CSCE 555 Bioinformatics Lecture 4 Sequence Alignment part

CSCE 555 Bioinformatics Lecture 4 Sequence Alignment (part 1) Meeting: MW 4: 00 PM-5: 15 PM SWGN 2 A 21 Instructor: Dr. Jianjun Hu Course page: http: //www. scigen. org/csce 555 University of South Carolina Department of Computer Science and Engineering 2008 www. cse. sc. edu.

Roadmap Sequence analysis applications Sequence alignment application cases Pairewise sequence alignment Summary 12/17/2021 2

180 Genomes Sequenced What does this mean? What we can do with all these sequence information?

Typical Sequence Analysis Tasks Comparison of sequences in order to find similar sequences (sequence alignment) Identification of gene-structures, reading frames, distributions of introns and exons and regulatory elements Prediction of protein structures genome mapping Comparison of homologous sequences to construct a molecular

Applications of Sequence Alignment Prediction of functions of (gene/protein/promoters) homology Database search ◦ Find similar sequences that are similar to our query sequence (e. g. new gene) Gene finding by genome comparison Sequence divergence/phylogeny Sequence Assembly

What Biological Sequence Similarity Tells Us Homology is an evolutionary relationship that either exists or does not. It cannot be partial. An ortholog is a homolog with shared function. A paralog is a homolog that arose through a gene duplication event. Paralogs often have divergent function. Similarity is a measure of the quality of alignment between two sequences. High similarity is evidence for homology. Similar sequences may be orthologs or paralogs. Timothy Bailey 6

What is Sequence Alignment? Given two sequences, how to measure their similarity? ATAACTTTAA ATCC-TTTACTAA ATAACTTTAA ATCC-TTTAC-TAA
What is Sequence Alignment? Arranging the primary sequences of DNA, RNA, or protein to identify regions of similarity that may be a consequence of functional, structural, or evolutionary relationships between the sequences

What is Sequence Alignment? sequence alignment of instances of the acidic ribosomal protein P 0 (L 10 E) from several organisms

Tasks of Sequence Alignment Pairwise alignment Multiple sequence alignment Global alignment Local assignment Approximate alignment algorithms versus optimal/exact alignment algorithms
Web Sequence Alignment Tool

Web Tool to Align Two Sequences http: //www. ebi. ac. uk/emboss/align

Multiple Sequence Alignment
Sequence Blasting on Web

Roadmap Sequence analysis applications Sequence alignment application cases Pairwise Sequence Alignment Summary 15

Pairwise Sequence Alignment Pairwise sequence alignment primary tool in sequence analysis, used for database search and as a component of other algorithms. What? To detect similarity Why? ◦ ◦ Infer phylogeny (similarity ~ distance) Predict function Predict structure Predict “signals” (binding sites, splicing signals etc. 16

Problem Definition: (Optimal) pairwise alignment consists of considering all possible alignments of two sequences and choosing the optimal one. Sub-optimal (heuristic) alignment algorithms are also very important: eg BLAST We will focus on optimal alignment methods in this class.

Key Issues The key issues are: Types of alignments (local vs. global) The scoring system The alignment algorithm Measuring alignment significance

Example Alignment True homology: alpha globin and beta globin HBA_HUMAN GSAQVKGHGKKVADALTNAVAHVDDMPNALSALSDLHAHKL G+ +VK+HGKKV HBB_HUMAN A+++++AH+D++ +++++LS+LH KL GNPKVKAHGKKVLGAFSDGLAHLDNLKGTFATLSELHCDKL Spurious “homology”: alpha globin protein with different strucure and function HBA_HUMAN LHAHKL GSAQVKGHGKKVADALTNAVAHVDDMPNALSALSD---GS+ + G + +D L ++ H+ D+ ++AH+ F 11 G 11. 2 GSGYLVGDSLTFVDLL-VAQHTADLLAANAALLDEFPQFKAHQE 19 A +AL D

Types of Alignments Global—sequences aligned from end -to-end. Local—alignments may start in the middle of either sequence Ungapped—no insertions or deletions are allowed Other types: overlap alignments, repeated match alignments 20

Local vs. Global Pairwise Alignments A global alignment includes all elements of the sequences and includes gaps. ◦ A global alignment may or may not include "end gap" penalties. ◦ Global alignments are better indicators of homology and take longer to compute. A local alignment includes only subsequences, and sometimes is computed without gaps. ◦ Local alignments can find shared domains in divergent proteins and are fast to compute 21

Scoring systems Match scores: s(a, b) assigns a score to each combination of aligned letters. Examples: transition/transversion, PAM, Blosum. Gap score: f(g) assigns a score to a gap of length g. Examples: linear, affine. Scoring systems are usually additive: the total score is the sum of the substitution scores and all the gap scores. 22

Match Scores Some amino acids are more “substitutable” for each other than others. Serine and threonine are more alike than tryptophan and alanine. We can introduce "mismatch costs" for handling different substitutions. We don't usually use mismatch costs in aligning nucleotide sequences, since often no substitution is per se better than any other. 23

Match Score Tables Match scores are computed using a lookup table with an entry for each possible pair of letters. Match scores are often calculated on the basis of the frequency of particular mutations in very similar sequences. 24

Log-odds Match Scores Match scores are usually log-odds scores. s(a, b) = log(Pr(a, b | M) / Pr(a, b | R)) b | M) = pab is the probability of the residues a and b appearing assuming the correct positions are aligned and the sequences descended from a common ancestor. Pr(a, b | R) is the probability of a and b if we pick two random sequence positions to align. Assuming independence (null model) gives Pr(a, b | R)=qaqb. Pr(a, 25

Adding log-odds substitution scores gives the log-odds of the alignment For ungapped alignments, the logodds alignment score of two sequence segments x and y is equal to the log-odds ratio of the segments. Let x = x 1 x 2. . . xn and y = y 1 y 2. . . yn. 26

Gap Scores Gap scores are negative (they are “costs”) Linear: gap score is proportional to length of gap. f(g) = -g · d Affine: gap score is a constant plus a linear score. With affine scores, many small gaps cost more than one large gap. f(g) = -d -(g-1)e, for g>0 27
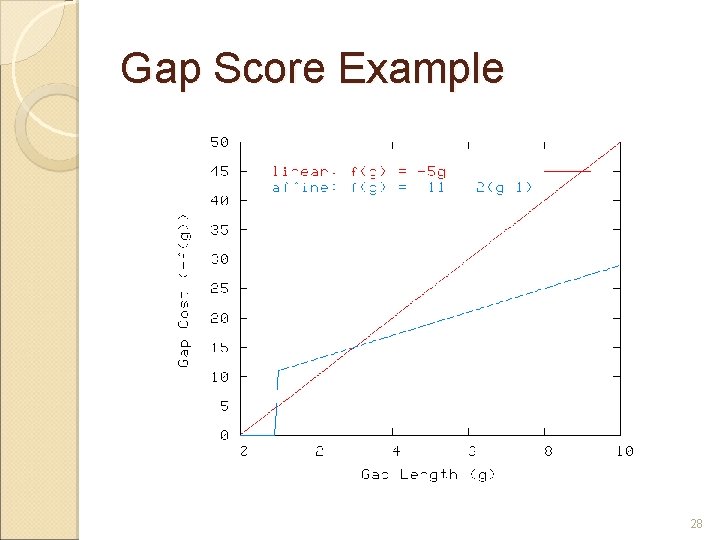
Gap Score Example 28

Summary Sequence alignment application cases Pairwise sequence alignment
- Slides: 29