafna
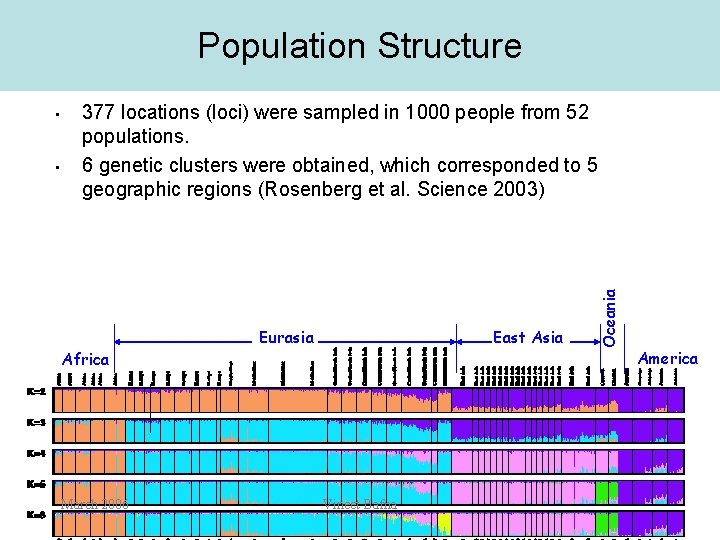
Population Structure • 377 locations (loci) were sampled in 1000 people from 52 populations. 6 genetic clusters were obtained, which corresponded to 5 geographic regions (Rosenberg et al. Science 2003) Africa March 2006 Eurasia East Asia Vineet Bafna Oceania • America

Population sub-structure: research problem • • Systematically explore the effect of admixture. Can admixture be predicted for a locus, or for an individual The sampling approach may or may not be appropriate. Formulate as an optimization/learning problem: – – (w/out admixture). Assign individuals to sub-populations so as to maximize linkage equilibrium, and hardy weinberg equilibrium in each of the sub-populations (w/ admixture) Assign (individuals, loci) to sub-populations March 2006 Vineet Bafna

Allowing for admixture • • Define qi, k as the fraction of individual i that originated from population k. Iteration – – Guess Z(0) For m = 1, 2, . . • • March 2006 Sample P(m), Q(m) from Pr(P, Q | X, Z(m-1)) Sample Z(m) from Pr(Z | X, P(m), Q(m)) Vineet Bafna
Estimating Z (admixture case) • Instead of estimating Pr(Z(i)=k|X, P, Q), (origin of individual i is k), we estimate Pr(Z(i, j, l)=k|X, P, Q) i, 1 i, 2 j March 2006 Vineet Bafna
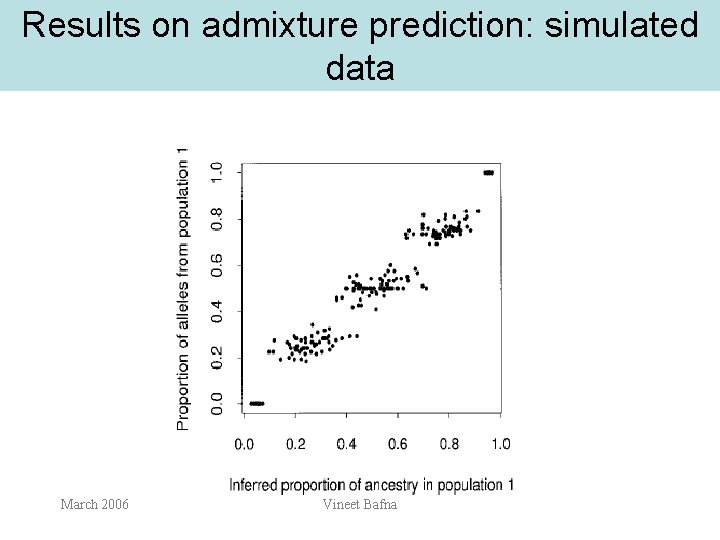
Results on admixture prediction: simulated data March 2006 Vineet Bafna

Results: Thrush data • • For each individual, q(i) is plotted as the distance to the opposite side of the triangle. The assignment is reliable, and there is evidence of admixture. March 2006 Vineet Bafna
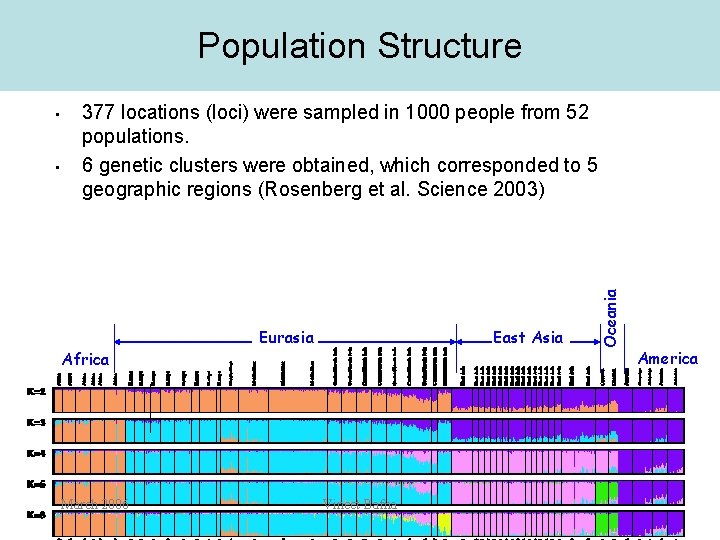
Population Structure • 377 locations (loci) were sampled in 1000 people from 52 populations. 6 genetic clusters were obtained, which corresponded to 5 geographic regions (Rosenberg et al. Science 2003) Africa March 2006 Eurasia East Asia Vineet Bafna Oceania • America
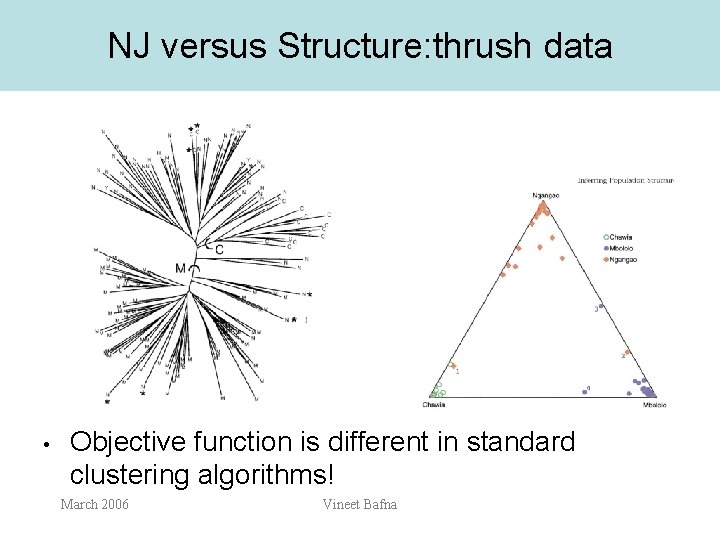
NJ versus Structure: thrush data • Objective function is different in standard clustering algorithms! March 2006 Vineet Bafna

Population sub-structure: research problem • • Systematically explore the effect of admixture. Can admixture be predicted for a locus, or for an individual The sampling approach may or may not be appropriate. Formulate as an optimization/learning problem: – – (w/out admixture). Assign individuals to sub-populations so as to maximize linkage equilibrium, and hardy weinberg equilibrium in each of the sub-populations (w/ admixture) Assign (individuals, loci) to sub-populations March 2006 Vineet Bafna

Admixture mapping March 2006 Vineet Bafna

Estimating Recombination Rates March 2006 Vineet Bafna
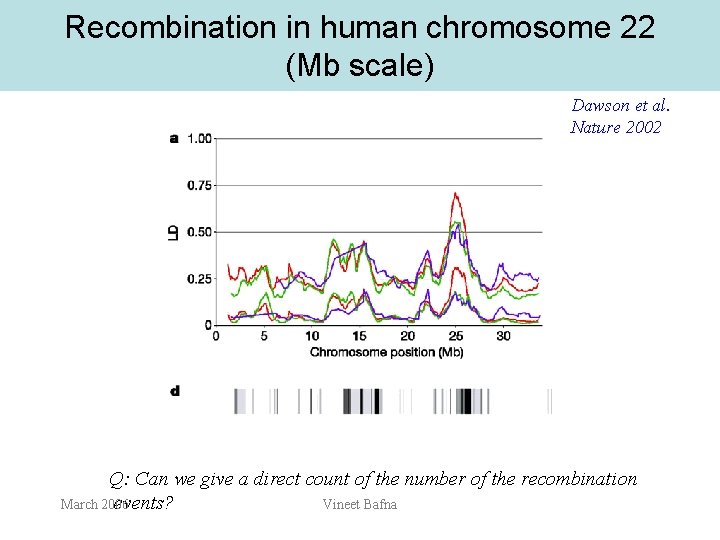
Recombination in human chromosome 22 (Mb scale) Dawson et al. Nature 2002 Q: Can we give a direct count of the number of the recombination events? March 2006 Vineet Bafna
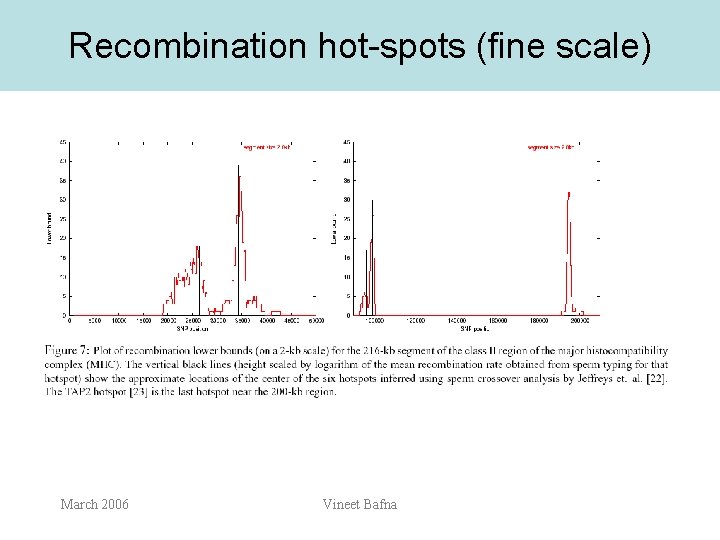
Recombination hot-spots (fine scale) March 2006 Vineet Bafna
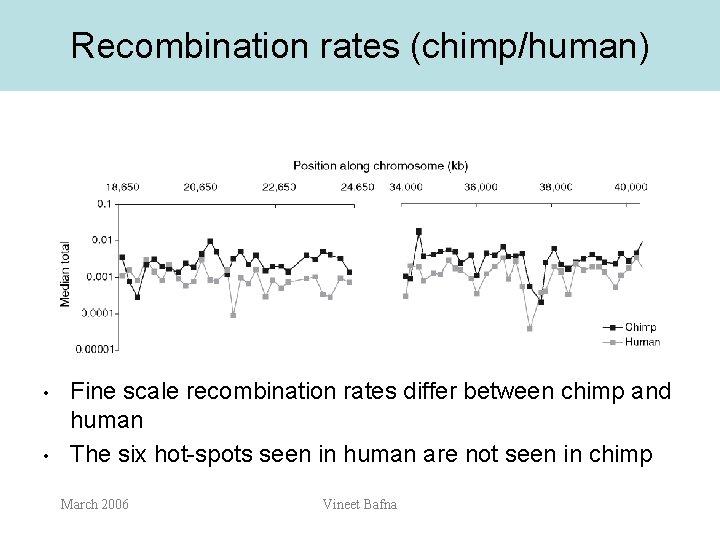
Recombination rates (chimp/human) • • Fine scale recombination rates differ between chimp and human The six hot-spots seen in human are not seen in chimp March 2006 Vineet Bafna

Combinatorial Bounds for estimating recombination rate • • Recall that expected #recombinations = log n Procedure • • Generate N random ARGs that results in the given sample Compute mean of the number of recombinations Alternatively, generate a summary statistic s from the population. For each , generate many populations, and compute the mean and variance of s (This only needs to be done once). Use this to select the most likely What is the correct summary statistic? Today, we talk about the min. number of recombination events as a possible summary statistic. It is not the most natural, but it is the most interesting computationally. March 2006 Vineet Bafna
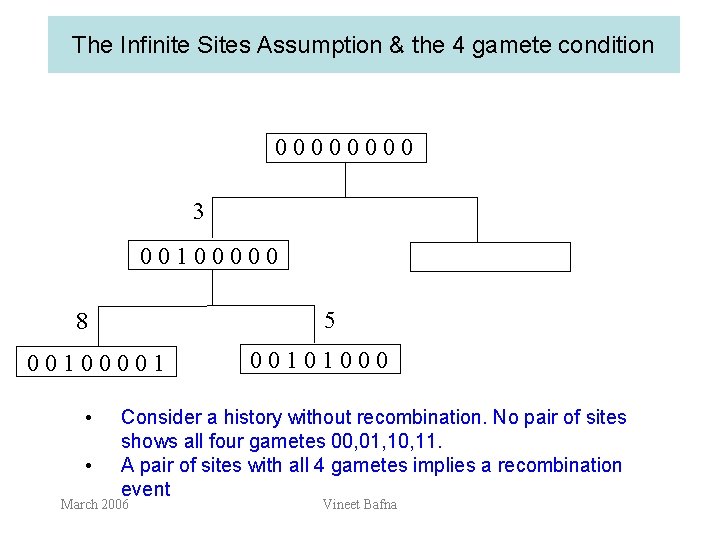
The Infinite Sites Assumption & the 4 gamete condition 0000 3 00100000 5 8 00100001 • • 00101000 Consider a history without recombination. No pair of sites shows all four gametes 00, 01, 10, 11. A pair of sites with all 4 gametes implies a recombination event March 2006 Vineet Bafna
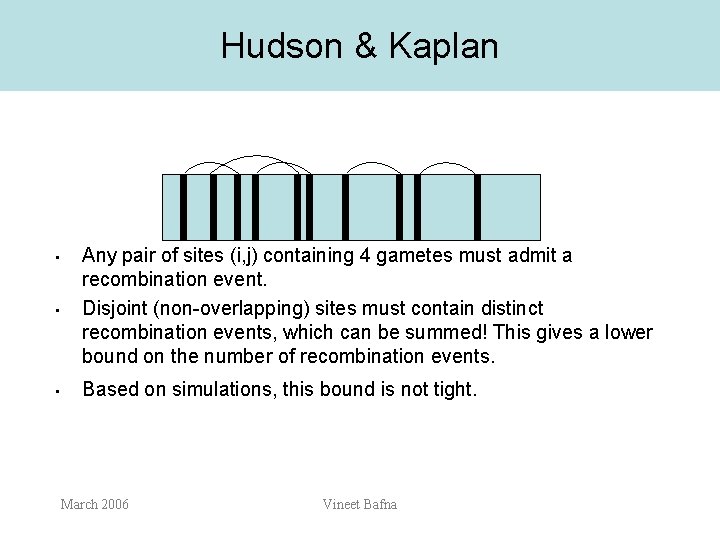
Hudson & Kaplan • Any pair of sites (i, j) containing 4 gametes must admit a recombination event. Disjoint (non-overlapping) sites must contain distinct recombination events, which can be summed! This gives a lower bound on the number of recombination events. • Based on simulations, this bound is not tight. • March 2006 Vineet Bafna
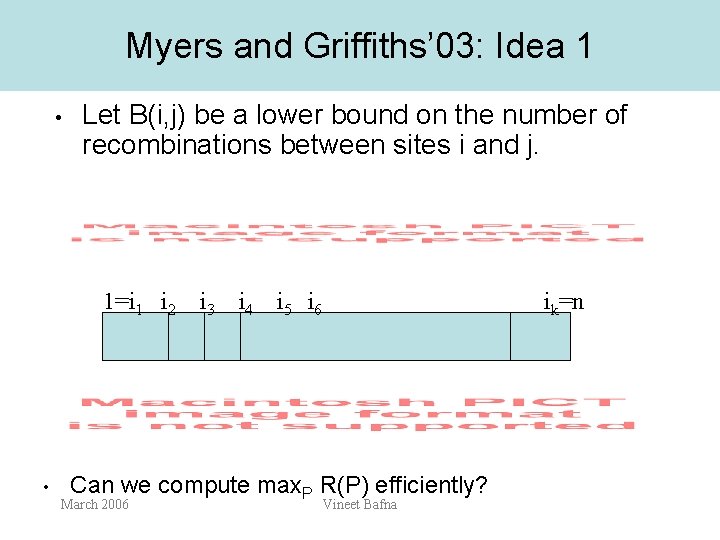
Myers and Griffiths’ 03: Idea 1 • Let B(i, j) be a lower bound on the number of recombinations between sites i and j. 1=i 1 i 2 i 3 i 4 i 5 i 6 • ik=n Can we compute max. P R(P) efficiently? March 2006 Vineet Bafna

The Rm bound March 2006 Vineet Bafna

Improved lower bounds • • • The Rm bound also gives a general technique for combining local lower bounds into an overall lower bound. In the example, Rm=2, but we cannot give any ARG with 2 recombination events. Can we improve upon Hudson and Kaplan to get better local lower bounds? March 2006 Vineet Bafna 000 001 010 011 100 101 110 111

Myers & Griffiths: Idea 2 • • Consider the history of individuals. Let Ht denote the number of distinct haplotypes at time t One of three things might happen at time t: – Mutation: Ht increase by at most 1 – Recombination: Ht increase by at most 1 – Coalescence: Ht does not increase March 2006 Vineet Bafna

The RH bound Ex: R>= 8 -3 -1=4 March 2006 Vineet Bafna 000 001 010 011 100 101 110 111

RH bound • • In general, RH can be quite weak: – consider the case when S>H However, it can be improved – Partitioning idea: sum RH over disjoint intervals – Apply to any subset of columns. Ex: Apply RH to the yellow columns 000000001 0000000100000001 100000001 10000000 11111111 (BB’ 05) March 2006 Vineet Bafna

The Rs bound • • Compute the minimum number of recombination events R in any ARG. Note that, we do not explicitly construct the ARG. Consider a matrix with M with H rows and S columns. – – The rows correspond to haplotypes. Columns correspond to sites. March 2006 Vineet Bafna

Rs bound: Observation I s § Non-informative column: If a site contains at most one 1, or one 0, then in any history, it can be obtained by adding a mutation to a branch. § § EX: if a is the haplotype containing a 1, It can simply be added to the branch without increasing number of recombination events R(M) = R(M-{s}) March 2006 Vineet Bafna 0 0 0 : : 1 a a b c
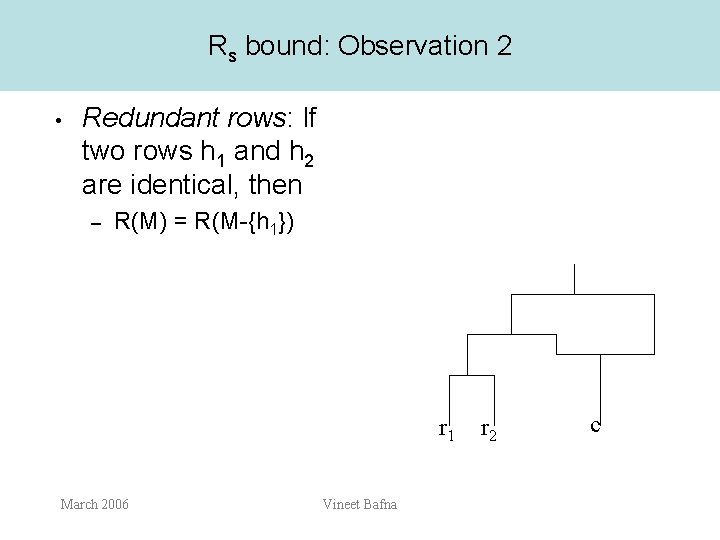
Rs bound: Observation 2 • Redundant rows: If two rows h 1 and h 2 are identical, then – R(M) = R(M-{h 1}) r 1 March 2006 Vineet Bafna r 2 c

Rs bound: Observation 3 • Suppose M has no noninformative columns, or redundant rows. – Then, at least one of the haplotypes is a recombinant. – There exists h s. t. R(M) = R(M-{h})+1 – Which h should you choose? March 2006 Vineet Bafna

Rs bound (Procedural) Procedure Compute_Rs(M) If non-informative column s return (Compute_Rs(M-{s})) Else if redundant row h return (Compute_Rs(M-{h})) Else return (1 + minh(Compute_Rs(M-{h})) March 2006 Vineet Bafna
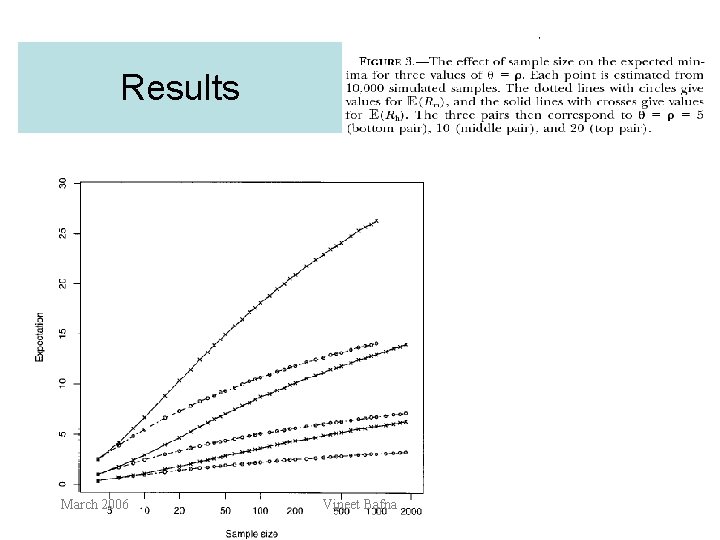
Results March 2006 Vineet Bafna

Additional results/problems • • • Using dynamic programming, Rs can be computed in 2^n poly(mn) time. Also, Rs can be augmented to handle intermediates. Are there poly. time lower bounds? – • The number of connected components in the conflict graph is a lower bound (BB’ 04). Fast algorithms for computing ARGs with minimum recombination. – Poly. Time to get ARG with 0 recombination – Poly. Time to get ARGs that are galled trees (Gusfield’ 03) March 2006 Vineet Bafna
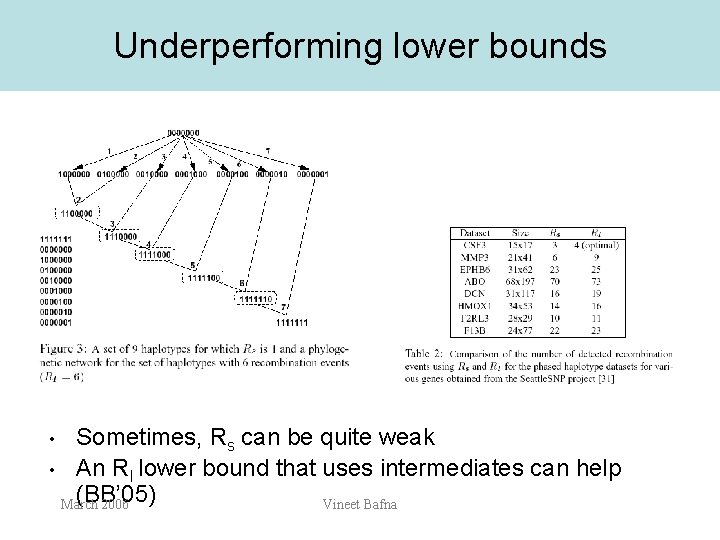
Underperforming lower bounds Sometimes, Rs can be quite weak • An RI lower bound that uses intermediates can help (BB’ 05) March 2006 Vineet Bafna •
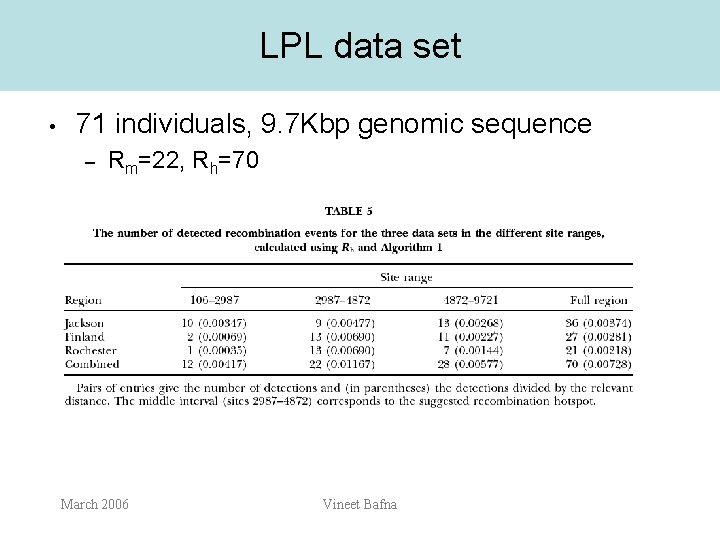
LPL data set • 71 individuals, 9. 7 Kbp genomic sequence – Rm=22, Rh=70 March 2006 Vineet Bafna

March 2006 Vineet Bafna
- Slides: 51