CSCE 555 Bioinformatics Lecture 2 Meeting MW 4

CSCE 555 Bioinformatics Lecture 2 Meeting: MW 4: 00 PM-5: 15 PM SWGN 2 A 21 Instructor: Dr. Jianjun Hu Course page: http: //www. scigen. org/csce 555 University of South Carolina Department of Computer Science and Engineering 2008 www. cse. sc. edu.

Roadmap � DNA, Chromosomes, Genomes � Genome � DNA Sequencing and whole genomes Sequence Representation, Models � Sequence � Basic Retrieval, Manipulation Analysis and Questions of Genomes � Summary 2/14/2022 2

Tools to Learn Concepts Quickly �Wikipedia. org ◦ Search “Genome” bringing up many related information ◦ In google, type “keywards wiki” �Google search tips ◦ Find info from university websites �Genome, site: edu ◦ Find info as powerpoint files �Genome, tutorial, filetype: ppt
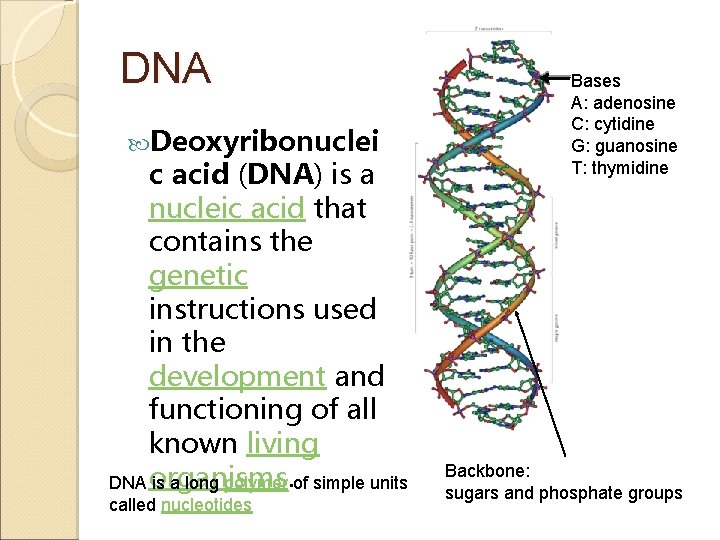
DNA Deoxyribonuclei c acid (DNA) is a nucleic acid that contains the genetic instructions used in the development and functioning of all known living DNA organisms. is a long polymer of simple units called nucleotides Bases A: adenosine C: cytidine G: guanosine T: thymidine Backbone: sugars and phosphate groups
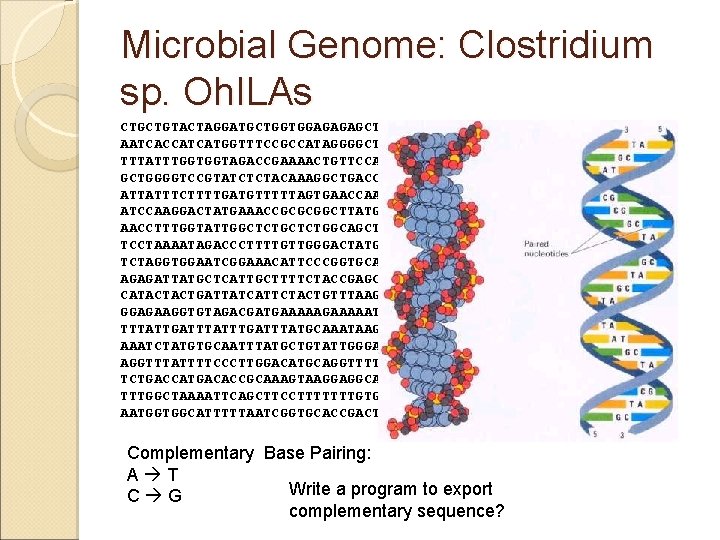
Microbial Genome: Clostridium sp. Oh. ILAs CTGCTGTACTAGGATGCTGGTGGAGAGAGCTGCATATAAATCTTTGAGAGATGCACCAAG AATCACCATCATGGTTTCCGCCATAGGGGCTTCTTTTTTTATTCAAAATCTTGCCATTGT TTTATTTGGTGGTAGACCGAAAACTGTTCCAACGGTGGAGGTATTGTCCGGGGTGATAAA GCTGGGGTCCGTATCTCTACAAAGGCTGACCTTAGTGATTCCAGTAGTAACCATACTGCT ATTATTTCTTTTGATGTTTTTAGTGAACCAAACGAAAACTGGAATGGCAATGCGTGCCGT ATCCAAGGACTATGAAACCGCGCGGCTTATGGGAATTGACGTCAATAAAATTATTACCAT AACCTTTGGTATTGGCTCTGGCAGCTATTGGTGGCATCATGTGGGGCGCAAAATT TCCTAAAATAGACCCTTTTGTTGGGACTATGCCGGGTATTAAATGCTTTATTGCTGCAGT TCTAGGTGGAATCGGAAACATTCCCGGTGCAGTAATCGGGGGGTTCATCTTAGGGATTGG AGAGATTATGCTCATTGCTTTTCTACCGAGCCTAACTGGCTATCGAGATGCCTTTGCTTT CATACTACTGATTATCATTCTACTGTTTAAGCCAACAGGAATCATGGGTGAAAAAATTGC GGAGAAGGTGTAGACGATGAAAAATACCATATTAACTGGATTAGCAGTATTGCT TTTATTGATTTATGCAAATAAGAATTATGATTCTTATAAAATTAGAGTTCT AAATCTATGTGCAATTTATGCTGTATTGGGACTCAGTATGAATTTGATCAATGGATTTAC AGGTTTATTTTCCCTTGGACATGCAGGTTTTATGGCAGTAGGTGCCTATACTACCGCTCT TCTGACCATGACACCGCAAAGTAAGGAGGCAACATTCTTCTTAGTGCCCATTGTAGAGCC TTTGGCTAAAATTCAGCTTCCTTTTTTTGTGGCACTGATCATCGGTGGACTACTTTCAGC AATGGTGGCATTTTTAATCGGTGCACCGACTTTAAGGCTGAAGGGCGATTATTTAGCCAT Complementary Base Pairing: A T Write a program to export C G complementary sequence?
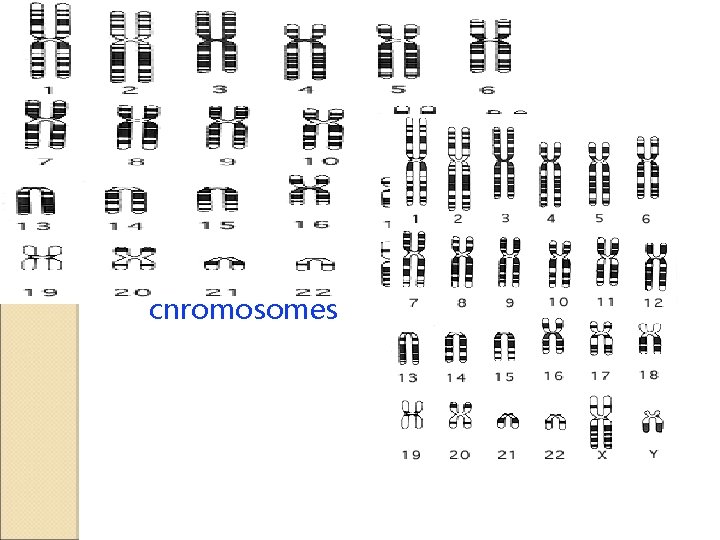
Genome of organisms genome of an organism is a complete DNA sequence of one set of chromosomes

Sequencing: Basic Ideas � Current lab techniques can sequence small (say 700 base pairs) DNA pieces. ◦ Use restriction enzymes to cut DNA pieces ◦ Sort pieces of different sizes using gel electrophoresis and use the sorting to read them � Mapping and Walking ◦ Sequence one piece, get 700 letters, make a primer that allowed you to read the next 700, and work sequentially down the clone ◦ Estimate for human genome sequencing using this method: 100 years � Shotgun sequencing (introduced by Sanger et al. 1977) for sequencing genomes ◦ Obtain random sequence reads from a genome ◦ Assemble them into contigs on the basis of sequence overlaps � Straightforward for simple genomes (with no or few repeat sequences) � Merge reads containing overlapping sequence � Shotgun sequencing is more challenging for complex (repeat-rich) genomes: two approaches

How Sequencing Works Beckman CEQ 8000

Sequencing small DNA pieces G G A T ------- A ------- C -------------- T ------- A G -------------- T ------- A ------- C ------- A -------------- G -------------- A -------------- C ------- T -------------- A G C Use DNA cloning or PCR to make multiple copies. Put in 4 testtubes marked G, A, T and C In testtube G use restriction enzymes that cuts at G. Do the above step for the other testubes. Use gel electrophoresis separately for the content in each testtube. The data results in the table on the left. Reading the table we get G has lengths 1, 7, 12, 13, 19; A has lengths 2, 6, 8, 11, 14, 15, 16; T has length 4, 5, 9, 18 and C has length 3, 10, 17. This gives us the sequence.

Methods for very large scale sequencing �A hierarchical approach ◦ Map on a large scale (physical mapping), sequence specific clones whose position in the genome is known �Shot gun sequencing ◦ “Tear up” the genome and sequence random fragments until it is done �Sequence tagged connectors (STC) ◦ Sequence the ends of many clones and use this info to pick overlapping clones
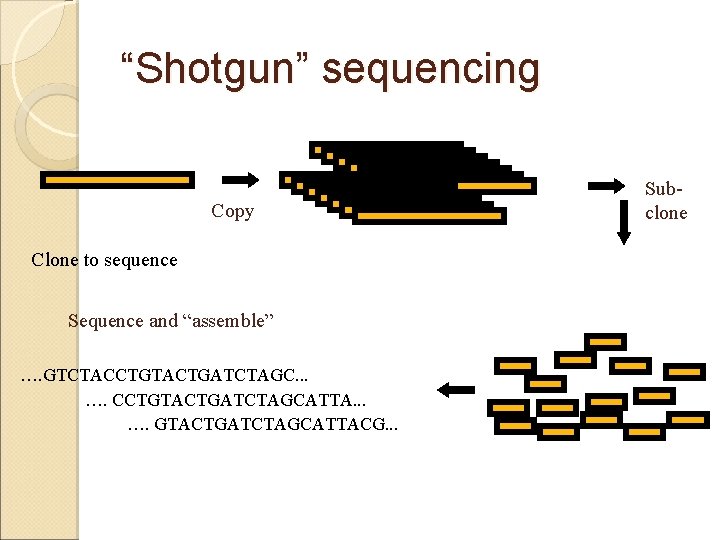
“Shotgun” sequencing Copy Clone to sequence Sequence and “assemble” …. GTCTACCTGTACTGATCTAGC. . . …. CCTGTACTGATCTAGCATTA. . . …. GTACTGATCTAGCATTACG. . . Subclone

Emerging Sequence Methods Sequencing by Hybridization (SBH). Mass Spectrophotometric Sequences. Direct Visualization of Single DNA Molecules by Atomic force Microscopy (AFM ) Single Molecule Sequencing Techniques Single nucleotide Cutting Nanopore sequencing Readout of Cellular Gene Expression

Whole Genomes of Species �Bacterial Genomes �Eukaryotic Genomes �Human Genome Project �Other Animal and Plant Genomes �Model Genomes The genomes of more than 180 organisms have been sequenced since 1995 http: //www. genomenewsnetwork. org/resou rces/sequenced_genomes/genome_guide_ p 1. shtml

Sizes of Genomes You will learn to download all these genomes into your computer’s harddrive Refer to Table 1. 1 Page 2 of Intro to Comp Genomics book.

Roadmap � DNA, Chromosomes, Genomes � Genome � DNA Sequencing and whole genomes Sequence Representation, Models � Sequence � Basic Retrieval, Manipulation Analysis and Questions of Genomes � Summary 2/14/2022 15

DNA Sequence Representation �DNA Sequence: a string of letters with alphabet {A, C, G, T} �Protein sequence: a string of amino acids with alphabet {ARNDCEQGHILKMFPSTWYV} ◦ 20 standard amino acids �Genetic code:
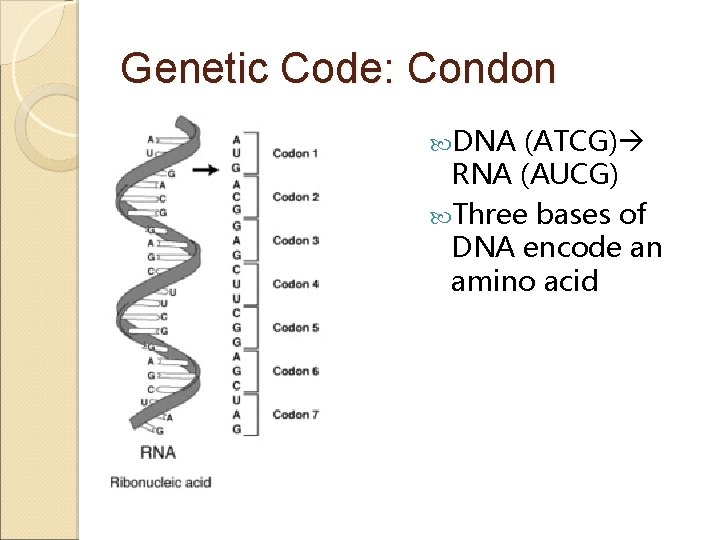
Genetic Code: Condon DNA (ATCG) RNA (AUCG) Three bases of DNA encode an amino acid
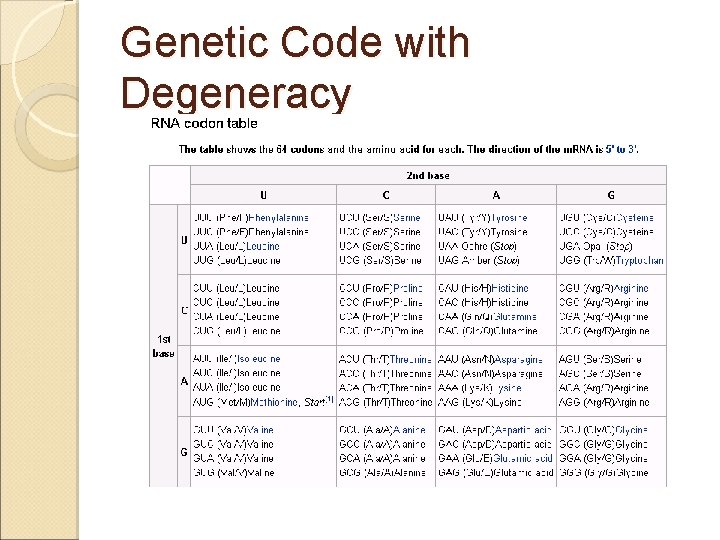
Genetic Code with Degeneracy

Representation of Sequences �Single DNA sequence ◦ ATCCTTAAGGAAA �Multiple sequences with similarity ◦ Regular Expression ◦ ATAAA ◦ ACAAAA ◦ ATAAAAAA ◦ A[TC]A+
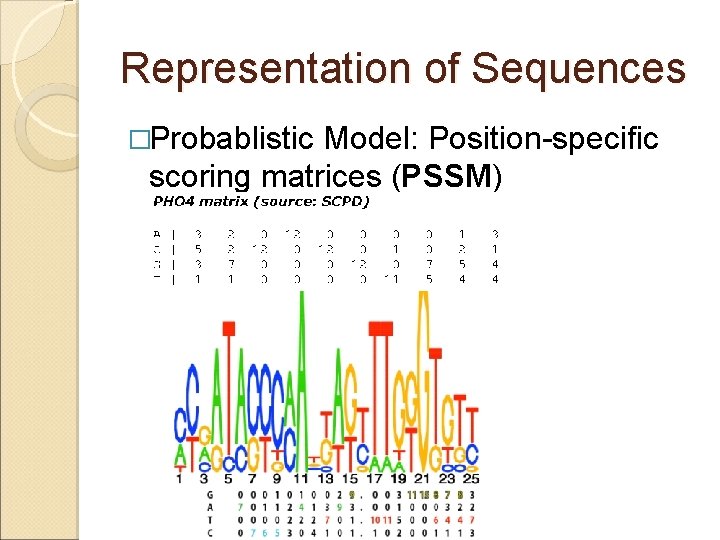
Representation of Sequences �Probablistic Model: Position-specific scoring matrices (PSSM)

Representation of Sequence: FASTA format �text-based format for representing either nucleic acid sequences or peptide sequences, �allows for sequence names and comments to precede the sequences.

Roadmap � DNA, Chromosomes, Genomes � Genome � DNA Sequencing and whole genomes Sequence Representation, Models � Sequence � Basic Retrieval, Manipulation Analysis and Questions of Genomes � Summary 2/14/2022 22

Sequence Retrieval, Manipulation �Where to download genome/sequence data ◦ Online databases: EMBL, Gen. Bank ◦ Entrez cross-database search (life science search engine) ◦ Goolge -

Example: Download H. influenzae Genome �First bacterial genome: H. influenzae, 1830 Kb �http: //www. ncbi. nlm. nih. gov/sites/entre z � NC_007146 Links. Haemophilus influenzae 86 -028 NP, complete genome DNA; circular; Length: 1, 914, 490 nt Replicon Type: chromosome Created: 2005/06/27
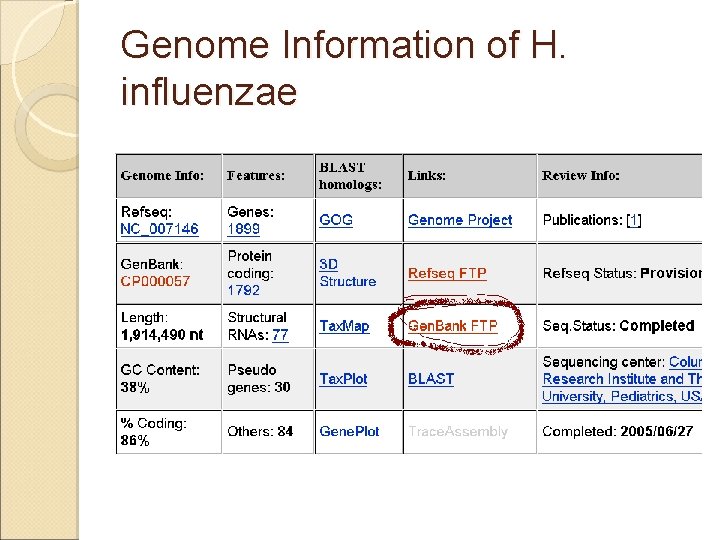
Genome Information of H. influenzae

Download the Complete Genome Sequence in Fasta Format

Roadmap � DNA, Chromosomes, Genomes � Genome � DNA Sequencing and whole genomes Sequence Representation, Models � Sequence � Basic Retrieval, Manipulation Analysis and Questions of Genomes � Summary 2/14/2022 28

Simple Questions and Analysis of Genome Sequence �Frequencies of Bases A/C/G/T by simple counting �Sliding windows to check local density �AT AG AC TA TG TC �K-mers frequent/unusual words ◦ 2 -mers AT AG AC TA TG TC etc. ◦ 3 -mers

Genomic landscape: GC content analysis �The overall GC content of the human genome is 41%. �A plot of GC content versus number of 20 kb windows shows a broad profile with skewing to the right. Page 627
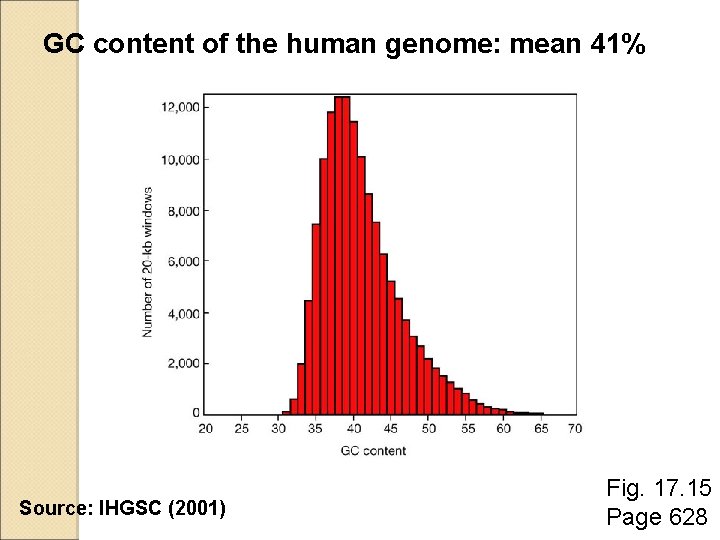
GC content of the human genome: mean 41% Source: IHGSC (2001) Fig. 17. 15 Page 628

Genomic landscape: Cp. G islands � Dinucleotides of Cp. G are under-represented in genomic DNA, occuring at one fifth the expected frequency. � Cp. G dinucleotides are often methylated on cytosine (and subsequently may be deamination to thymine). � Methylated Cp. G residues are often associated with house-keeping genes in the promoter and exonic regions. � Methyl-Cp. G binding proteins recruit histone deacetylases and are thus responsible for transcriptional repression. � They have roles in gene silencing, genomic imprinting, and X-chromosome inactivation.

Broad genomic landscape: Cp. G islands �Findings: ◦ 50, 267 Cp. G islands in human genome ◦ 28, 890 after masking repeats with Repeat. Masker ◦ 5 -15 Cp. G islands per megabase ◦ (about <40 genes per megabase)

Summary �DNA, Chromosome, Genome �Sequence models �Sequence database, retrieval �Whole genome sequence analysis

Slides Credits �Slides in this presentation are partially based on the work of slides from Internet.
- Slides: 35