CS 581 BIOE 540 Algorithmic Computational Genomics Tandy

CS 581 / BIOE 540: Algorithmic Computational Genomics Tandy Warnow Departments of Bioengineering and Computer Science http: //tandy. cs. illinois. edu

Course Details Office hours: Tuesdays 12: 30‐ 1: 30 (Siebel 3235) Course webpage: http: //tandy. cs. illinois. edu/581‐ 2017. html Textbook: Computational Phylogenetics, available for download at http: //tandy. cs. illinois. edu/textbook. pdf TA: Pranjal Vachaspati (to be confirmed)

Today • Describe some important problems in computational biology, for which students in this course could develop improved methods • Explain how the course will be run • Answer questions

This Course Topics: computational and statistical problems in sequence analysis (e. g. , multiple sequence alignment, phylogeny estimation, metagenomics, etc. ). Focus: understanding the mathematical foundations, and designing algorithms with outstanding accuracy and speed on large, complex datasets. This is not a course about how to use the tools.

Prerequisites No background in biology is needed. However, the course has the following prerequisites: • CS 374: computational complexity, algorithm design techniques, and proving theorems about algorithms • CS 361: probability and statistics • By recursion, CS 225: programming
If you haven’t satisfied the pre‐reqs: You need permission to stay in the course. • The first homework is due (by email) on Saturday at 1 PM. See the homework webpage http: //tandy. cs. illinois. edu/cs 581‐ 2017‐hw. html • Then make an appointment to see me to review the homework.

This course • Phylogeny estimation based on stochastic models of sequence evolution and genome evolution • Multiple sequence alignment • Applications to metagenomics, protein structure prediction, and other biological problems

Species Tree Orangutan From the Tree of the Life Website, University of Arizona Gorilla Chimpanzee Human

Evolution informs about everything in biology • Big genome sequencing projects just produce data so‐‐‐‐‐‐ what? • Evolutionary history relates all organisms and genes, and helps us understand predict – interactions between genes (genetic networks) – drug design – predicting functions of genes – influenza vaccine development – origins and spread of disease – origins and migrations of humans
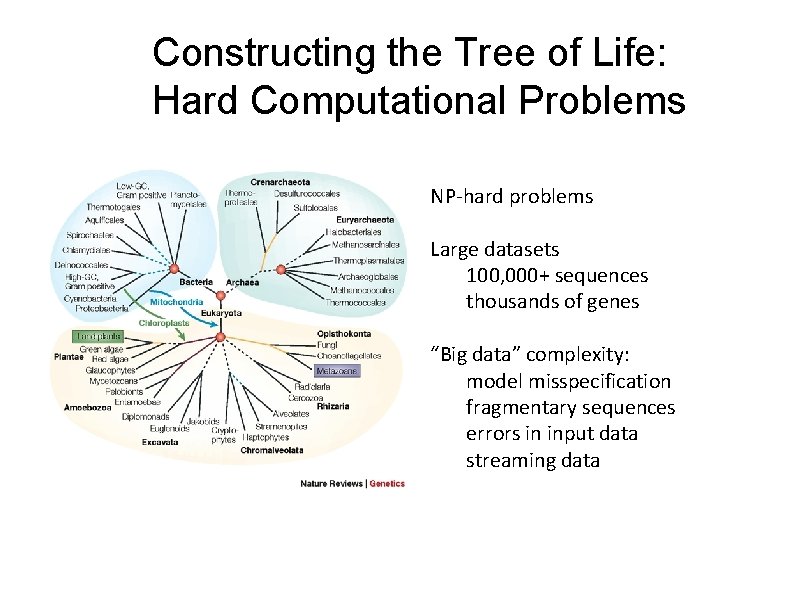
Constructing the Tree of Life: Hard Computational Problems NP‐hard problems Large datasets 100, 000+ sequences thousands of genes “Big data” complexity: model misspecification fragmentary sequences errors in input data streaming data

Phylogenomic pipeline Select taxon set and markers Gather and screen sequence data, possibly identify orthologs Compute multiple sequence alignments for each locus Compute species tree or network: Compute gene trees on the alignments and combine the estimated gene trees, OR Estimate a tree from a concatenation of the multiple sequence alignments Get statistical support on each branch (e. g. , bootstrapping) Estimate dates on the nodes of the phylogeny Use species tree with branch support and dates to understand biology

Phylogenomic pipeline Select taxon set and markers Gather and screen sequence data, possibly identify orthologs Compute multiple sequence alignments for each locus Compute species tree or network: Compute gene trees on the alignments and combine the estimated gene trees, OR Estimate a tree from a concatenation of the multiple sequence alignments Get statistical support on each branch (e. g. , bootstrapping) Estimate dates on the nodes of the phylogeny Use species tree with branch support and dates to understand biology

Research Strategies Improved algorithms through: • • Divide‐and‐conquer “Bin‐and‐conquer” Iteration Bayesian statistics Hidden Markov Models Graph theory Combinatorial optimization Statistical modelling Massive Simulations High Performance Computing

Avian Phylogenomics Project Erich Jarvis, HHMI • Approx. MTP Gilbert, Copenhagen G Zhang, BGI 50 species, whole genomes • 14, 000 loci T. Warnow UT‐Austin S. Mirarab Md. S. Bayzid, UT‐Austin UT‐ Plus many other people… Science, December 2014 (Jarvis, Mirarab, et al. , and Mirarab et al. )

1 kp: Thousand Transcriptome Project G. Ka-Shu Wong U Alberta J. Leebens-Mack U Georgia N. Wickett Northwestern N. Matasci i. Plant T. Warnow, UIUC S. Mirarab, UT-Austin N. Nguyen, UT-Austin Plus many other people… Plant Tree of Life based on transcriptomes of ~1200 species More than 13, 000 gene families (most not single copy) First paper: PNAS 2014 (~100 species and ~800 loci) Gene Tree Incongruence Upcoming Challenges (~1200 species, ~400 loci)
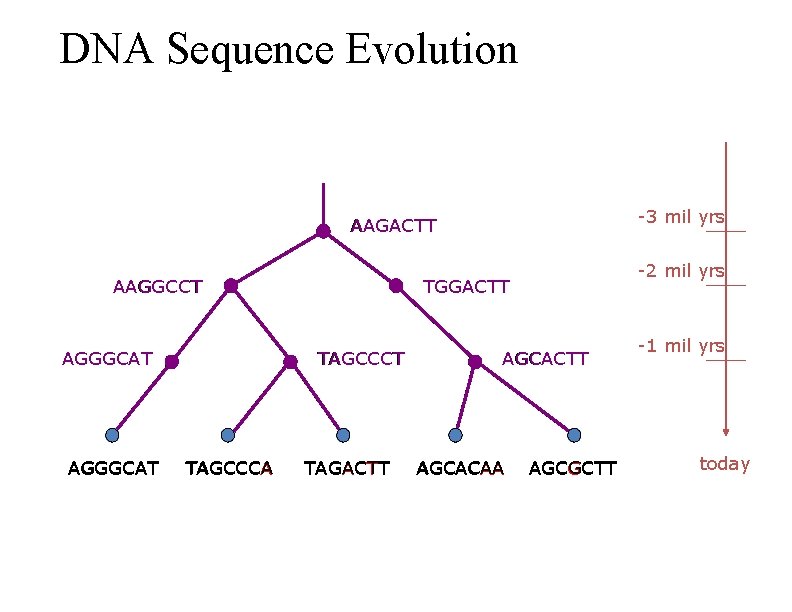
DNA Sequence Evolution -3 mil yrs AAGACTT AAGGCCT AGGGCAT TGGACTT TAGCCCA -2 mil yrs TAGACTT AGCACAA AGCGCTT -1 mil yrs today
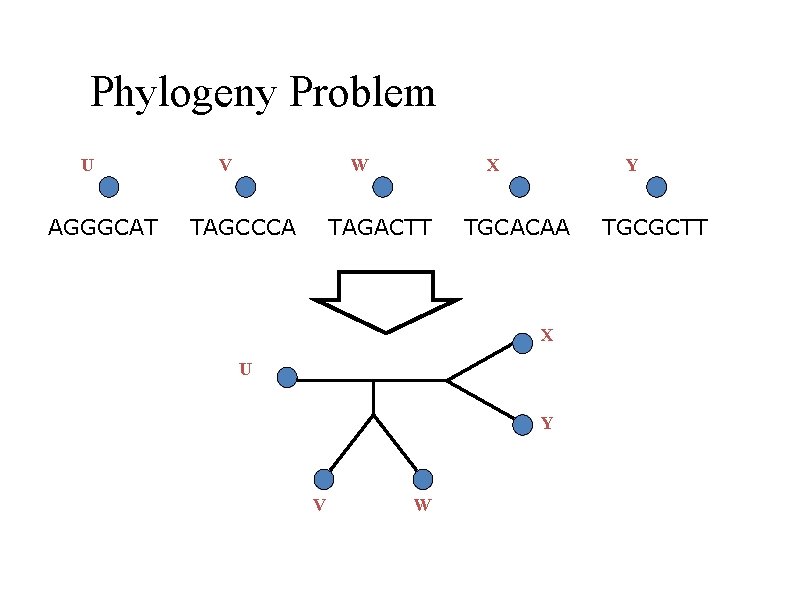
Phylogeny Problem U AGGGCAT V W TAGCCCA X TAGACTT Y TGCACAA X U Y V W TGCGCTT

Performance criteria • Running time • Space • Statistical performance issues (e. g. , statistical consistency) with respect to a Markov model of evolution • “Topological accuracy” with respect to the underlying true tree or true alignment, typically studied in simulation • Accuracy with respect to a particular criterion (e. g. maximum likelihood score), on real data
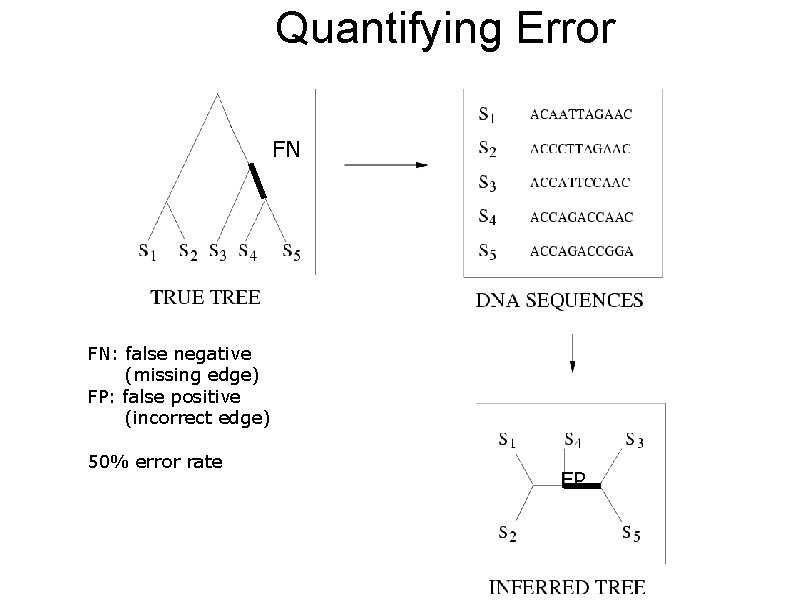
Quantifying Error FN FN: false negative (missing edge) FP: false positive (incorrect edge) 50% error rate FP
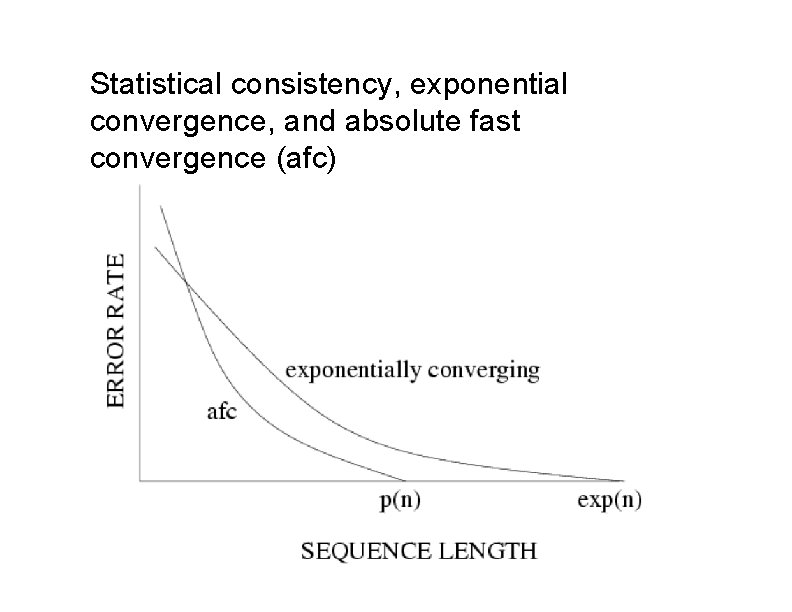
Statistical consistency, exponential convergence, and absolute fast convergence (afc)
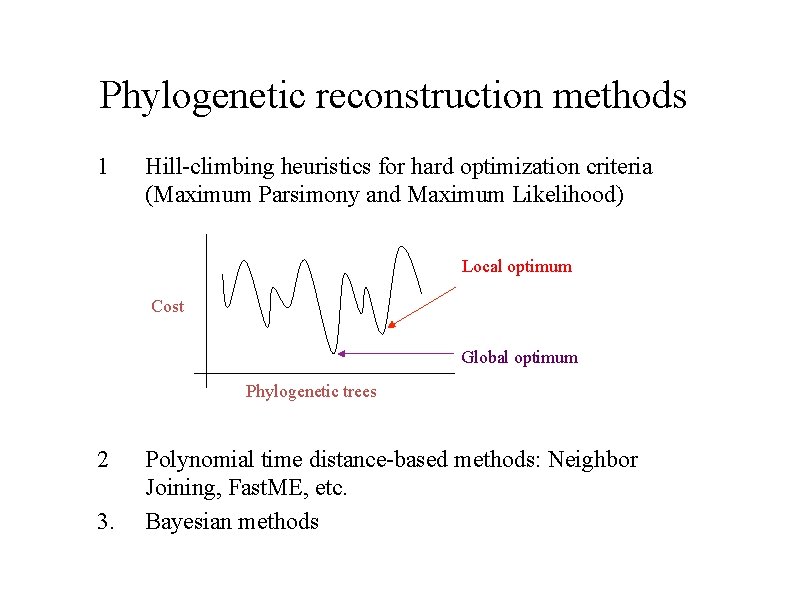
Phylogenetic reconstruction methods 1 Hill-climbing heuristics for hard optimization criteria (Maximum Parsimony and Maximum Likelihood) Local optimum Cost Global optimum Phylogenetic trees 2 3. Polynomial time distance-based methods: Neighbor Joining, Fast. ME, etc. Bayesian methods

Solving maximum likelihood (and other hard optimization problems) is… unlikely # of Taxa # of Unrooted Trees 4 3 5 15 6 105 7 945 8 10395 9 135135 10 2027025 20 2. 2 x 1020 100 4. 5 x 10190 1000 2. 7 x 102900
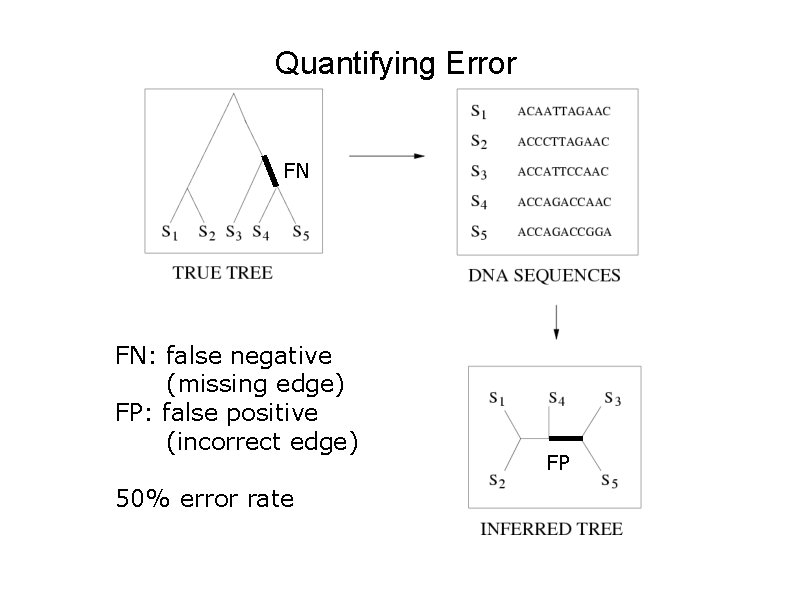
Quantifying Error FN FN: false negative (missing edge) FP: false positive (incorrect edge) 50% error rate FP
![Neighbor joining has poor performance on large diameter trees [Nakhleh et al. ISMB 2001] Neighbor joining has poor performance on large diameter trees [Nakhleh et al. ISMB 2001]](http://slidetodoc.com/presentation_image/99e92df184ce6162796b91f8304a9b72/image-24.jpg)
Neighbor joining has poor performance on large diameter trees [Nakhleh et al. ISMB 2001] 0. 8 NJ Error Rate Theorem (Atteson): Exponential sequence length requirement for Neighbor Joining! 0. 6 0. 4 0. 2 0 0 400 800 No. Taxa 1200 1600

Major Challenges • Phylogenetic analyses: standard methods have poor accuracy on even moderately large datasets, and the most accurate methods are enormously computationally intensive (weeks or months, high memory requirements)
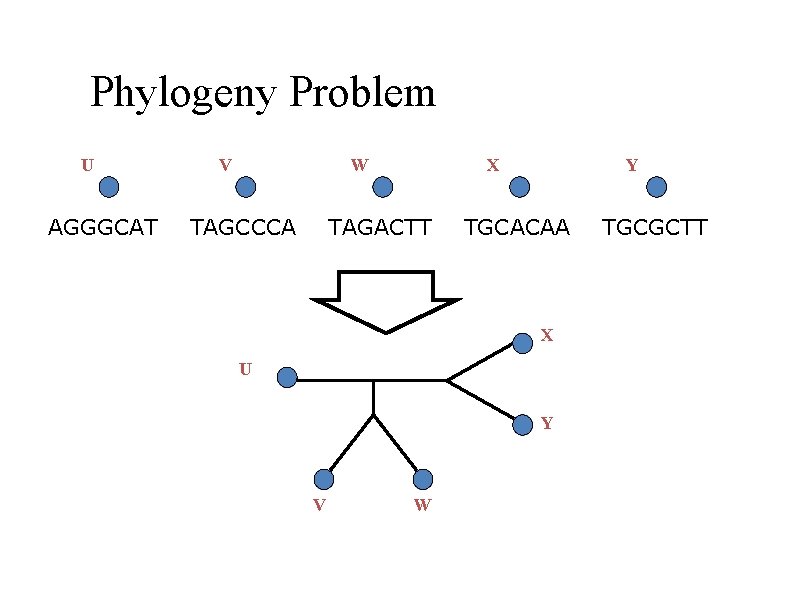
Phylogeny Problem U AGGGCAT V W TAGCCCA X TAGACTT Y TGCACAA X U Y V W TGCGCTT
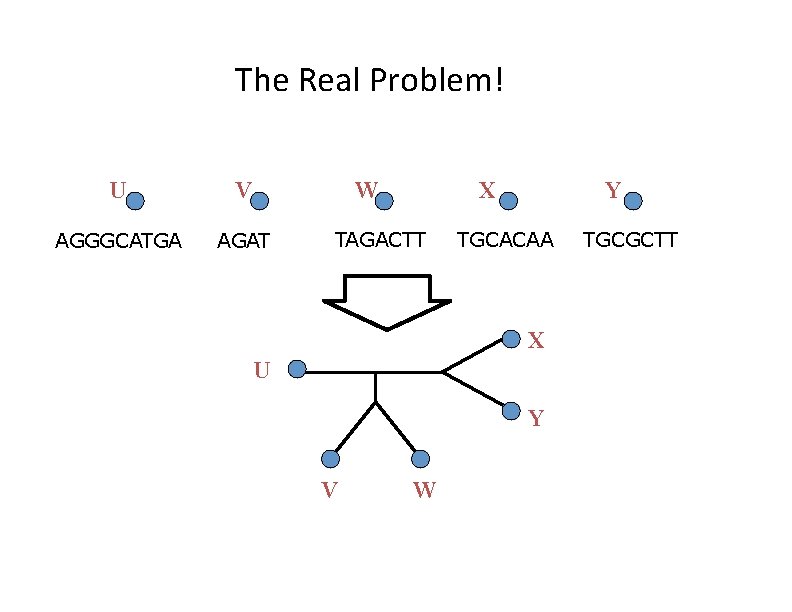
The Real Problem! U V W AGGGCATGA AGAT X TAGACTT Y TGCACAA X U Y V W TGCGCTT
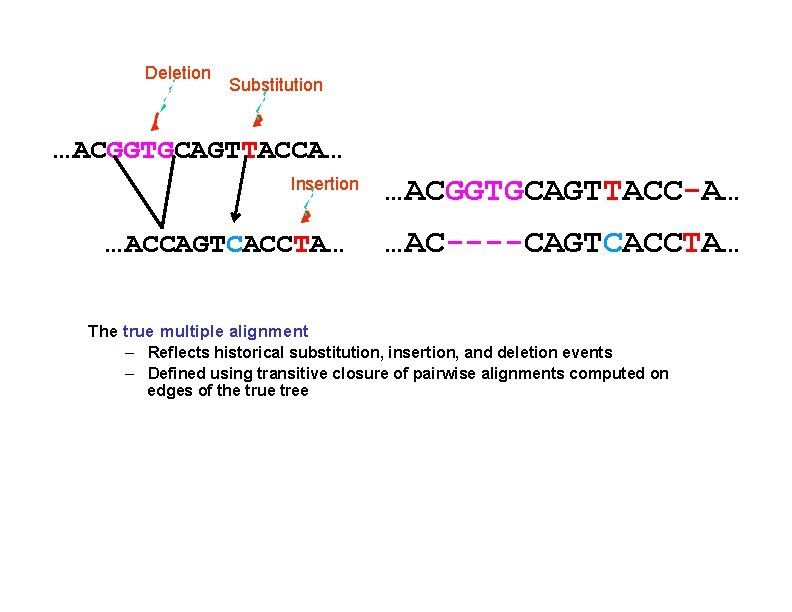
Deletion Substitution …ACGGTGCAGTTACCA… Insertion …ACCAGTCACCTA… …ACGGTGCAGTTACC-A… …AC----CAGTCACCTA… The true multiple alignment – Reflects historical substitution, insertion, and deletion events – Defined using transitive closure of pairwise alignments computed on edges of the true tree

Input: unaligned sequences S 1 S 2 S 3 S 4 = = AGGCTATCACCTGACCTCCA TAGCTATCACGACCGC TAGCTGACCGC TCACGACA
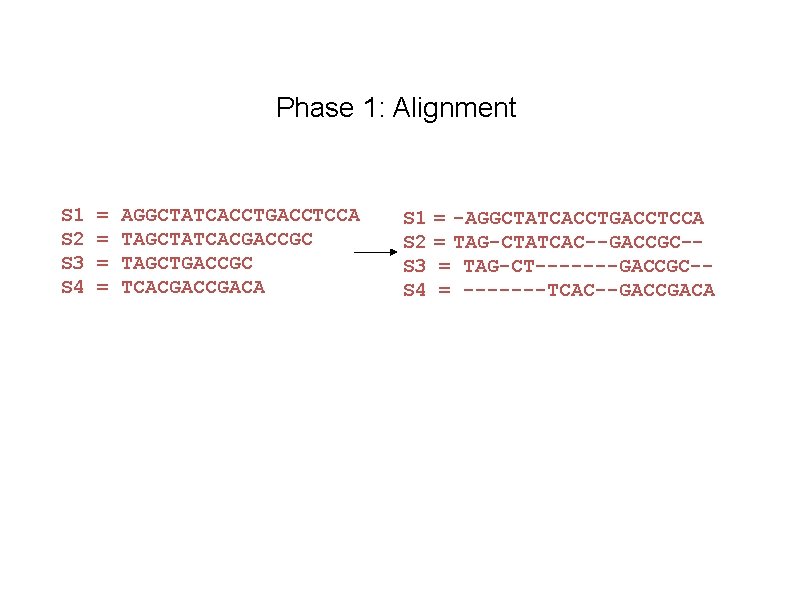
Phase 1: Alignment S 1 S 2 S 3 S 4 = = AGGCTATCACCTGACCTCCA TAGCTATCACGACCGC TAGCTGACCGC TCACGACA S 1 S 2 S 3 S 4 = -AGGCTATCACCTGACCTCCA = TAG-CTATCAC--GACCGC-= TAG-CT-------GACCGC-= -------TCAC--GACCGACA
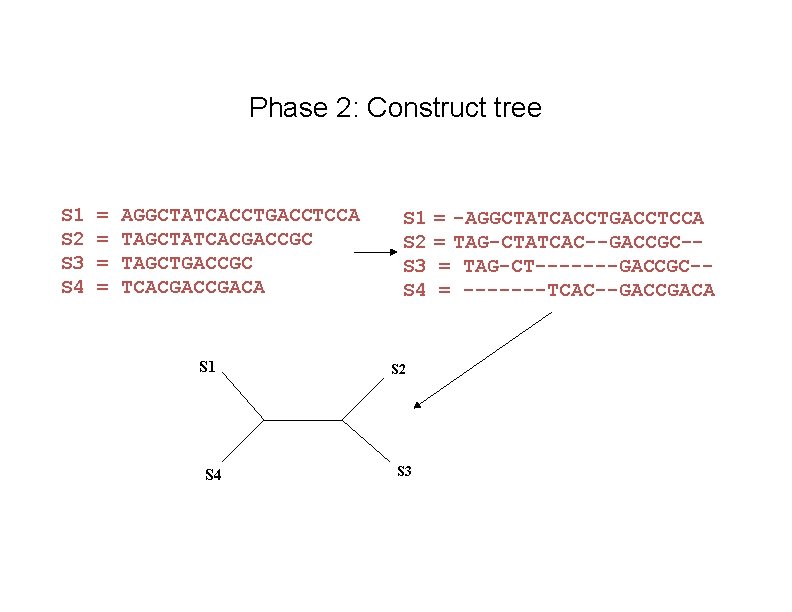
Phase 2: Construct tree S 1 S 2 S 3 S 4 = = AGGCTATCACCTGACCTCCA TAGCTATCACGACCGC TAGCTGACCGC TCACGACA S 1 S 4 S 1 S 2 S 3 S 4 S 2 S 3 = -AGGCTATCACCTGACCTCCA = TAG-CTATCAC--GACCGC-= TAG-CT-------GACCGC-= -------TCAC--GACCGACA

Two-phase estimation Alignment methods • Clustal • POY (and POY*) • Probcons (and Probtree) • Probalign • MAFFT • Muscle • Di-align • T-Coffee • Prank (PNAS 2005, Science 2008) • Opal (ISMB and Bioinf. 2007) • FSA (PLo. S Comp. Bio. 2009) • Infernal (Bioinf. 2009) • Etc. Phylogeny methods • Bayesian MCMC • Maximum parsimony • Maximum likelihood • Neighbor joining • Fast. ME • UPGMA • Quartet puzzling • Etc.
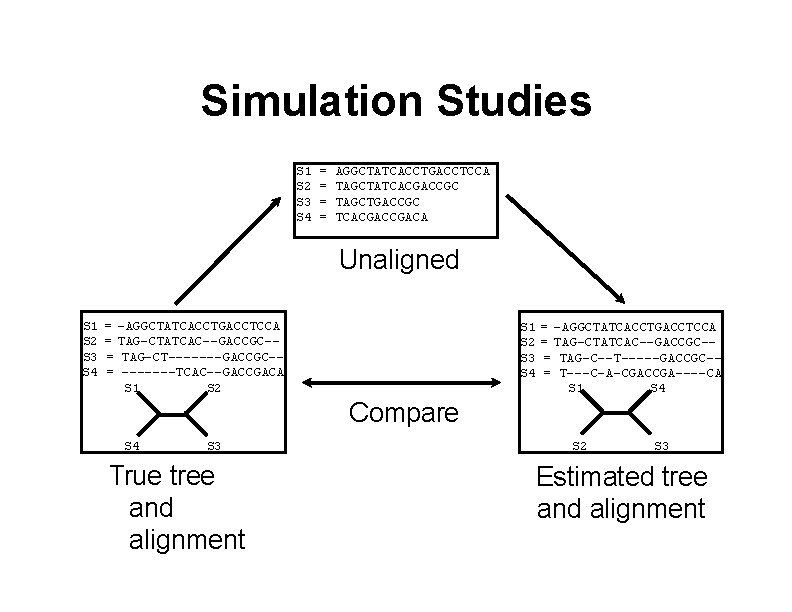
Simulation Studies S 1 S 2 S 3 S 4 = = AGGCTATCACCTGACCTCCA TAGCTATCACGACCGC TAGCTGACCGC TCACGACA Unaligned S 1 S 2 S 3 S 4 = -AGGCTATCACCTGACCTCCA = TAG-CTATCAC--GACCGC-= TAG-CT-------GACCGC-= -------TCAC--GACCGACA S 1 S 2 S 3 S 4 = -AGGCTATCACCTGACCTCCA = TAG-CTATCAC--GACCGC-= TAG-C--T-----GACCGC-= T---C-A-CGACCGA----CA S 1 S 4 Compare S 4 S 3 True tree and alignment S 2 S 3 Estimated tree and alignment
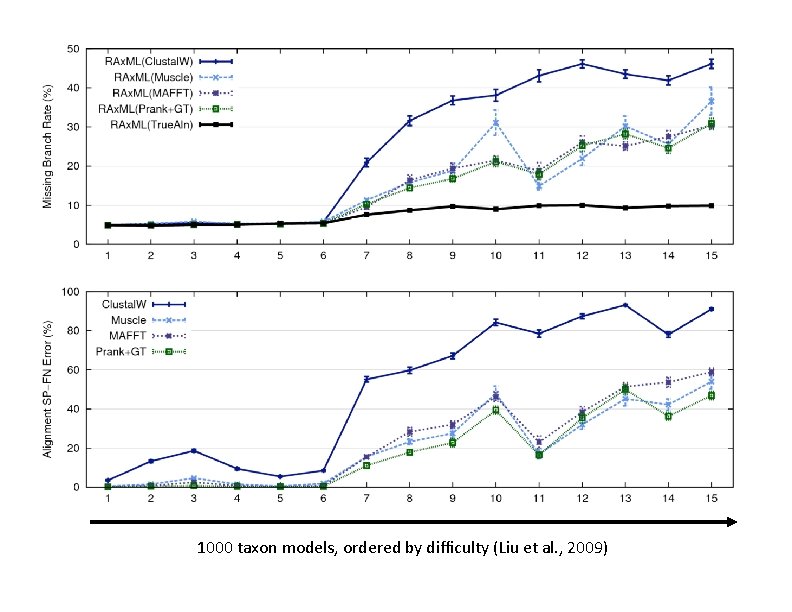
1000 taxon models, ordered by difficulty (Liu et al. , 2009)

Multiple Sequence Alignment (MSA): another grand challenge 1 S 1 = S 2 = S 3 = … Sn = AGGCTATCACCTGACCTCCA TAGCTATCACGACCGC TAGCTGACCGC TCACGACA S 1 S 2 S 3 … Sn = -AGGCTATCACCTGACCTCCA = TAG-CTATCAC--GACCGC-= TAG-CT-------GACCGC-= -------TCAC--GACCGACA Novel techniques needed for scalability and accuracy NP-hard problems and large datasets Current methods do not provide good accuracy Few methods can analyze even moderately large datasets Many important applications besides phylogenetic estimation 1 Frontiers in Massive Data Analysis, National Academies Press, 2013

Major Challenges • Phylogenetic analyses: standard methods have poor accuracy on even moderately large datasets, and the most accurate methods are enormously computationally intensive (weeks or months, high memory requirements) • Multiple sequence alignment: key step for many biological questions (protein structure and function, phylogenetic estimation), but few methods can run on large datasets. Alignment accuracy is generally poor for large datasets with high rates of evolution.
(Phylogenetic estimation from whole genomes)

Species Tree Estimation requires multiple genes! Orangutan From the Tree of the Life Website, University of Arizona Gorilla Chimpanzee Human

Two basic approaches for species tree estimation • Concatenate (“combine”) sequence alignments for different genes, and run phylogeny estimation methods • Compute trees on individual genes and combine gene trees
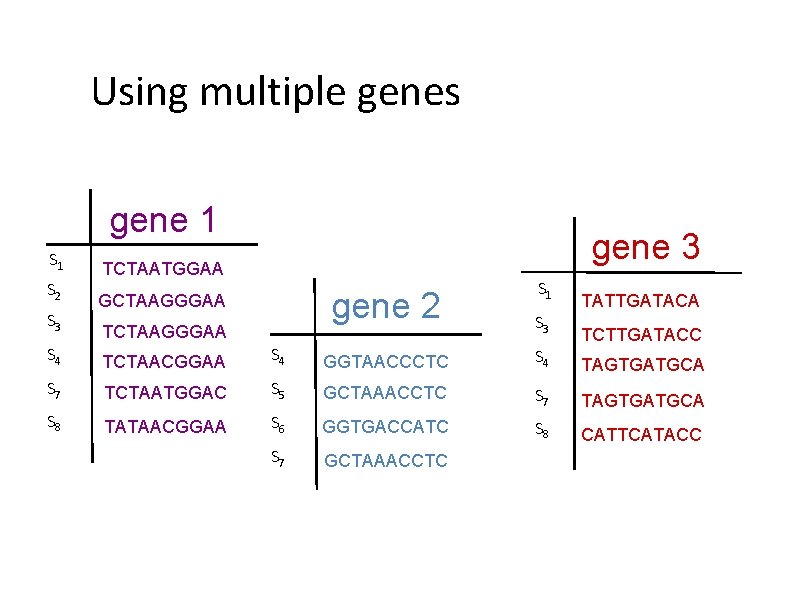
Using multiple genes gene 1 S 2 S 3 gene 3 TCTAATGGAA gene 2 GCTAAGGGAA TCTAAGGGAA S 1 TATTGATACA S 3 TCTTGATACC S 4 TCTAACGGAA S 4 GGTAACCCTC S 4 TAGTGATGCA S 7 TCTAATGGAC S 5 GCTAAACCTC S 7 S 8 TAGTGATGCA TATAACGGAA S 6 GGTGACCATC S 8 CATTCATACC S 7 GCTAAACCTC
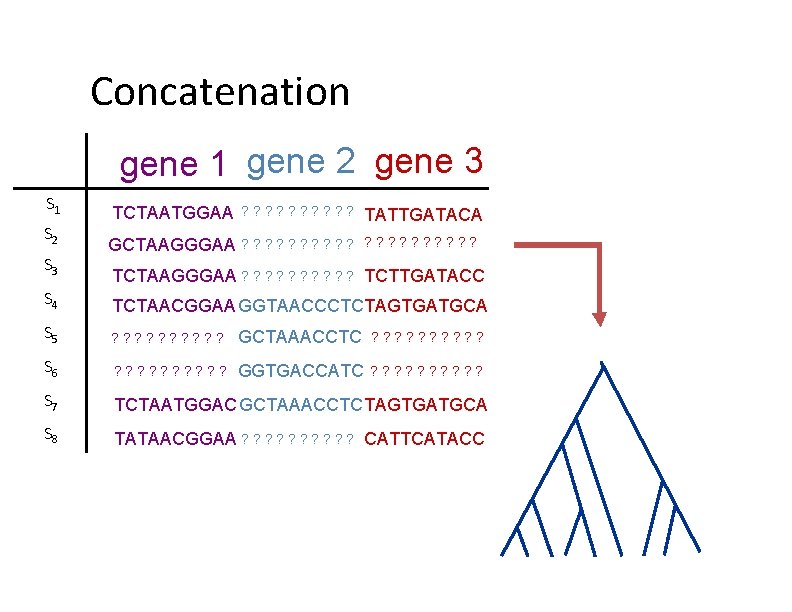
Concatenation gene 1 gene 2 gene 3 S 1 S 2 S 3 TCTAATGGAA ? ? ? ? ? GCTAAGGGAA ? ? ? ? ? TATTGATACA ? ? ? ? ? TCTAAGGGAA ? ? ? ? ? TCTTGATACC S 4 TCTAACGGAA GGTAACCCTCTAGTGATGCA S 5 ? ? ? ? ? GCTAAACCTC ? ? ? ? ? S 6 ? ? ? ? ? GGTGACCATC ? ? ? ? ? S 7 TCTAATGGAC GCTAAACCTC TAGTGATGCA S 8 TATAACGGAA ? ? ? ? ? CATTCATACC
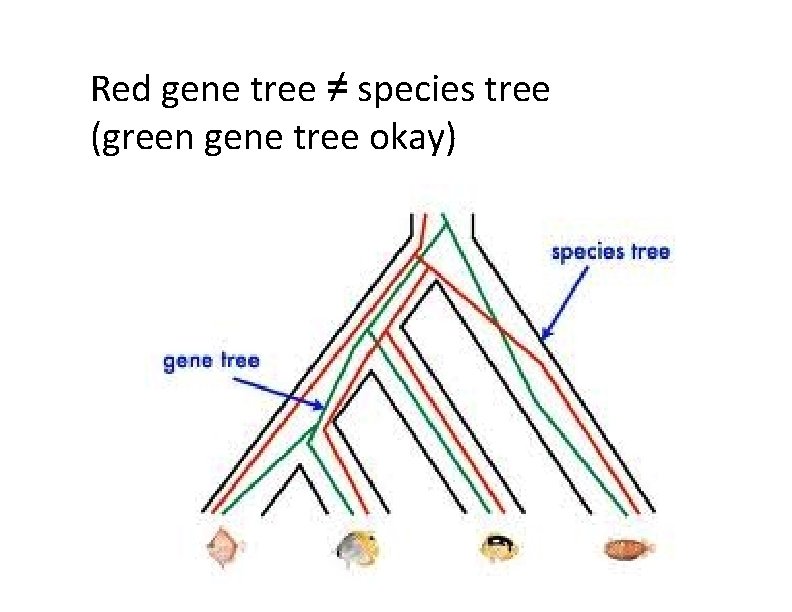
Red gene tree ≠ species tree (green gene tree okay)

Gene Tree Incongruence Gene trees can differ from the species tree due to: Duplication and loss Horizontal gene transfer Incomplete lineage sorting (ILS)

Incomplete Lineage Sorting (ILS) Confounds phylogenetic analysis for many groups: Hominids Birds Yeast Animals Toads Fish Fungi There is substantial debate about how to analyze phylogenomic datasets in the presence of ILS.

Lineage Sorting Population‐level process, also called the “Multi‐species coalescent” (Kingman, 1982) Gene trees can differ from species trees due to short times between speciation events or large population size; this is called “Incomplete Lineage Sorting” or “Deep Coalescence”.
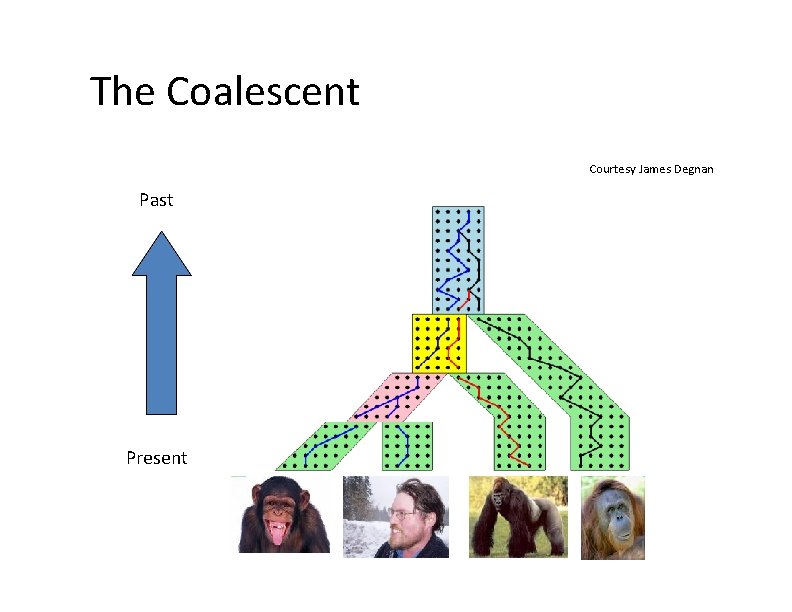
The Coalescent Courtesy James Degnan Past Present

Gene tree in a species tree Courtesy James Degnan

Key observation: Under the multi‐species coalescent model, the species tree defines a probability distribution on the gene trees, and is identifiable from the distribution on gene trees Courtesy James Degnan
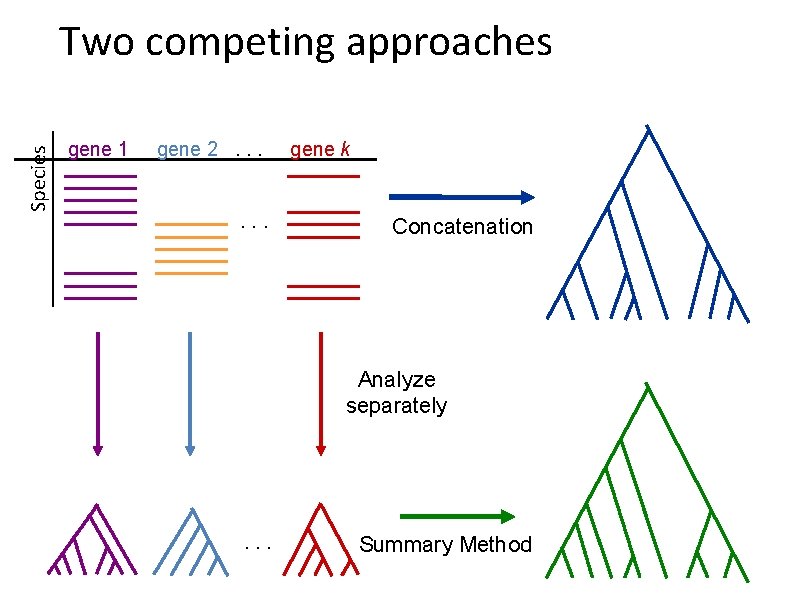
Species Two competing approaches gene 1 gene 2. . . gene k Concatenation Analyze separately . . . Summary Method

Species tree estimation: difficult, even for small datasets! Orangutan From the Tree of the Life Website, University of Arizona Gorilla Chimpanzee Human

Major Challenges: large datasets, fragmentary sequences • Multiple sequence alignment: Few methods can run on large datasets, and alignment accuracy is generally poor for large datasets with high rates of evolution. • Gene Tree Estimation: standard methods have poor accuracy on even moderately large datasets, and the most accurate methods are enormously computationally intensive (weeks or months, high memory requirements). • Species Tree Estimation: gene tree incongruence makes accurate estimation of species tree challenging. • Phylogenetic Network Estimation: Horizontal gene transfer and hybridization requires non-tree models of evolution Both phylogenetic estimation and multiple sequence alignment are also impacted by fragmentary data.

Major Challenges: large datasets, fragmentary sequences • Multiple sequence alignment: Few methods can run on large datasets, and alignment accuracy is generally poor for large datasets with high rates of evolution. • Gene Tree Estimation: standard methods have poor accuracy on even moderately large datasets, and the most accurate methods are enormously computationally intensive (weeks or months, high memory requirements). • Species Tree Estimation: gene tree incongruence makes accurate estimation of species tree challenging. • Phylogenetic Network Estimation: Horizontal gene transfer and hybridization requires non-tree models of evolution Both phylogenetic estimation and multiple sequence alignment are also impacted by fragmentary data.

Avian Phylogenomics Project Erich Jarvis, HHMI • Approx. MTP Gilbert, Copenhagen G Zhang, BGI 50 species, whole genomes • 14, 000 loci T. Warnow UT‐Austin S. Mirarab Md. S. Bayzid, UT‐Austin UT‐ Plus many other people… Challenges: • Species tree estimation under the multi‐species coalescent model, from 14, 000 poor estimated gene trees, all with different topologies (we used “statistical binning”) • Maximum likelihood estimation on a million‐site genome‐scale alignment – 250 CPU years Science, December 2014 (Jarvis, Mirarab, et al. , and Mirarab et al. )

1 kp: Thousand Transcriptome Project G. Ka-Shu Wong U Alberta J. Leebens-Mack U Georgia N. Wickett Northwestern N. Matasci i. Plant T. Warnow, UIUC S. Mirarab, UT-Austin N. Nguyen, UT-Austin Plus many other people… Plant Tree of Life based on transcriptomes of ~1200 species More than 13, 000 gene families (most not single copy) First paper: PNAS 2014 (~100 species and ~800 loci) Gene Tree Incongruence Upcoming Challenges (~1200 species, ~400 loci): • Species tree estimation under the multi‐species coalescent from hundreds of conflicting gene trees on >1000 species (we will use ASTRAL – Mirarab et al. 2014, Mirarab & Warnow 2015) • Multiple sequence alignment of >100, 000 sequences (with lots of fragments!) – we will use UPP (Nguyen et al. , 2015)
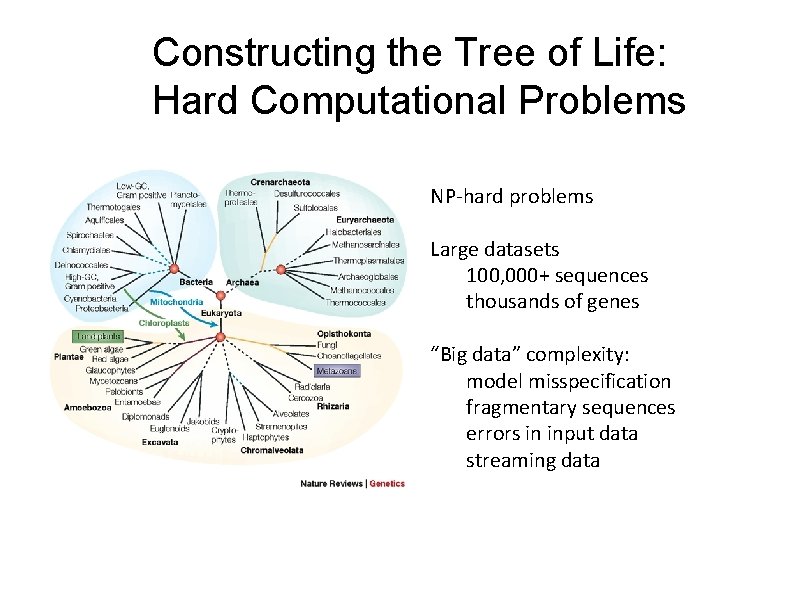
Constructing the Tree of Life: Hard Computational Problems NP‐hard problems Large datasets 100, 000+ sequences thousands of genes “Big data” complexity: model misspecification fragmentary sequences errors in input data streaming data

Research Strategies Improved algorithms through: • • Divide‐and‐conquer “Bin‐and‐conquer” Iteration Bayesian statistics Hidden Markov Models Graph theory Combinatorial optimization Statistical modelling Massive Simulations High Performance Computing

Evolution informs about everything in biology • Big genome sequencing projects just produce data so‐‐‐‐‐‐ what? • Evolutionary history relates all organisms and genes, and helps us understand predict – interactions between genes (genetic networks) – drug design – predicting functions of genes – influenza vaccine development – origins and spread of disease – origins and migrations of humans

Metagenomics: Venter et al. , Exploring the Sargasso Sea: Scientists Discover One Million New Genes in Ocean Microbes

Metagenomic data analysis NGS data produce fragmentary sequence data Metagenomic analyses include unknown species Taxon identification: given short sequences, identify the species for each fragment Applications: Human Microbiome Issues: accuracy and speed Mihai Pop, Univ Maryland

Metagenomic taxon identification Objective: classify short reads in a metagenomic sample
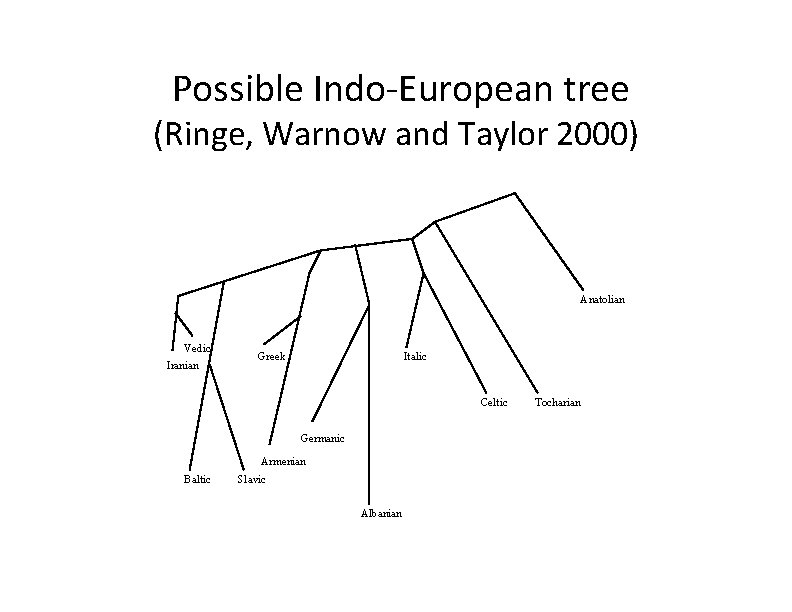
Possible Indo‐European tree (Ringe, Warnow and Taylor 2000) Anatolian Vedic Iranian Greek Italic Celtic Germanic Armenian Baltic Slavic Albanian Tocharian
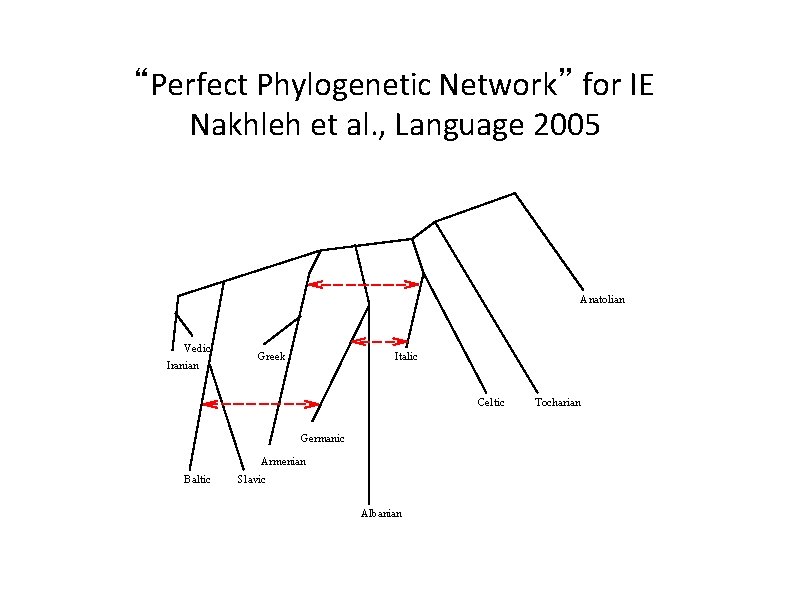
“Perfect Phylogenetic Network” for IE Nakhleh et al. , Language 2005 Anatolian Vedic Iranian Greek Italic Celtic Germanic Armenian Baltic Slavic Albanian Tocharian

Grading • • Homework: 25% (one hw dropped) Midterm: 40% (March 30) Final Project: 25% (due May 6) Course Participation: 10% No final exam.

Homework Assignments • Homework assignments are listed at http: //tandy. cs. illinois. edu/cs 581‐ 2017‐ hw. html and are due at 1 PM (in person or via email) – late homeworks have reduced credit and will not be accepted after 48 hours past the deadline. • You are encouraged to work with others on your homework, but you must write up solutions by yourself and indicate who you worked with on each homework.

Course Schedule A detailed course schedule is athttp: //tandy. cs. illinois. edu/cs 581‐ 2017‐ detailed‐syllabus. html This schedule includes material you are expected to have looked at before coming to class: • assigned reading (from textbook and/or scientific literature) • PPT and/or PDF of my lecture

Final Project and Class Presentation Either research project (can be with another student) or survey paper (done by yourself). Many interesting and publishable problems to address: see http: //tandy. cs. illinois. edu/topics. html Your class presentation should be related to your final project.

Academic Integrity Please see course website at http: //tandy. cs. illinois. edu/581‐ 2017. html and also http: //tandy. cs. illinois. edu/ethics. pdf For this course: • Examine the policy about collaboration • Learn and understand what plagiarism is (and then don’t do it). This applies to homework, all writing assignments, and the final project.

Course Research Projects • Evaluating existing methods on simulated and real (biological or linguistic) datasets • Designing a new method, and establishing its performance (using theory and data) • Analyzing a biological dataset using several different methods, to address biology

Examples of published course projects Md S. Bayzid, T. Hunt, and T. Warnow. "Disk Covering Methods Improve Phylogenomic Analyses". Proceedings of RECOMB‐CG (Comparative Genomics), 2014, and BMC Genomics 2014, 15(Suppl 6): S 7. T. Zimmermann, S. Mirarab and T. Warnow. "BBCA: Improving the scalability of *BEAST using random binning". Proceedings of RECOMB‐CG (Comparative Genomics), 2014, and BMC Genomics 2014, 15(Suppl 6): S 11. J. Chou, A. Gupta, S. Yaduvanshi, R. Davidson, M. Nute, S. Mirarab and T. Warnow. “A comparative study of SVDquartets and other coalescent‐based species tree estimation methods”. RECOMB‐Comparative Genomics and BMC Genomics, 2015, 16 (Suppl 10): S 2. P. Vachaspati and T. Warnow (2016). Fast. RFS: Fast and Accurate Robinson‐Foulds Supertrees using Constrained Exact Optimization Bioinformatics 2016; doi: 10. 1093/bioinformatics/btw 600. (Special issue for papers from RECOMB‐CG)

Research Projects you could join • Phylogenomics projects (Avian and the 1 KP) • Species tree and network estimation from conflicting genes • Large‐scale multiple sequence alignment • Large‐scale maximum likelihood tree estimation • Improving gene tree estimation using whole genomes • Metagenomics (with Mihai Pop, University of Maryland, and Bill Gropp) • Identifying genes and taxa from short sequences • Metagenomic assembly • Applications to clinical diagnostics • Protein Sequence Analysis (with Jian Peng) • What function and structure does this protein have? • How did structure and function evolve? • Historical Linguistics (with Donald Ringe, UPenn) • How did Indo‐European evolve? • Designing and implementing statistical estimation methods for language phylogenies
- Slides: 70