CSCE 555 Bioinformatics Lecture 9 Gene Finding Comparative

CSCE 555 Bioinformatics Lecture 9 Gene Finding & Comparative genomics HAPPY CHINESE NEW YEAR Meeting: MW 4: 00 PM-5: 15 PM SWGN 2 A 21 Instructor: Dr. Jianjun Hu Course page: http: //www. scigen. org/csce 555 University of South Carolina Department of Computer Science and Engineering 2008 www. cse. sc. edu.

Outline Performance Evaluation of Gene Finding programs Comparative genomics: ◦ What to do ◦ Tools ◦ Databases ◦ Application case 1/4/2022 2

Accuracy Measures of Gene-Finding Programs Sensitivity vs. Specificity (adapted from Burset&Guigo 1996) TP FP TN FN TP FN TN Actual Predicted Actual No Coding / Coding Predicted Coding / No Coding TP FP FN TN ØSensitivity (Sn) Fraction of actual coding regions that are correctly predicted as coding ØSpecificity (Sp) Fraction of the prediction that is actually correct ØCorrelation Coefficient (CC) Combined measure of Sensitivity & Specificity Range: -1 (always wrong) +1 (always right)

Test Datasets Sample Tests reported by Literature ◦ Test on the set of 570 vertebrate gene seqs (Burset&Guigo 1996) as a standard for comparison of gene finding methods. ◦ Test on the set of 195 seqs of human, mouse or rat origin (named HMR 195) (Rogic 2001).
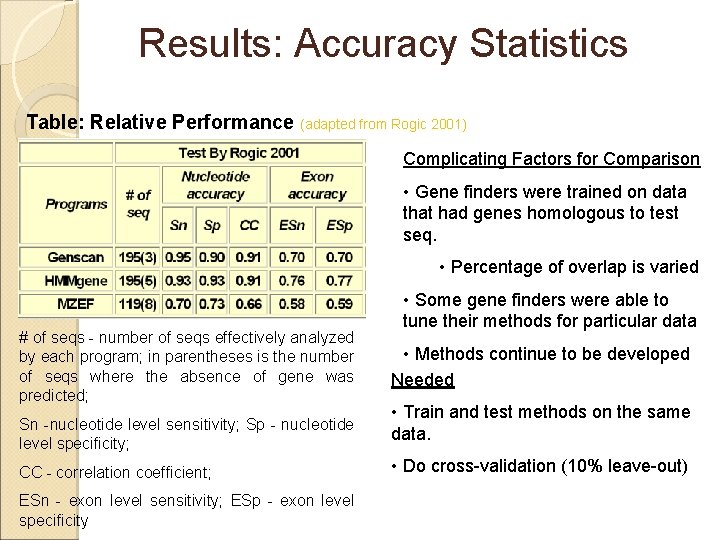
Results: Accuracy Statistics Table: Relative Performance (adapted from Rogic 2001) Complicating Factors for Comparison • Gene finders were trained on data that had genes homologous to test seq. • Percentage of overlap is varied # of seqs - number of seqs effectively analyzed by each program; in parentheses is the number of seqs where the absence of gene was predicted; Sn -nucleotide level sensitivity; Sp - nucleotide level specificity; CC - correlation coefficient; ESn - exon level sensitivity; ESp - exon level specificity • Some gene finders were able to tune their methods for particular data • Methods continue to be developed Needed • Train and test methods on the same data. • Do cross-validation (10% leave-out)
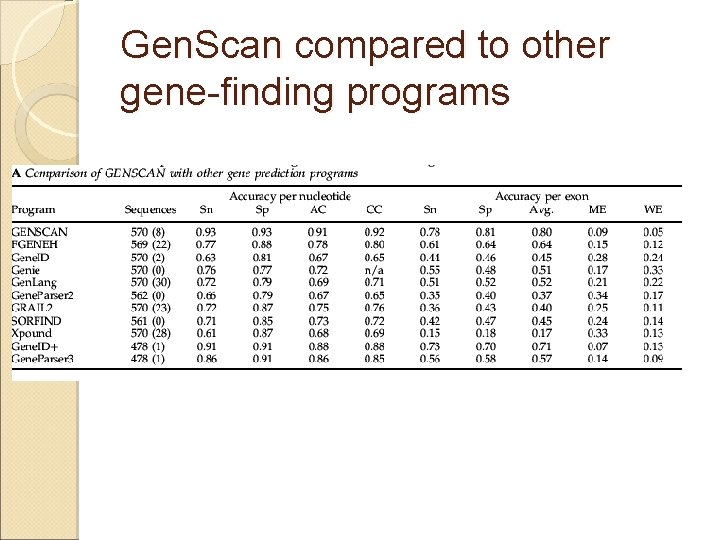
Gen. Scan compared to other gene-finding programs

Why not Perfect? Gene Number usually approximately correct, but may not Organism primarily for human/vertebrate seqs; maybe lower accuracy for non-vertebrates. ‘Glimmer’ & ‘Gene. Mark’ for prokaryotic or yeast seqs Exon and Feature Type Internal exons: predicted more accurately than Initial or Terminal exons; Exons: predicted more accurately than Poly-A or Promoter signals Biases in Test Set (Resulting statistics may not be representative)

Eukaryotic Gene Finding Tools Genscan (ab initio), Genome. Scan (hybrid) (http: //genes. mit. edu/) Twinscan (hybrid) (http: //genes. cs. wustl. edu/) FGENESH (ab initio) (http: //www. softberry. com/berry. phtml? topic=gfind) Gene. Mark. hmm (ab initio) (http: //opal. biology. gatech. edu/Gene. Mark/eukhmm. cgi) MZEF (ab initio) (http: //rulai. cshl. org/tools/genefinder/) Grail. EXP (hybrid) (http: //grail. lsd. ornl. gov/grailexp/) Gene. ID (hybrid) (http: //www 1. imim. es/geneid. html)
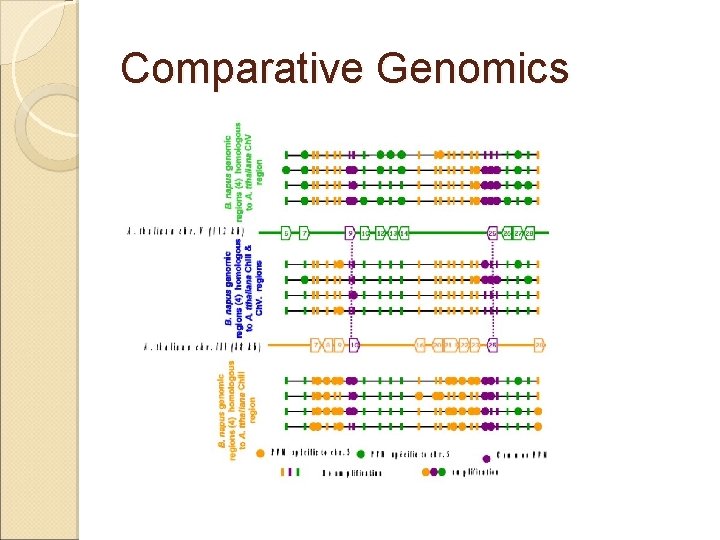
Comparative Genomics

Outline for Comparative Genomics Overview Why do comparative genomic analysis? Assumptions/Limitations Genome Analysis and Annotation Standard Procedure General Purposes Databases for Comparative Genomics Organism Specific Databases Genome Analysis Environments Genome Sequence Alignment Programs Genomic Comparison Visualization Tools

What is comparative genomics? Analyzing & comparing genetic material from different species to study evolution, gene function, and inherited disease Understand the uniqueness between different species

What is compared? Gene location Gene structure ◦ ◦ Exon number Exon lengths Intron lengths Sequence similarity Gene characteristics ◦ Splice sites ◦ Codon usage ◦ Conserved synteny
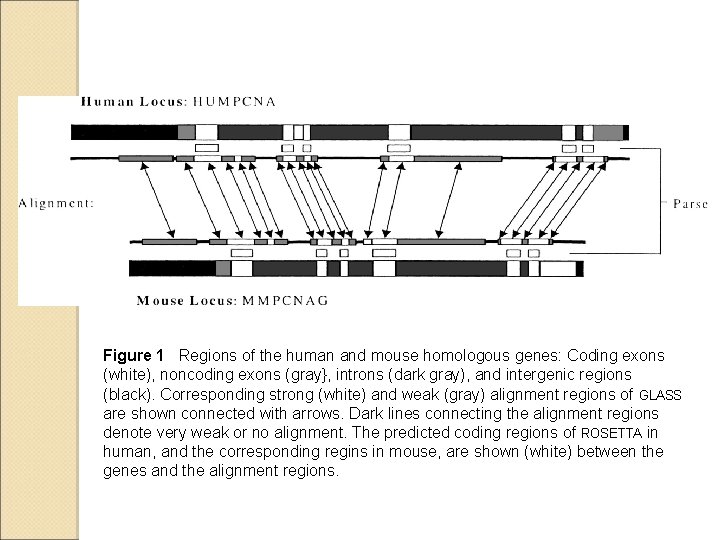
Figure 1 Regions of the human and mouse homologous genes: Coding exons (white), noncoding exons (gray}, introns (dark gray), and intergenic regions (black). Corresponding strong (white) and weak (gray) alignment regions of GLASS are shown connected with arrows. Dark lines connecting the alignment regions denote very weak or no alignment. The predicted coding regions of ROSETTA in human, and the corresponding regins in mouse, are shown (white) between the genes and the alignment regions.

Sequenced prokaryotic genomes Bacteroides fragilis Bordetella bronchiseptica Bordetella parapertussis Bordetella pertussis Burkholderia cepacia Burkholderia pseudomallei Chlamidophila abortus Clostridium botulinum Clostridium difficile Corynebacterium diphtheriae Erwinia carotovora Escherichia/Shigella spp. (5) Mycobacterium bovis Mycobacterium marinum Neisseria meningitidis (serogroup C) Salmonella typhi Salmonella spp. (5) Staphylococcus aureus (MRSA) Staphylococcus aureus (MSSA) Streptococcus pneumoniae Streptococcus pyogenes Streptococcus suis Streptococcus uberis Streptomyces coelicolor Tropheryma whipelli Wolbachia (Culex quinquefasciatus) Wolbachia (Onchocerca volvulus) Yersinia enterocolitica Yersinia pestis Opportunistic Veterinary Whooping cough Lung infections in CF Melliodosis Veterinary Botulism Colitis Diphtheria Plant pathogen Various Tuberculosis Various Bacterial meningitis Typhoid fever Various (Nosocomial) Various (Community acquired) Bacterial meningitis Various (ARF-associated) Veterinary Non-pathogenic Whipple’s disease Vector (Bancroftian filariasis) River Blindness Food poisoning Plague In progress Complete In progress Funded In progress Complete Funded In progress In progress Complete In progress Funded In progress Complete

Sequenced eukaryotic genomes Aspergillus fumigatus Dictyostelium discoideum Entamoeba histolitica Leishmania major Plasmodium falciparum Schistosoma mansoni Schizosaccharomyces pombe Theileria annulata Toxoplasma gondii Trypanosoma brucei Farmer’s lung Soil amoeba Amoebic dysentry Leishmaniasis Malaria Bilharzia Fission yeast Veterinary Toxoplasmosis Sleeping sickness In progress In progress Complete In progress

Bioinformatics Flow Chart 1 a. Sequencing 1 b. Analysis of nucleic acid seq. 6. Gene & Protein expression data 7. Drug screening 2. Analysis of protein seq. 3. Molecular structure prediction Ab initio drug design OR Drug compound screening in database of molecules 4. molecular interaction 8. Genetic variability 5. Metabolic and regulatory networks
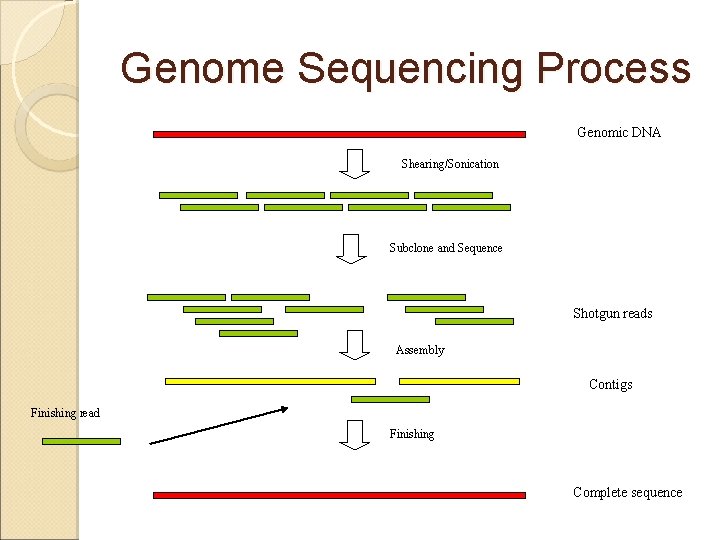
Genome Sequencing Process Genomic DNA Shearing/Sonication Subclone and Sequence Shotgun reads Assembly Contigs Finishing read Finishing Complete sequence

Genome Sequencing - Review Strategy Clone by clone vs whole genome shotgun Libraries Subcloning; generate small insert libraries Sequencing Assembly Closure Annotation Release • Most genome will be sequenced and can be sequenced; few problem are unsolvable. Assembly: Process of taking raw single-pass reads into contiguous • consensus sequence (Phred/Phrap) Problem lies in understanding what you have: Closure: Process of ordering and merging consensus finding sequences into • a. Gene singleprediction/gene contiguous sequence • Annotation -DNA features (repeats/similarities) -Gene finding Release to the public e. g. EMBL or Gen. Bank -Peptidedata features -Initial role assignment -Others- regulatory regions
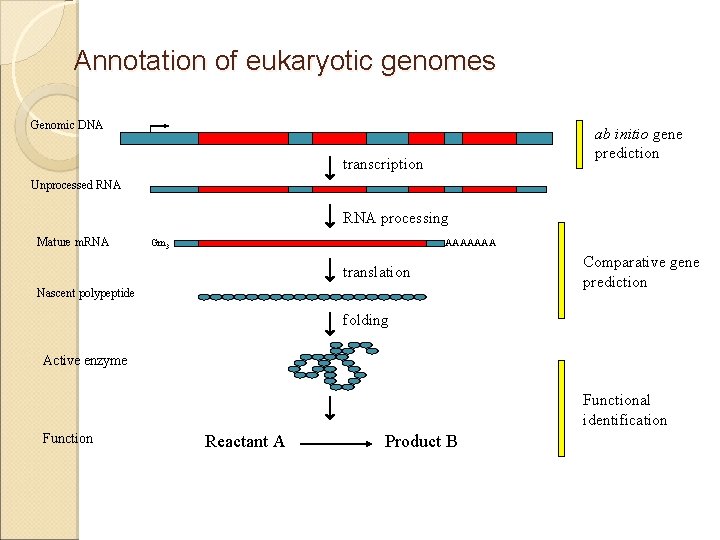
Annotation of eukaryotic genomes Genomic DNA ab initio gene prediction transcription Unprocessed RNA processing Mature m. RNA Gm 3 AAAAAAA translation Nascent polypeptide Comparative gene prediction folding Active enzyme Functional identification Function Reactant A Product B

Why do comparative genomics? Many of the genes encoded in each genome from the genome projects had no known or predictable function Analysis of protein set from completely sequenced genomes Uniform evolutionary conservation of proteins in microbial genomes, 70% of gene products from sequenced genomes have homologs in distant genomes (Koonin et al. , 1997) Function of many of these genes can be predicted by comparing different genomes of known functional annotation and transferring functional annotation of proteins from better studied organisms to their orthologs in lesser studied organisms. Cross species comparison to help reveal conserved coding regions No prior knowledge of the sequence motif is necessary Complement to algorithmic analysis

Assumptions/Limitation Homologous genes are relatively well preserved while noncoding regions tend to show varying degrees of conservation. Conserved noncoding regions are believed to be important in regulating gene expression, maintaiing structural organization of the genome and most likely other possible functions. Cross species comparative genomics is influenced by the evolutionary distance of the compared species.

Genome Analysis and Annotation: General Procedure Basic procedure to determine the functional and structural annotation of uncharacterized proteins: Use a sequence similarity search programs such as BLAST or FASTA to identify all the functional regions in the sequence. If greater sensitivity is required then the Smith-Waterman algorithm based programs are preferred with the trade-off greater analysis time. Identify functional motifs and structural domains by comparing the protein sequence against PROSITE, BLOCKS, SMART, CDD, or Pfam. Predict structural features of the protein such as signal peptides, transmembrane segments, coiled-coil regions, and other regions of low sequence complexity Generate a secondary and tertiary (if possible) structure prediction Annotation: ◦ Transfer of function information from a well-characterized organism to a lesser studied organism and/or ◦ Use phylogenetic patterns (or profiles) and/or ◦ Use the phylogenetic pattern search tools (e. g. through COGs) to perform a systematic formal logical operations (AND, OR, NOT) on gene sets -- differential genome display (Huynen et al. , 1997).

Automated Genome Annotation Gene. Quiz – limited number of searches/day MAGPIE – outside users cannot submit own seq PEDANT – commercial version allow for full capacity SEALS – semi automated

General Databases Useful for Comparative Genomics Locus Link/Ref. Seq: http: //www. ncbi. nih. gov/Locus. Link/ PEDANT -Protein Extraction Description ANalysis Tool http: //pedant. gsf. de/ MIPS – http: //mips. gsf. de/ COGs - Cluster of Orthologous Groups (of proteins) http: //www. ncbi. nih. gov/COG/ KEGG - Kyoto Encyclopedia of Genes and Genomes http: //www. genome. ad. jp/kegg/ MBGD - Microbial Genome Database http: //mbgd. genome. ad. jp/ GOLD - Genome On. Line Database http: //wit. integratedgenomics. com/GOLD/ TOGA – http: //www. tigr. org/xxxxx

Problems with existing sequence alignments algorithms for genomic analysis Most algorithms were developed for comparing single protein sequences or DNA sequences containing a single gene Most algorithms were based on assigning a score to all the possible alignments (usually by the sum of the similarity/identity values for each aligned residue minus a penalty for the introduction of gaps) and then finding the optimal or near-optimal alignment based on the chosen scoring scheme. Unfortunately, most of these programs cannot accurately handle long alignments. Linear-space type of Smith-Waterman variants are too computationally intensive requiring specialized hardware (memory-limited) or very time-consuming. Higher speed vs increased sensitivity.

Genome-size comparative alignment tools ASSIRC - Accelerated Search for SImilarity Regions in Chromosomes ◦ ftp: //ftp. biologie. ens. fr/pub/molbio/ (Vincens et al. 1998) BLAT – ◦ http: //genome. ucsc. edu/cgi-bin/hg. Blat? command=start (Kent xxx) DIALIGN - DIagonal ALIGNment ◦ http: //www. gsf. de/biodv/dialign. html (Morgenstern et al. 1998; Morgenstern 1999( DBA - DNA Block Aligner ◦ http: //www. sanger. ac. uk/Software/Wise 2/dba. shtml (Jareborg et al. 1999( GLASS - GLobal Alignment Sy. Stem ◦ http: //plover. lcs. mit. edu/ (Batzoglou et al. 2000) LSH-ALL-PAIRS - Locality -Sensitve Hashing in ALL PAIRS ◦ Email: jbuhler@cs. washington. edu (Buhler 2001) Mega. Blast ◦ http: //www. ncbi. nih. gov/blast/ (Zhang 2000) MUMmer - Maximal Unique Match (mer) ◦ http: //www. tigr. org/softlab/ (Delcher et al. 1999) PIPMaker - Percent Identity Plot MAKER ◦ http: //biocse. psu. edu/pipmaker/ (Schwartz et al. 2000) SSAHA – Sequence Search and Alignment by Hashing Algorithm ◦ http: //www. sanger. ac. uk/Software/analysis/SSAHA/ WABA - Wobble Aware Bulk Aligner ◦ http: //www. cse. ucsc. edu/~kent/xeno. Ali/ (Kent & Zahler 2000)

SSAHA Sequence Search and Alignment by Hashing Algorithm Software tool for very fast matching and alignment of DNA sequences. Achieves fast search speed by converting sequence information into a hash table data structure which can then be searched very rapidly for matches http: //www. sanger. ac. uk/Software/analysis/SSAHA/ Run from the Unix command line Need > 1 GB RAM (needs a lot of memory) SSAHA algorithm best for application requiring exact or “almost exact” matches between two sequences – e. g. SNP detection, fast sequence assembly, ordering and orientation of contigs

Genome Analysis Environment MAGPIE - Automated Genome Project Investigation Environment PEDANT SEALS

Problems with Visualizing Genomes Alignment programs output often were visualized by text file, which can be intuitively difficult to interpret when comparing genomes. Visualization tools needed to handle the complexity and volume of data and present the information in a comprehensive and comprehensible manner to a biologist for interpretation. Genome Alignment Visualization tools need to provide: ◦ interpretable alignments, ◦ gene prediction and database homologies from different sources ◦ Interactive features: real time capabilities, zooming, searching specific regions of homologies ◦ Represent breaks in synteny ◦ Multiple alignments display ◦ Displaying contigs of unfinished genomes with finished genomes ◦ Handle various data formats ◦ Software availabilty (no black box)

Genome Comparison Visualization Tool ACT - Artemis Comparison Tool (displays parsed BLAST alignments; based on Artemis – an annotation tool) ◦ http: //www. sanger. ac. uk/Software/ACT/ Alfresco (displays DBA alignments and. . . ) ◦ http: //www. sanger. ac. uk/Software/Alfresco/ (Jareborg & Durbin 2000) Pip. Maker (displays Blast. Z alignments) ◦ http: //bio. cse. psu. edu/pipmaker/ (Schwartz et al. 2000) Enteric/Menteric/Maj (displays Blastz alignments) ◦ http: //glovin. cse. psu. edu/enterix/ (Florea et al. 2000; Mc. Clelland et al. 2000) Intronerator (displays WABA alignments and. . . ) ◦ http: //www. cse. ucsc. edu/~kent/intronerator/ (Kent & Zahler 2000 b) VISTA (Visualization Tool for Alignment) (displays GLASS alignments) ◦ http: //www-gsd. lbl. gov/vista/ Syn. Plot (displays DIALIGN and GLASS alignments) ◦ http: //www. sanger. ac. uk/Users/igrg/Syn. Plot/

Artemis Comparison Tool (ACT) - ACT is a DNA sequence comparison viewer based on Artemis Can read complete EMBL and Gen. Bank entries or sequence in FASTA or raw format Additional sequence feature can be in EMBL, Gen. Bank, GFF format ACT is free software and is distributed under the GNU Public License Java based software Latest release 2. 0 better support Eukaryotic Genome Comparison http: //www. sanger. ac. uk/Software/ACT/
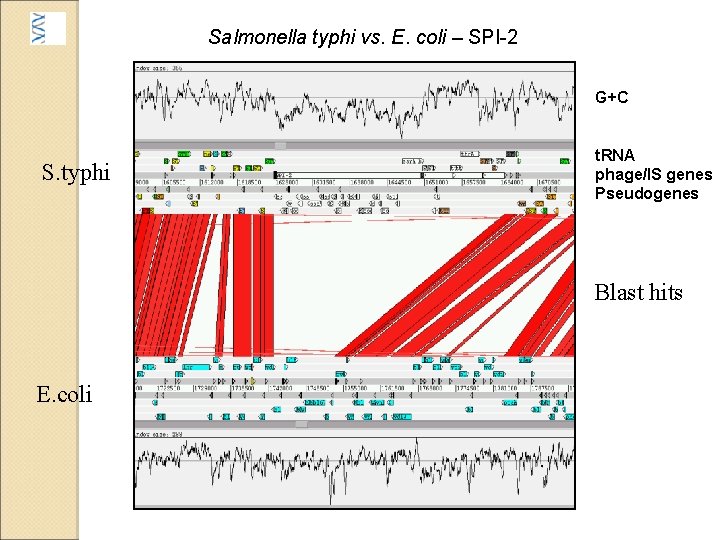
Salmonella typhi vs. E. coli – SPI-2 G+C S. typhi t. RNA phage/IS genes Pseudogenes Blast hits E. coli
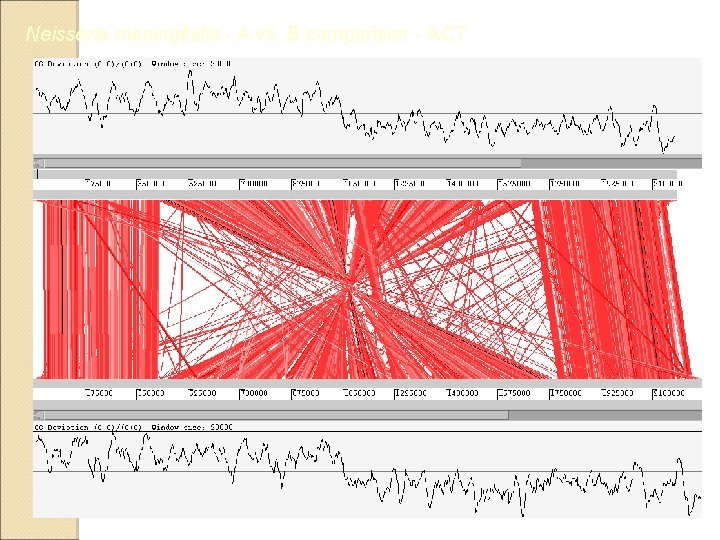
Neisseria meningitidis - A vs. B comparison - ACT
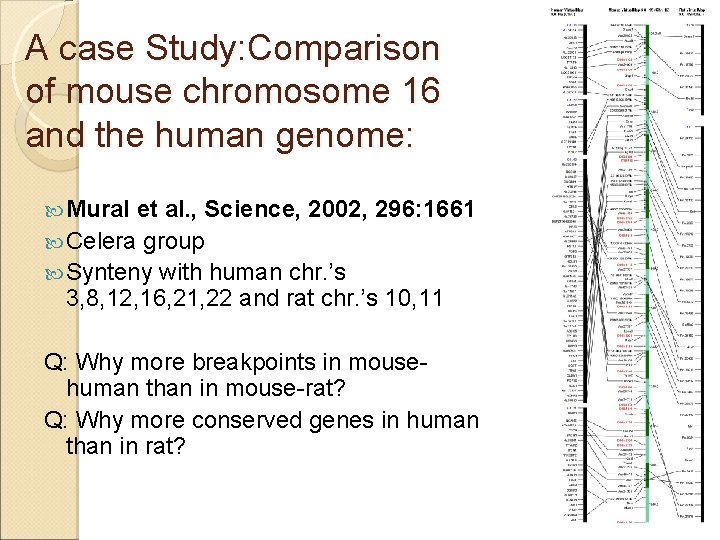
A case Study: Comparison of mouse chromosome 16 and the human genome: Mural et al. , Science, 2002, 296: 1661 Celera group Synteny with human chr. ’s 3, 8, 12, 16, 21, 22 and rat chr. ’s 10, 11 Q: Why more breakpoints in mousehuman than in mouse-rat? Q: Why more conserved genes in human than in rat?
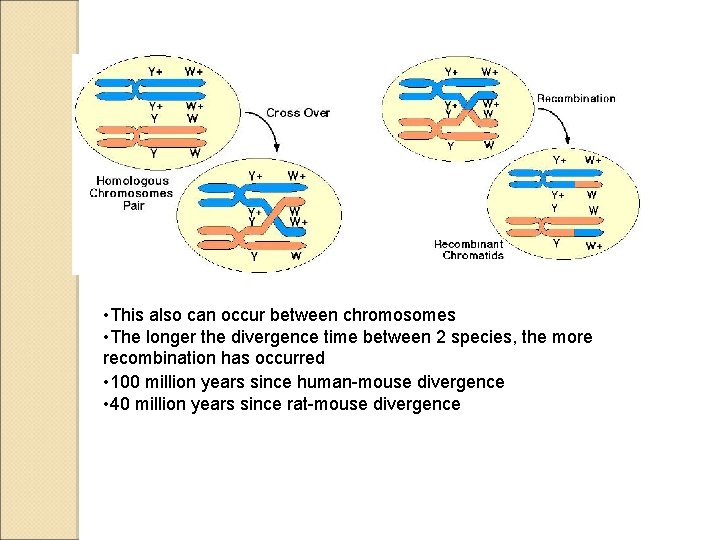
• This also can occur between chromosomes • The longer the divergence time between 2 species, the more recombination has occurred • 100 million years since human-mouse divergence • 40 million years since rat-mouse divergence

Whole-genome shotgun sequencing: 1. Genome is cut into small sections 2. Each section is hundreds or a few thousand bp of DNA 3. Each section is sequenced and put in a database 4. A computer aligns all sequences together (millions of them from each chromosome) to form contigs 5. Contigs are arranged (using markers, etc) to form scaffolds Q: What are the advantages of this over the traditional method? Q: What are the potential sources of error?

1. Assembly of Mmu 16 1. 2. 3. 4. 5. 6. Total size: 99 Mbp Not one contiguous sequence (contig) 8, 635 contigs on 20 “scaffolds” Average scaffold size: 10 Mbp Number of gaps: 8615 Total size of gaps: ~6 Mbp Total coverage: ~93 Mbp

2. Identify genes in Mmu 16 1. 2. 3. 4. 5. Scaffolds of >10 kbp were examined (scaffolds larger than 1 Mbp were chopped) Regions with repeat motifs were ignored using Repeat. Masker Several gene prediction engines use (Gen. Scan, Grail, Fgenes) Amino acid sequences from open reading frames searched against nr protein db (NCBI) Nucleotide searchers (using DNA from across scaffolds) performed against: 1. 2. 3. 4. 5. Celera’s gene clusters Mmu, Rno, & Hsa EST db’s NCBI’s Ref. Seq m. RNA db Celera’s dog genomic db Public pufferfish genomic db

2. Identify genes in Mmu 16 6. 7. 8. 9. 1055 genes with high & medium confidence were predicted Other efforts have identified 1142 genes After visual annotation inspection, psuedogenes and annotation errors removed, leaving 731 homologues genes The genes found were mostly orthologues because they were reciprocal best matches by BLAST searches.

3. Identify regions of conserved synteny between Mmu 16 and Hsa 1. 2. Regions of conserved synteny predicted by sequence similarity and by protein comparisons Synteny based on sequence comparisons: Syntenic anchors were located - regions with high (80%) similarity over short distances (~200 bp or more). Average distance between anchors is 8 kbp, but there are gaps as large as 707 kbp in the mouse and 3. 4 Mbp in the human

3. Identify regions of conserved synteny between Mmu 16 and Hsa 3. 4. 5. 56% of anchors were in mouse genes - exons mostly 44% in intergenic regions Relatively density is independent of coding/noncoding - making the anchors an important marker of synteny (in addition to genes) Human chr. 16 8 12 22 3 q 27 -29 3 q 11. 1 -13. 3 21 Mmu len. 10, 461 1, 284 363 2, 081 13, 557 41, 660 22, 327 Hsa len. 12, 329 1, 491 306 2, 273 16, 461 46, 493 28, 421 No. anchors 1, 429 121 31 418 1, 714 5, 485 2, 127 bad anch. (% incon. ) 21 (1. 5) 1 (0. 8) 3 (9. 7) 8 (1. 9) 18 (1. 0) 63 (1. 1) 27 (1. 3) Orthologues 87 6 3 30 107 165 111

Summary Performance evaluation of gene-finding programs Comparative genomics analysis example

Acknowledgement Chuong Huynh (NIH)
- Slides: 43