CSCI 6904 Genomics and Biological Computing Lecture 3

CSCI 6904 Genomics and Biological Computing Lecture 3 – Conceptual Biology Cells, Gene circuits Conceptual Biology

Overview Computing in Biological systems Cells are computing information and react programatically to various situations. We will have a brief look at what is a cell and how they “compute”. Evolutionary emergence of Networks These Circuits of gene products are arising in a stochastic manner. We will have a quick look on how this random walk results in a combinatorial strategy to evolve solutions. Investigating Networks None of these network is visible, investigating the relationships in the physical world is a resource consuming operation. Building Knowledge models of cells using text mining Present a test case called GENEWAY.
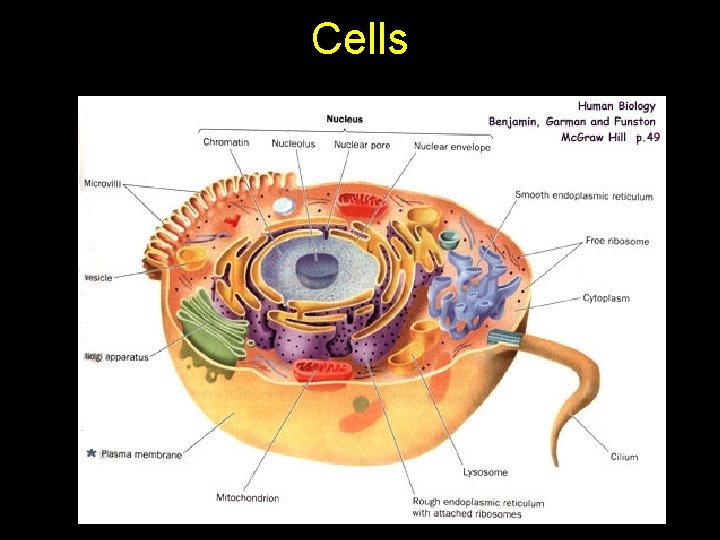
Cells

Scope of molecular Biology Molecular biology tries to organize a stochastically evolved system comprising hundreds of thousands components. None of these components can be seen, even under the most powerful microscopes. They are usually present in the 10 -8 – 10 -12 grams scale. They degrade in a matter of second to hours. The bottomline is: Everything we know about this system comes from fragments of information. Many of these are going to be refuted over time.
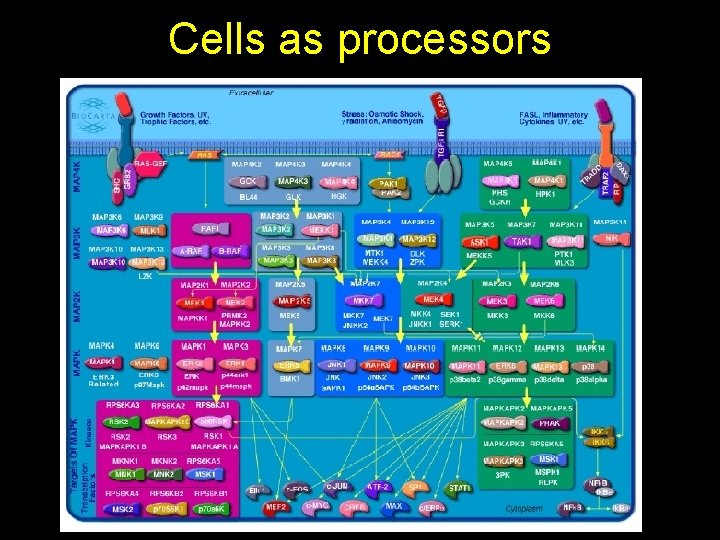
Cells as processors
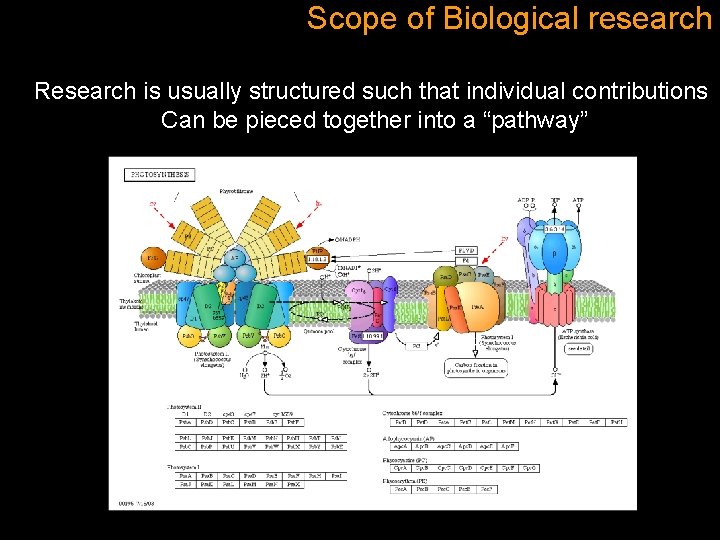
Scope of Biological research Research is usually structured such that individual contributions Can be pieced together into a “pathway”
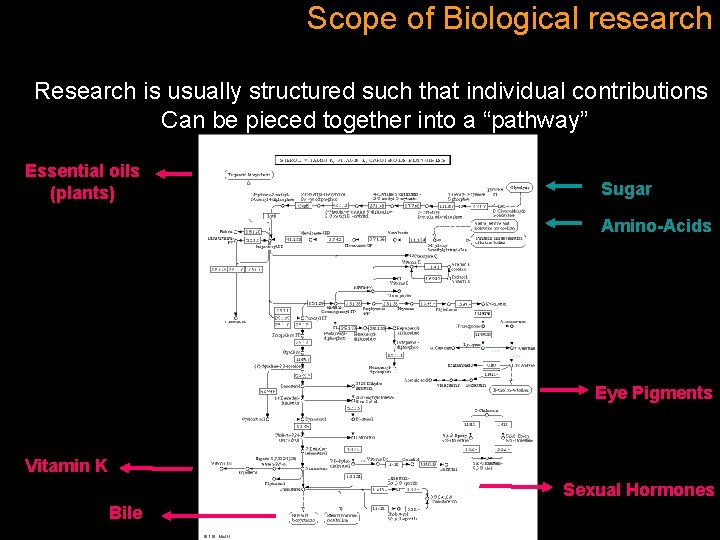
Scope of Biological research Research is usually structured such that individual contributions Can be pieced together into a “pathway” Essential oils (plants) Sugar Amino-Acids Eye Pigments Vitamin K Sexual Hormones Bile

Networks How do they come into being? Combinatorial assembly during a stochastic process. What is done to understand the main pathways? Grasping event the smallest facts about 1 edge in the graph is a feat.
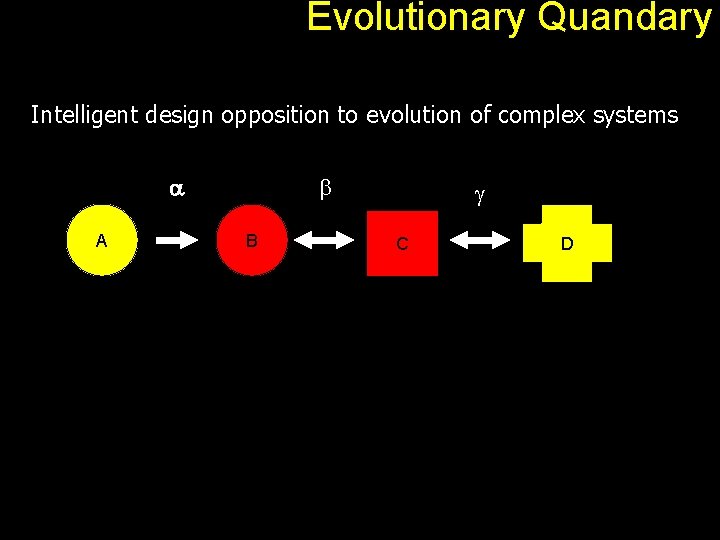
Evolutionary Quandary Intelligent design opposition to evolution of complex systems a A b B g C D
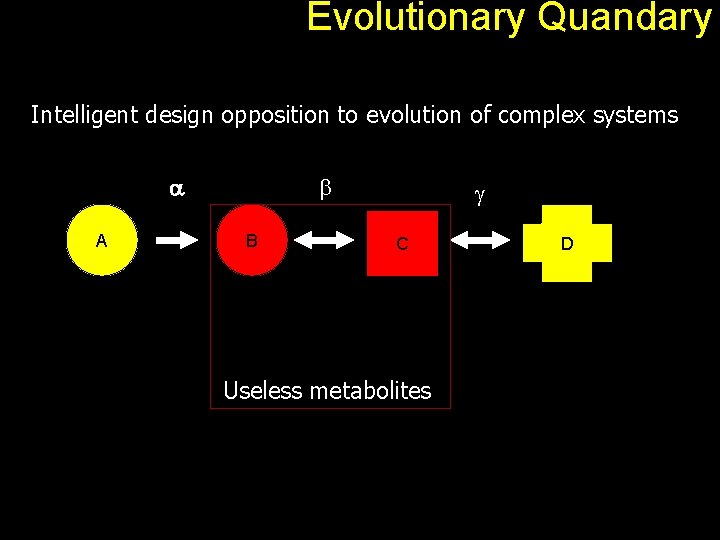
Evolutionary Quandary Intelligent design opposition to evolution of complex systems a A b B g C Useless metabolites D
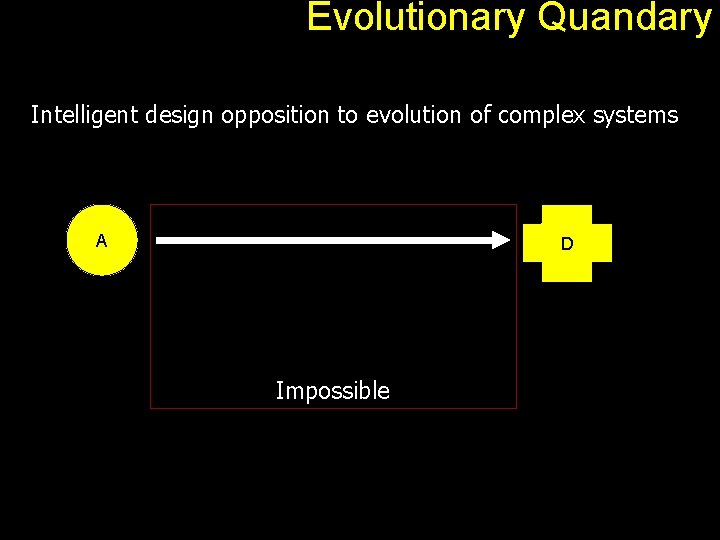
Evolutionary Quandary Intelligent design opposition to evolution of complex systems A D Impossible
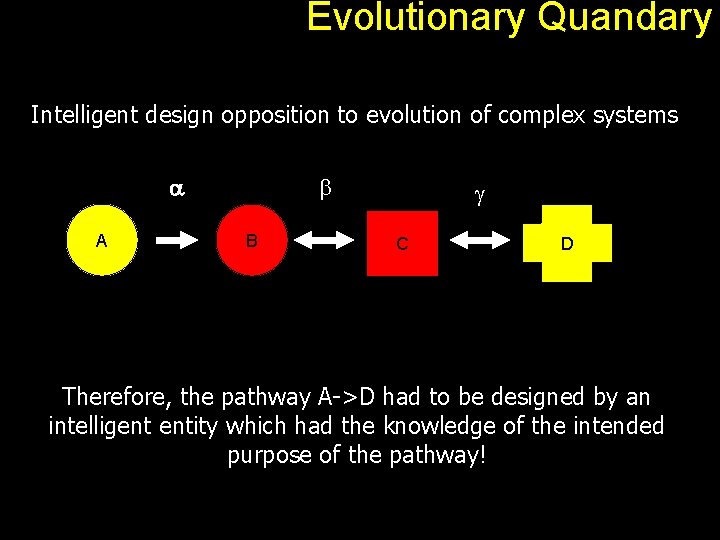
Evolutionary Quandary Intelligent design opposition to evolution of complex systems a A b B g C D Therefore, the pathway A->D had to be designed by an intelligent entity which had the knowledge of the intended purpose of the pathway!

Closer look at high-level genes organization A modular system Proteins can be broken down into domains. A combinatorial effect Domains can assemble in a combinatorial fashion to try together a vast array of potential biological activities.

Proteins are made of domains Proteins are organized into domains http: //www. ncbi. nlm. nih. gov Transcription factor e. F 1 / (PDB: 1 IJF)

Proteins are made of domains Domains have several interesting properties. http: //www. ncbi. nlm. nih. gov Transcription factor e. F 1 / (PDB: 1 IJF)
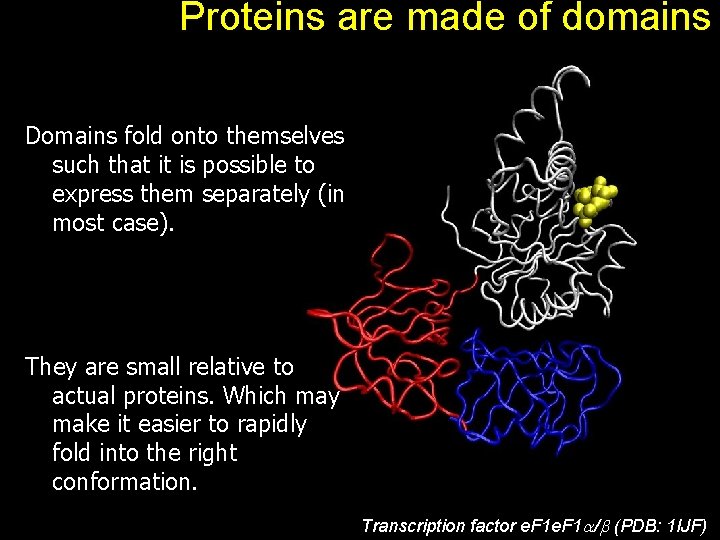
Proteins are made of domains Domains fold onto themselves such that it is possible to express them separately (in most case). They are small relative to actual proteins. Which may make it easier to rapidly fold into the right conformation. Transcription factor e. F 1 / (PDB: 1 IJF)
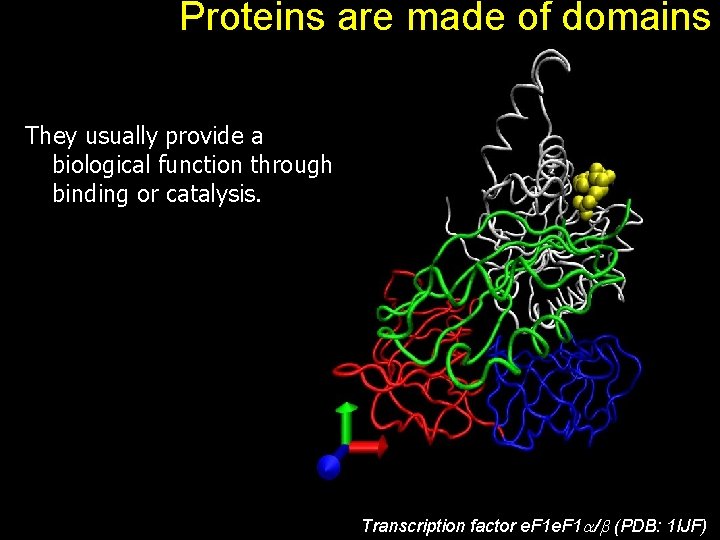
Proteins are made of domains They usually provide a biological function through binding or catalysis. Transcription factor e. F 1 / (PDB: 1 IJF)
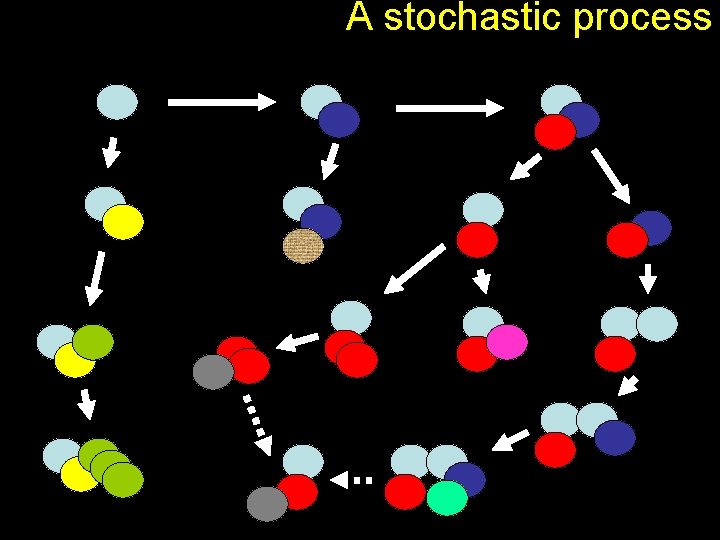
A stochastic process
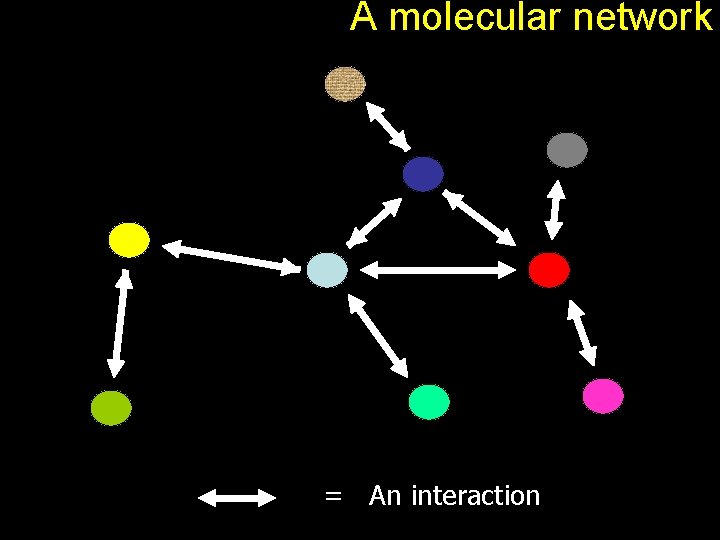
A molecular network = An interaction
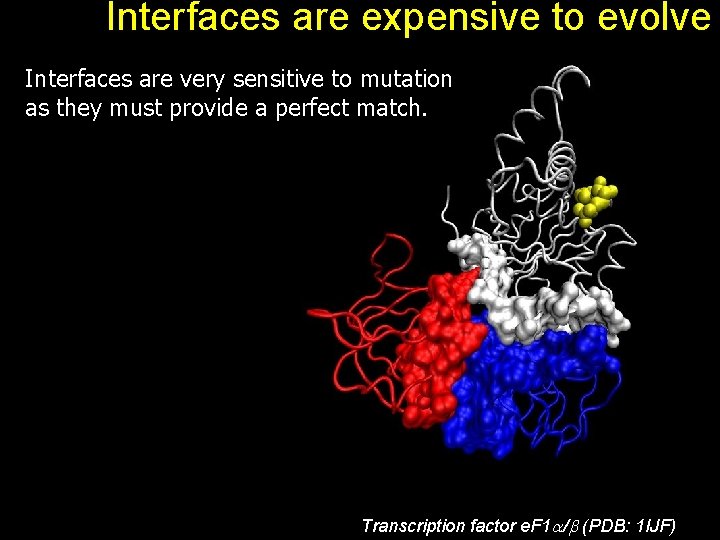
Interfaces are expensive to evolve Interfaces are very sensitive to mutation as they must provide a perfect match. Transcription factor e. F 1 / (PDB: 1 IJF)

Network of Metabolites are essentially forming network with a scale -free property, which parallels the stochastic assembly of domains. At least, this appears to be true with the data there are so far. http: //www. genego. com/about/products. shtml Rzhetsky and Gomez, 2001. Bioinformatics, 17: 988 -996
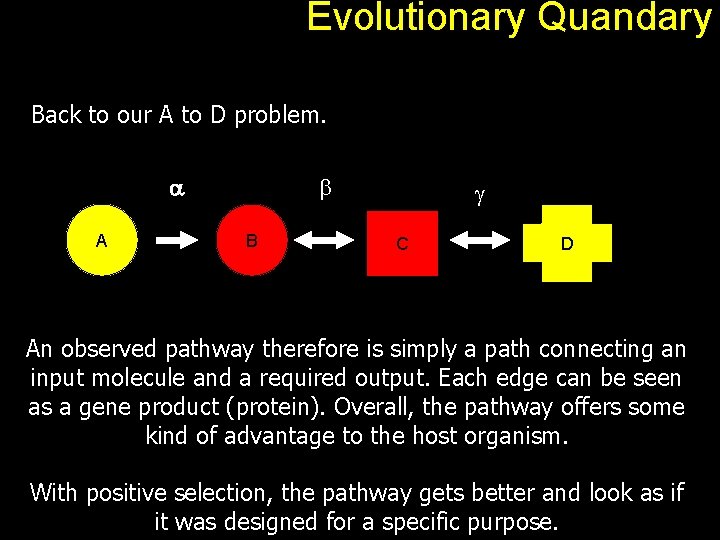
Evolutionary Quandary Back to our A to D problem. a A b B g C D An observed pathway therefore is simply a path connecting an input molecule and a required output. Each edge can be seen as a gene product (protein). Overall, the pathway offers some kind of advantage to the host organism. With positive selection, the pathway gets better and look as if it was designed for a specific purpose.
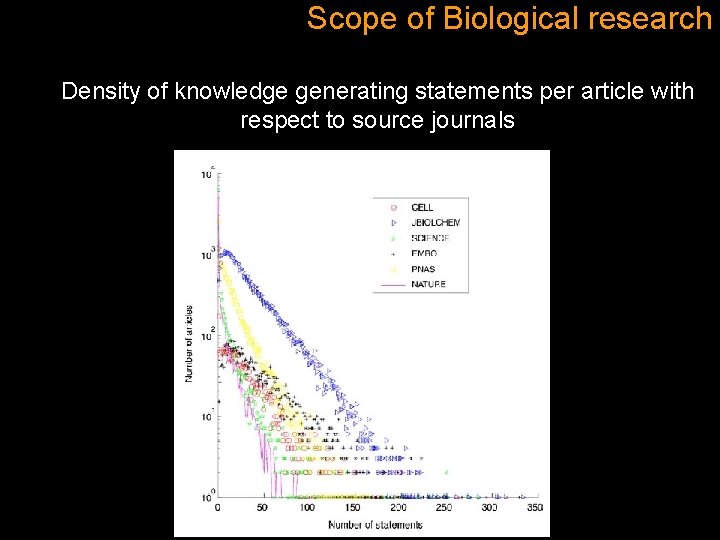
Scope of Biological research Density of knowledge generating statements per article with respect to source journals

Where it becomes a bioinformatic’s problem: Nature of the problem Building a global model from plain English text sources. Size Complexity What is done in the Gene. Ways project The workflow of their integrated system What I think it really means in the long run The relationship between research and researchers (The right information system will be the next big thing)

Motivation Human limitations and Data-heavy and knowledge-heavy Disciplines Synthesizing Hypothesis building Visualizing Records keeping Modeling Knowledge Streamlining Structuring (Directing) (Changing the way research is communicated? )
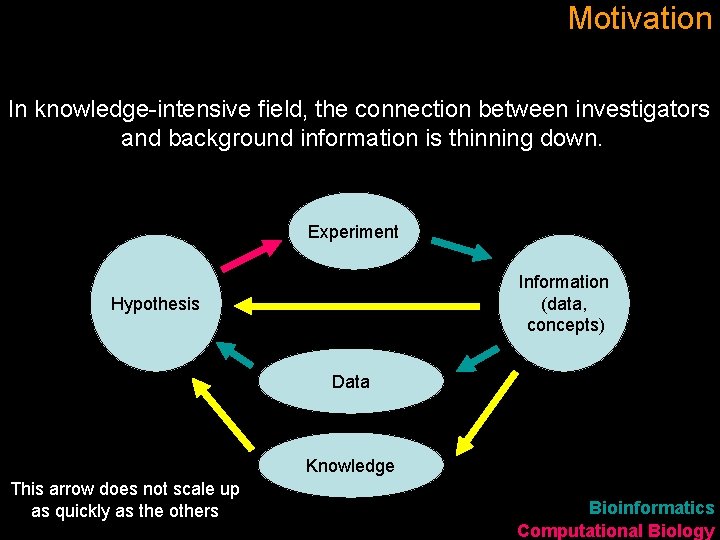
Motivation In knowledge-intensive field, the connection between investigators and background information is thinning down. Experiment Information (data, concepts) Hypothesis Data Knowledge This arrow does not scale up as quickly as the others Bioinformatics Computational Biology
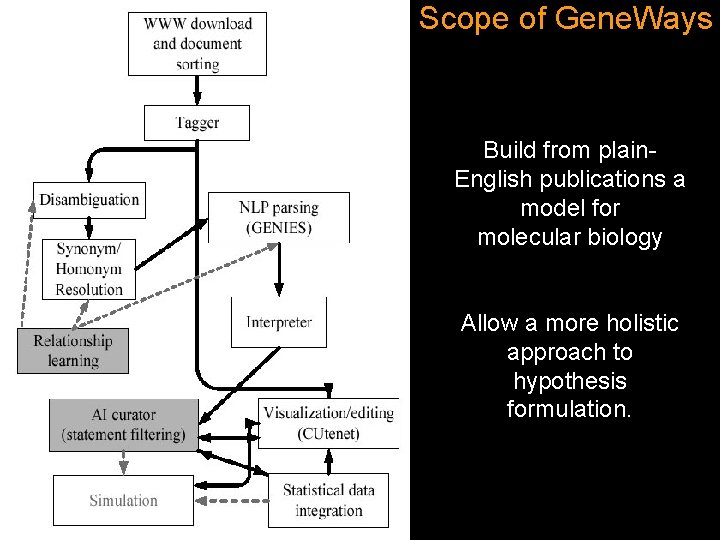
Scope of Gene. Ways Build from plain. English publications a model for molecular biology Allow a more holistic approach to hypothesis formulation.
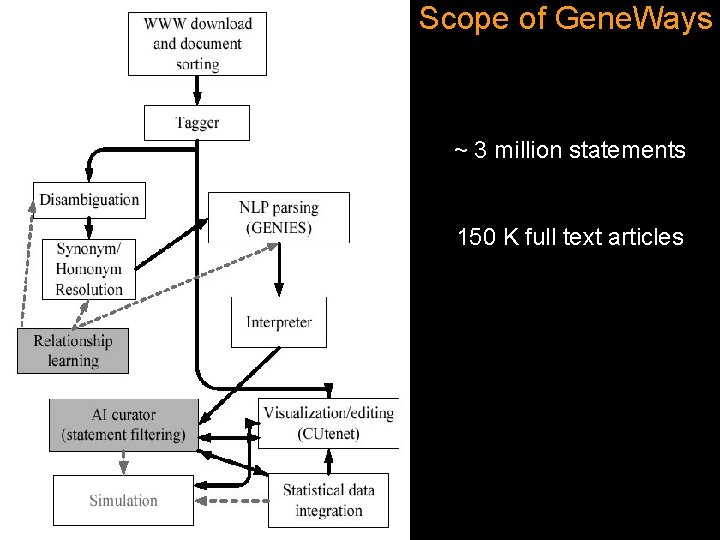
Scope of Gene. Ways ~ 3 million statements 150 K full text articles
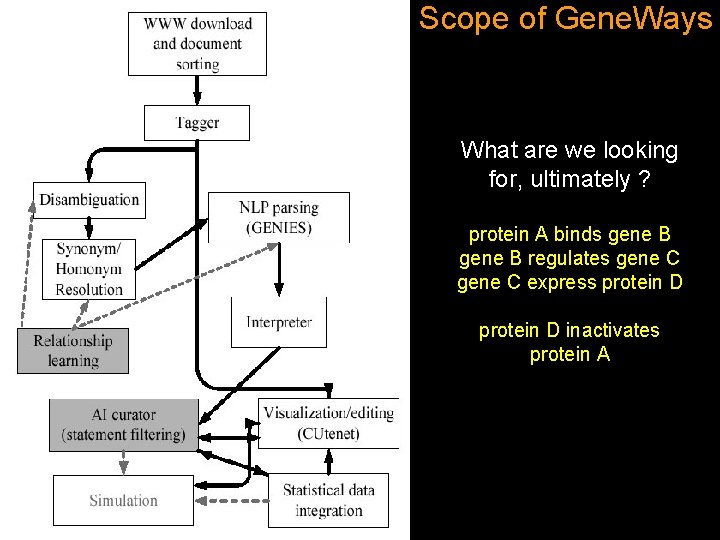
Scope of Gene. Ways What are we looking for, ultimately ? protein A binds gene B regulates gene C express protein D inactivates protein A
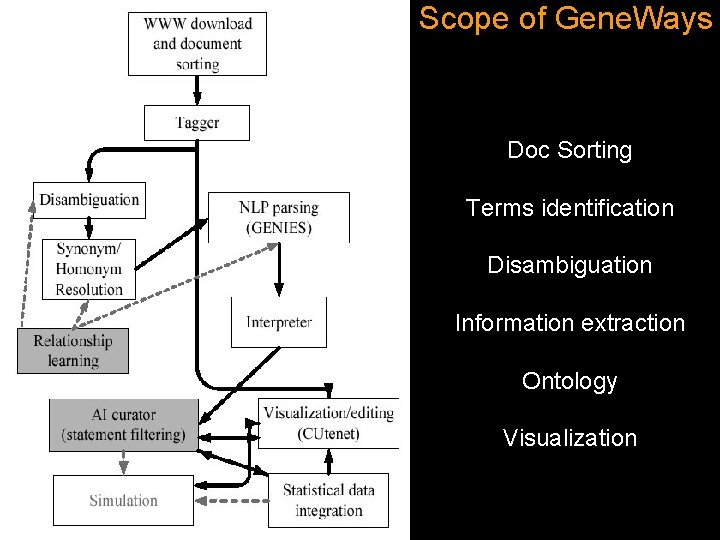
Scope of Gene. Ways Doc Sorting Terms identification Disambiguation Information extraction Ontology Visualization
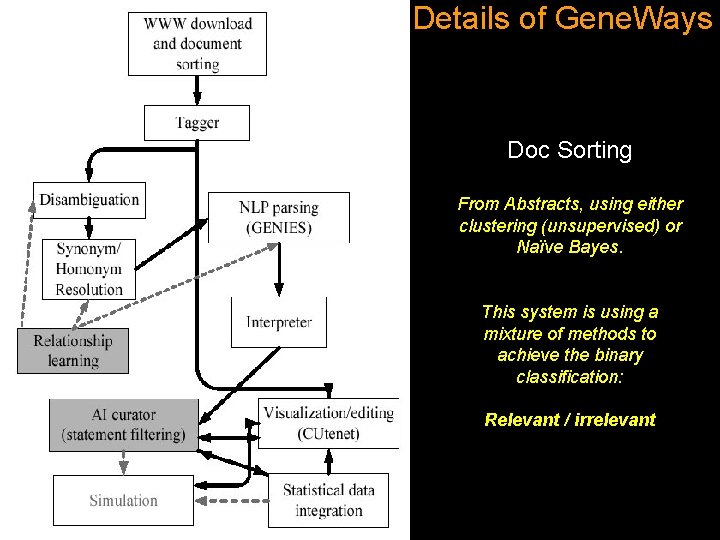
Details of Gene. Ways Doc Sorting From Abstracts, using either clustering (unsupervised) or Naïve Bayes. This system is using a mixture of methods to achieve the binary classification: Relevant / irrelevant
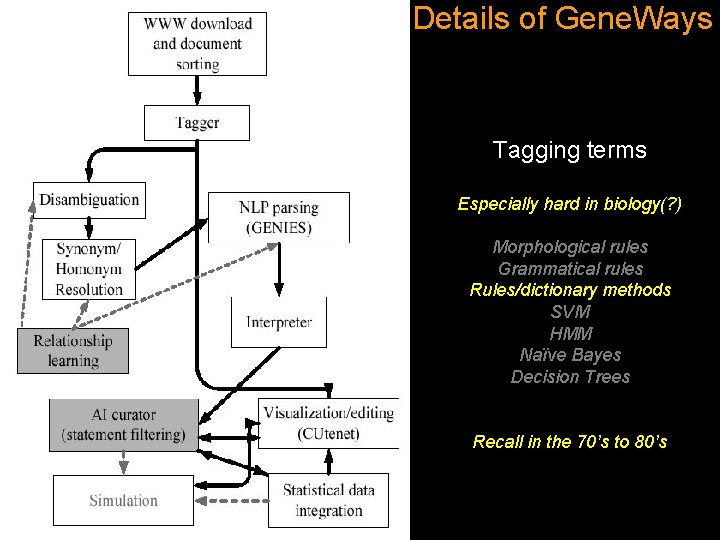
Details of Gene. Ways Tagging terms Especially hard in biology(? ) Morphological rules Grammatical rules Rules/dictionary methods SVM HMM Naïve Bayes Decision Trees Recall in the 70’s to 80’s
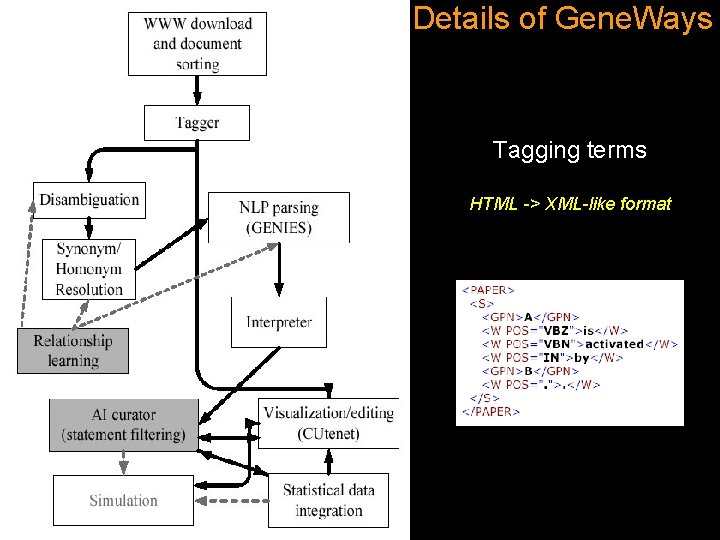
Details of Gene. Ways Tagging terms HTML -> XML-like format
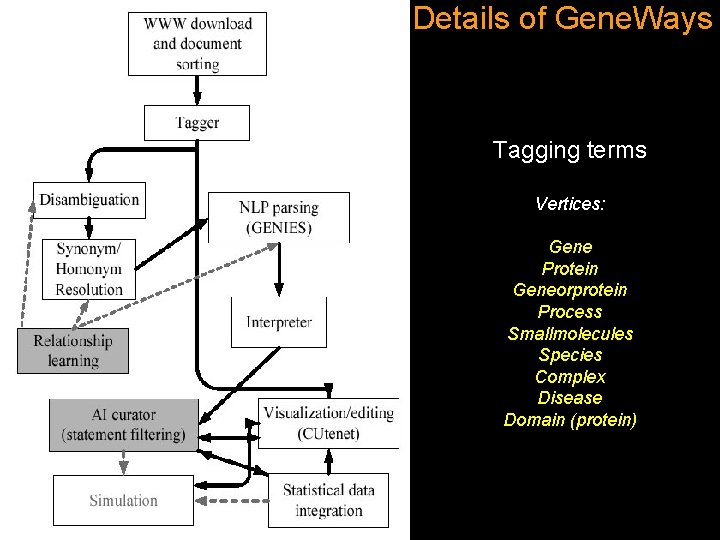
Details of Gene. Ways Tagging terms Vertices: Gene Protein Geneorprotein Process Smallmolecules Species Complex Disease Domain (protein)
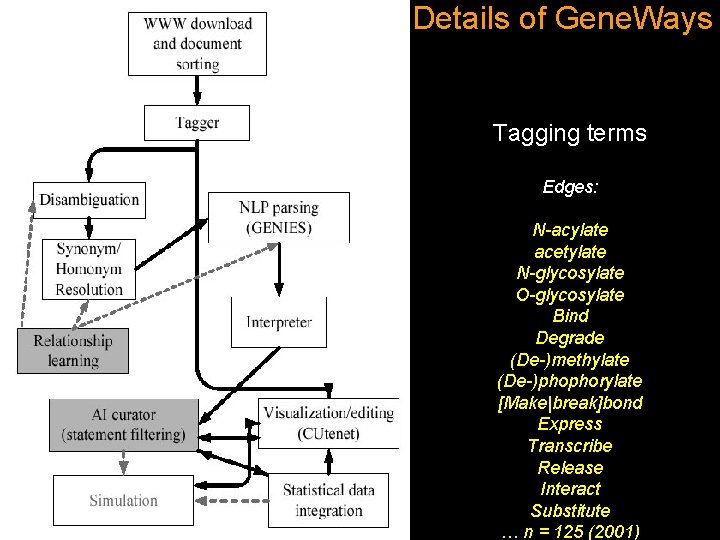
Details of Gene. Ways Tagging terms Edges: N-acylate acetylate N-glycosylate O-glycosylate Bind Degrade (De-)methylate (De-)phophorylate [Make|break]bond Express Transcribe Release Interact Substitute … n = 125 (2001)
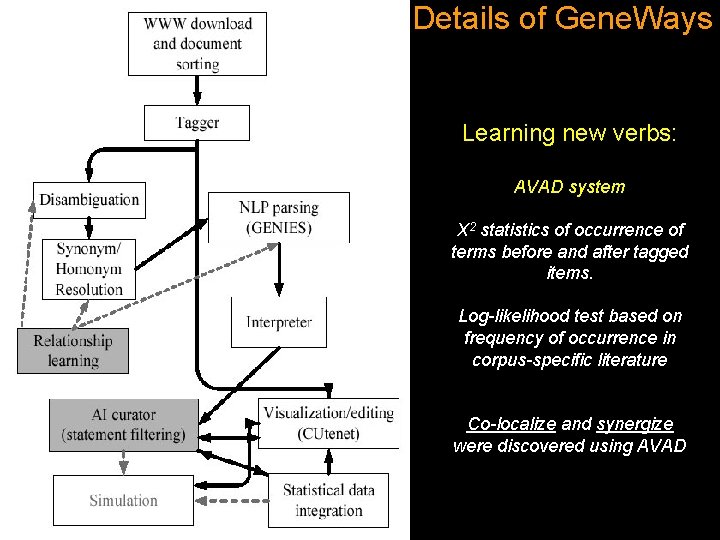
Details of Gene. Ways Learning new verbs: AVAD system Χ 2 statistics of occurrence of terms before and after tagged items. Log-likelihood test based on frequency of occurrence in corpus-specific literature Co-localize and synergize were discovered using AVAD

Nomenclature There are obscure ways to agree: Protein kinase A phosphorylates protein B Is the same as :

Nomenclature There are obscure ways, period: Gene named: “Forever Young” in Arabidopsis Thaliana (mustard familly) “Mother against decapentaplegic” in Fruit fly

Nevermind the jargon! Fight fire with fire: They developed a method that uses BLAST, a popular sequence database search algorithm to mine for biological terms. (Krauthammer et al. , 2000. Gene. 259: 245 -252)

Nevermind the jargon! Fight fire with fire: N-(2 -Hydroxyethyl)piperazine-N'-(2 -ethanesulfonic acid) (HEPES) 2 -(N-Morpholino)ethanesulfonic acid (MES) 3 -(N-Morpholino)propanesulfonic acid (MOPS) N-tris[Hydroxymethyl]methyl-3 -aminopropanesulfonic acid (TAPS) tris(Hydroxymethyl)aminomethane (TRIS)
Details of Gene. Ways Disambiguation il 2 and interleukine-2 can both be used to refer to either the gene, the protein or the m. RNA.
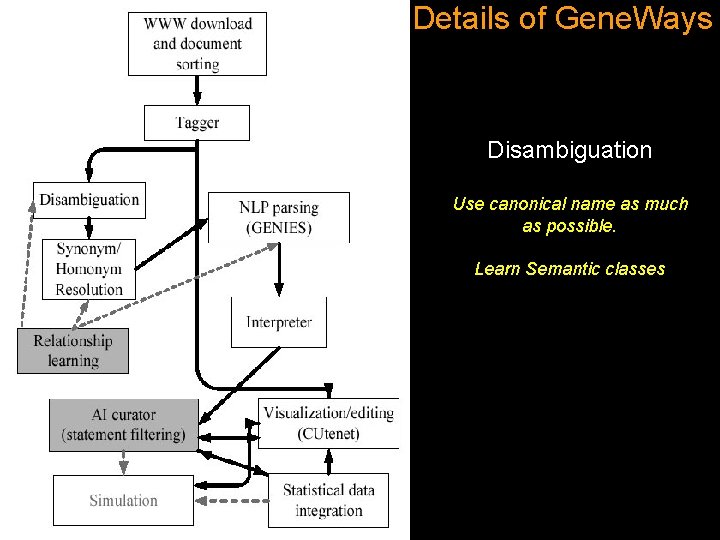
Details of Gene. Ways Disambiguation Use canonical name as much as possible. Learn Semantic classes
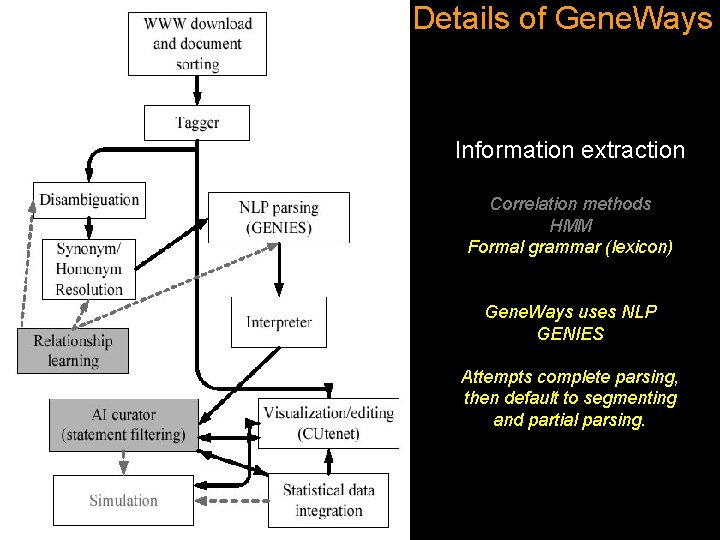
Details of Gene. Ways Information extraction Correlation methods HMM Formal grammar (lexicon) Gene. Ways uses NLP GENIES Attempts complete parsing, then default to segmenting and partial parsing.
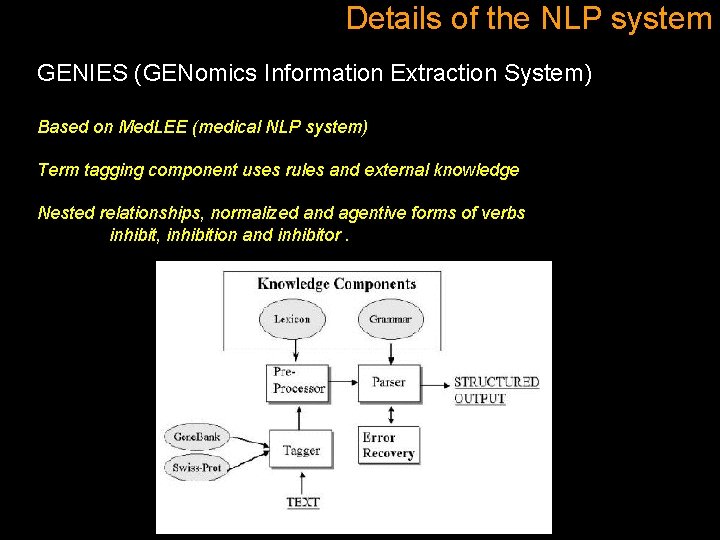
Details of the NLP system GENIES (GENomics Information Extraction System) Based on Med. LEE (medical NLP system) Term tagging component uses rules and external knowledge Nested relationships, normalized and agentive forms of verbs inhibit, inhibition and inhibitor.

Details of Gene. Ways Information simplification Convert nested relationships into a collection of binary statements.
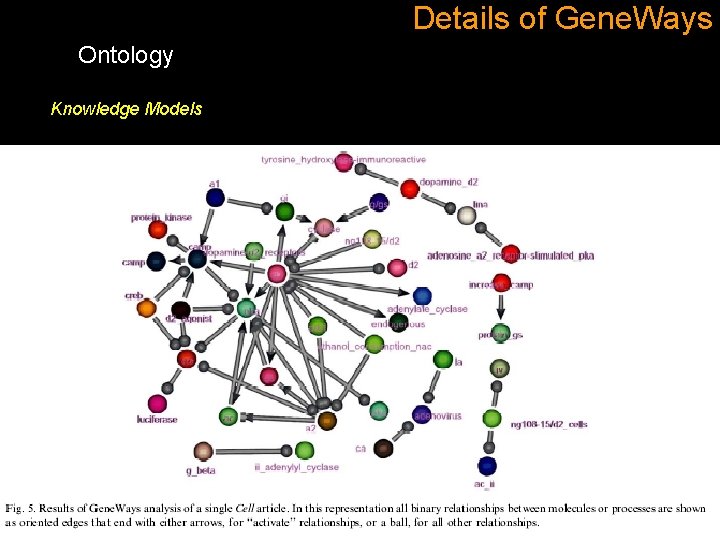
Details of Gene. Ways Ontology Knowledge Models
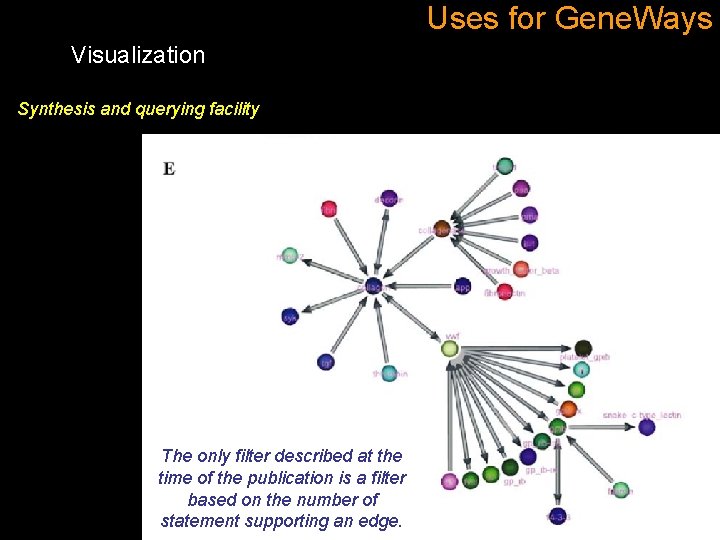
Uses for Gene. Ways Visualization Synthesis and querying facility The only filter described at the time of the publication is a filter based on the number of statement supporting an edge.
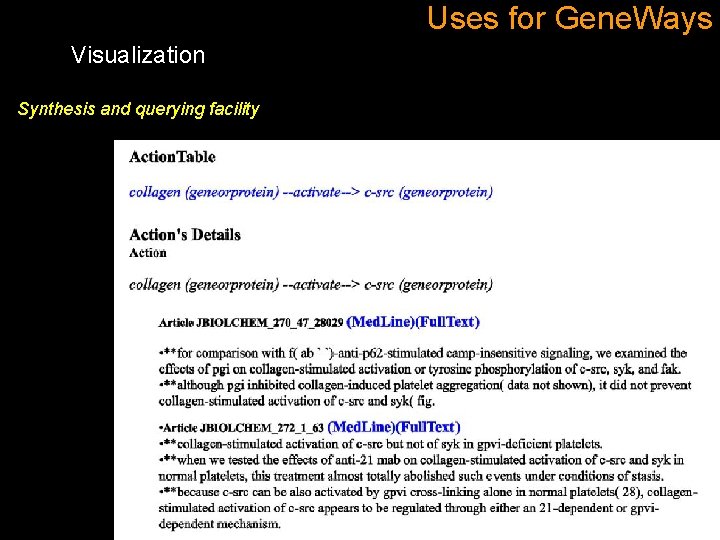
Uses for Gene. Ways Visualization Synthesis and querying facility

Validation of Gene. Ways Expert Review 125 statements / 2500 were erroneous or “phantoms”. Of these 125: - 100 due to term identification. 12 NLP errors. 5 Simplifier errors. 8 Actually correct! System’s precision: 95% Expert’s precision : 93. 5% Such as system should be seen as a mean to enrich

Validation of Gene. Ways Redundancy Redundant statements are not necessarily “more true”. Redundancy due to indirect relationships.

Validation of Gene. Ways A parser’s nightmare: Statement : “mitogen-activated protein kinase (MAPKKK) phosphorylates protein B” Interpretations: 1. 2. 3. 4. Protein kinase [protein] is activated by the mitogen [complex] MAPK[protein] phosphorylate MAPKK[protein] phosphorylate MAPKKK[protein] phosphorylate B [protein] Potential historical artifacts: 1. 2. 3. B[protein] is activated by the mitogen[complex] MAPKK[wrongly thought to be MAPK] phosphorylate B[protein] …

Perspective References Main: Rzhetski et al. , 2004. Gene. Ways: a system for extracting, analysing, visualizing, and integrating molecular pathway data. J. Biomed. Informatics, 37: 43 -53 Learning Verbs: Hatzivassiloglou, V. , Weng, W. Learning Anchor Verbs for Biological Interactions Patterns from published text articles www. cs. columbia. edu/nlp/papers/2002/ hatzivassiloglou_weng_02. pdf NLP processor: Friedman C, Kra P, Yu H, Krauthammer M, Rzhetsky A. 2001. GENIES: a natural-language processing system for the extraction of molecular pathways from journal articles. Bioinformatics, 17: S 74 -S 82 Acknowledgement: Aditya Aggarwal, the student who dug out this paper to present in CSCI 6904 (2004)
- Slides: 52