CryoElectron Microscopy James Conway University of Pittsburgh School

Cryo-Electron Microscopy James Conway University of Pittsburgh School of Medicine 1. Making images image formation contrast function detectors – film & scanning, CCD, DED energy filters phase plates 2. Making density maps (AUTO 3 DEM) calculating 3 D density maps from 2 D projections estimating resolution modelling Penn State Med School – Friday, 4 -Oct-2013

1. How does 3 D reconstruction work anyway? • Images are 2 D projections of the 3 D structure • Simple approach is to back-project into a 3 D volume • More computationally efficient is to perform the equivalent operation in Fourier (reciprocal) space • Easier to demonstrate back-projection using 1 D projections from a 2 D object Penn State Med School – Friday, 4 -Oct-2013
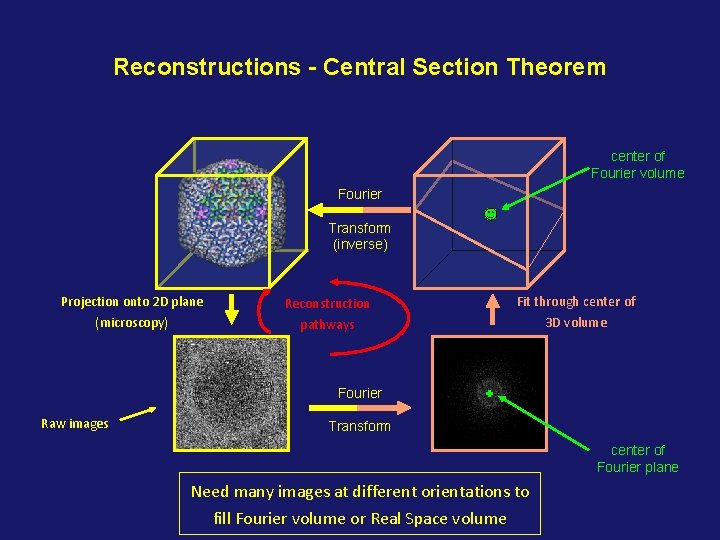
Reconstructions - Central Section Theorem center of Fourier volume Fourier Transform (inverse) Projection onto 2 D plane Reconstruction Fit through center of (microscopy) pathways 3 D volume Fourier Raw images Transform center of Fourier plane Need many images at different orientations to fill Fourier volume or Real Space volume
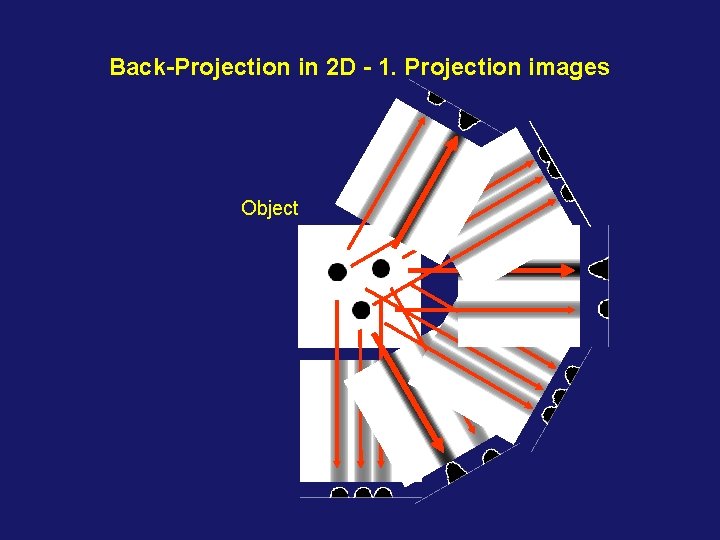
Back-Projection in 2 D - 1. Projection images Object
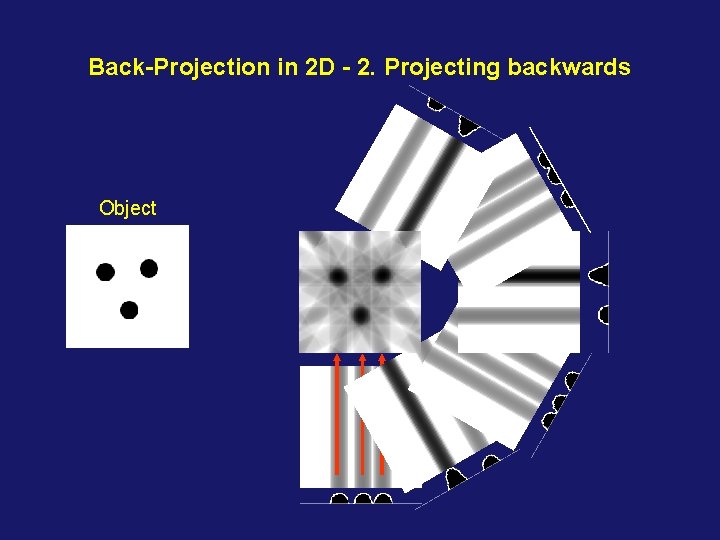
Back-Projection in 2 D - 2. Projecting backwards Object

Back-Projection in 2 D - 3. Finer sampling Object 30° 10° 5° 1°
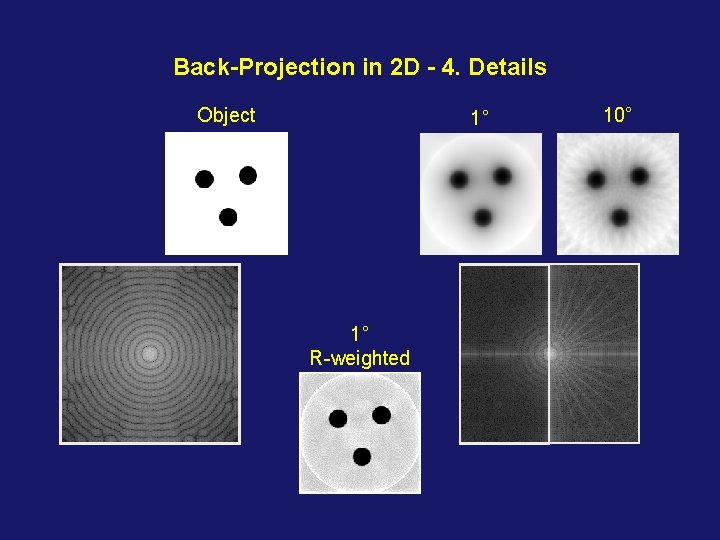
Back-Projection in 2 D - 4. Details Object 1° 1° R-weighted 10°

Back-Projection in 2 D - 5. Another Example Object 1° R-weighted
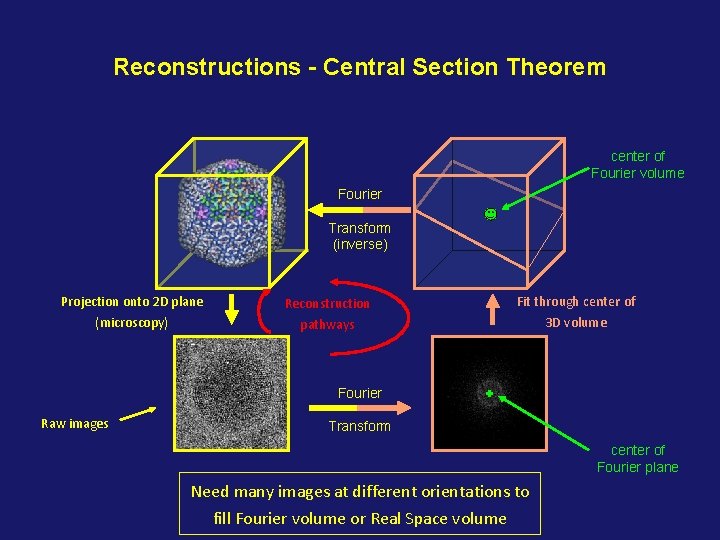
Reconstructions - Central Section Theorem center of Fourier volume Fourier Transform (inverse) Projection onto 2 D plane Reconstruction Fit through center of (microscopy) pathways 3 D volume Fourier Raw images Transform center of Fourier plane Need many images at different orientations to fill Fourier volume or Real Space volume

2. Reconstruction Process a. Particle Picking b. CTF estimation c. RMC – initial model search d. Auto 3 Dem Tim Baker Purdue/UCSD e. Resolution f. Interpretation & modelling Penn State Med School – Friday, 4 -Oct-2013

2 a. Particle Picking 1000Å 500Å Penn State Med School – Friday, 4 -Oct-2013
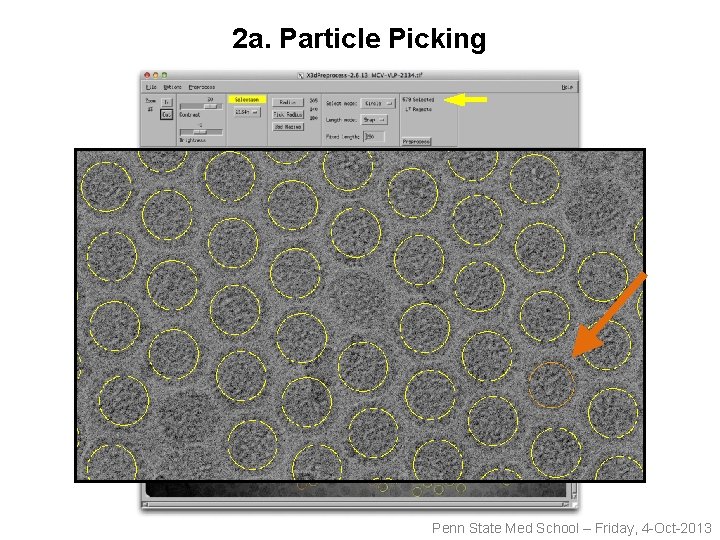
2 a. Particle Picking Penn State Med School – Friday, 4 -Oct-2013

2 a. Particle Picking 579 capsids 21 capsids 451 x 451 pixels Penn State Med School – Friday, 4 -Oct-2013
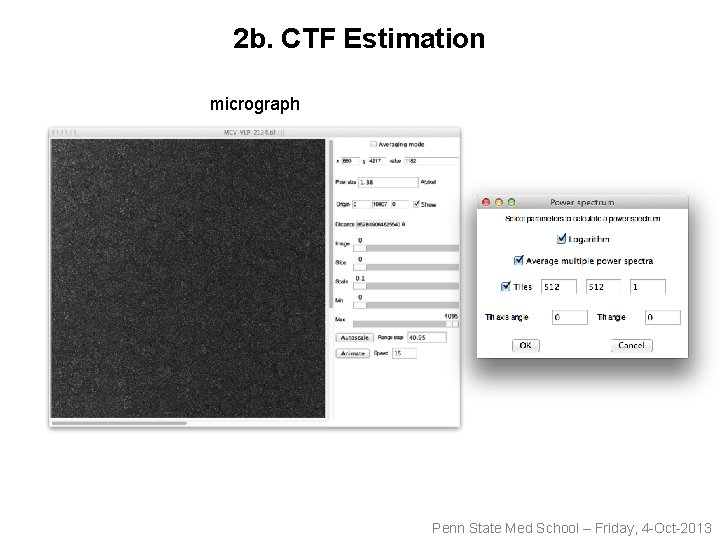
2 b. CTF Estimation micrograph Penn State Med School – Friday, 4 -Oct-2013
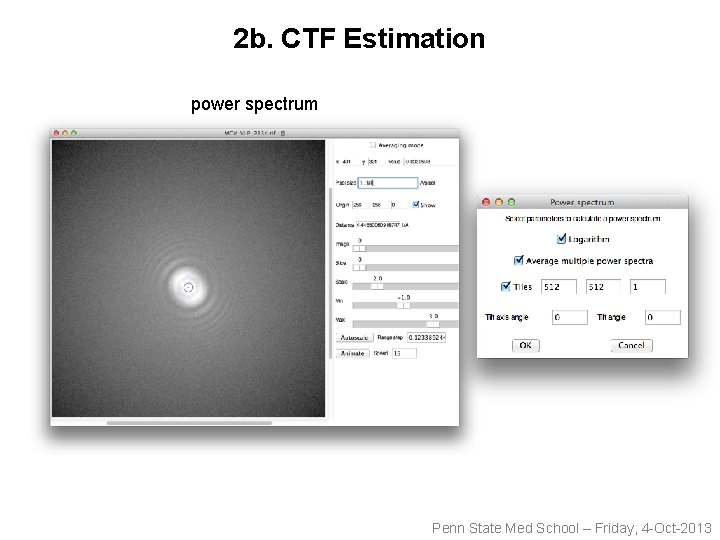
2 b. CTF Estimation power spectrum Penn State Med School – Friday, 4 -Oct-2013
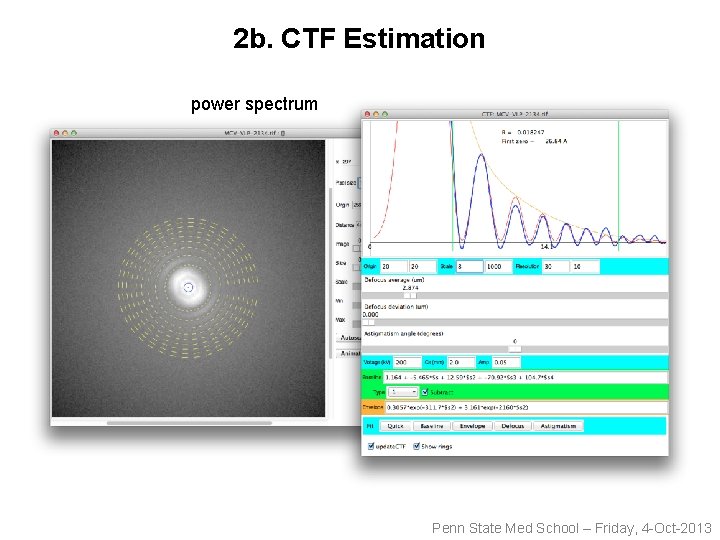
2 b. CTF Estimation power spectrum Penn State Med School – Friday, 4 -Oct-2013

2 a & b. Summary • Picked 579 biggies & 21 smallies from 1 micrograph (2134) • Repeat for other micrographs: ugraph 2134 2135 2136 2137 2138 Total Mean Big 579 508 528 490 540 2645 529 Smalltotal 21600 17525 30558 8498 12552 882733 17546 • Defocus estimates ugraph 2134 2135 2136 2137 2138 defocus 2. 77 2. 85 3. 08 3. 20 2. 87 • Continue with analysis of biggies… Penn State Med School – Friday, 4 -Oct-2013
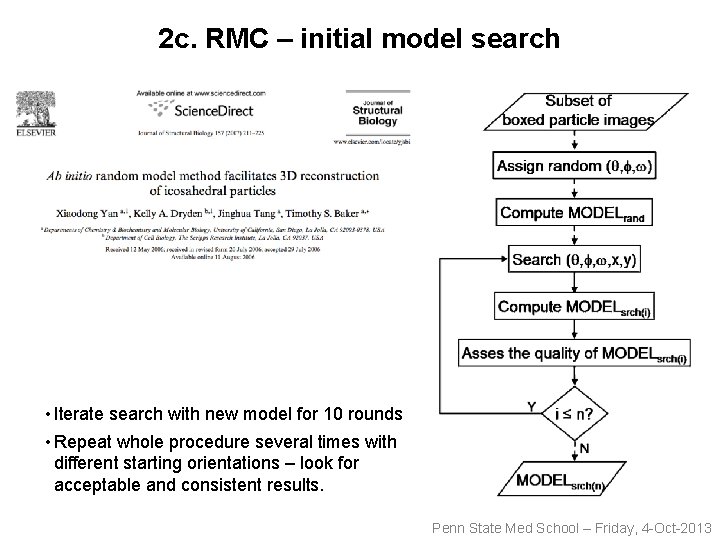
2 c. RMC – initial model search • Iterate search with new model for 10 rounds • Repeat whole procedure several times with different starting orientations – look for acceptable and consistent results. Penn State Med School – Friday, 4 -Oct-2013

2 c. RMC – initial model search conway% setup_rmc ------ SETUP_RMC version v 4. 03. 1 -----NAME setup_rmc - script for setting up random model calculation EXAMPLE setup_rmc -ncpu 4 -seed 123 -listfile setup_rmc -usedefaults DESCRIPTION All input specified using the syntax -key value. Defaults values are shown in brackets following descriptions. To use all default values, use the -usedefaults flag. 'n' = integer, 'f' = float, 's' = string -boxrad n image box size [obtained from image file] -fsc_nbins n number of bins for FSC calculation [50] -fsc_res_min f minimum resolution (A) for FSC calculation [60] -fsc_res_max f maximum resolution (A) for FSC calculation [30] -list s file or expression listing particle parameter files If not specified, then setup_rmc looks for (1) file named 'list' or (2) *00 n files in rundir -multi run multiple RMCs in parallel -ncpu n number of CPUs for each RMC computation[4] -nimages n number of images to use in constructing model [150] -nmodels n number of random models [10] -noctf turn off CTF corrections -nodefile s file containing list of compute nodes -res f highest resolution used in constructing map -rmax n maximum capsid radius -rmin n minimum capsid radius -rundir s directory containing particle parameter files [dat] -seed n seed for random number generator. Use an integer for reproducible result or -seed=time for seed based on seconds since 1/1/1970 [1] -symm_code n symmetry of reconstruction [532] -traditional RMC calculation (best map chosen using FSC) Penn State Med School – Friday, 4 -Oct-2013

2 c. RMC – initial model search conway% setup_rmc –nmodels 5 --- setup_rmc starting --setup_rmc parameters: Default rundir = dat Default maximum map resolution = 29 list not specified - setup_rmc will first check for file 'list', then look for files dat/*00 n Default ncpu = 4 Default fsc_nbins = 50 Default fsc_res_min = 60. 0 Default fsc_res_max = 30. 0 Default nimages = 150 User specified nmodels = 5 Default symm_code = 532 Default seed = 1 Note – 150 particles only Parameter files in dat considered for model construction: all 5. dat_000 Box radius from PIF/MRC file = 112 Instructions: Run RMC_run to construct starting model Run RMC_cleanup to remove temporary files after calculations complete Starting model will be found at dat/rmc. pif auto 3 dem input file auto-bin 2_master automatically generated --- setup_rmc done --conway% RMC_run Penn State Med School – Friday, 4 -Oct-2013

2 c. RMC – initial model search Trial 1 Iteration 0 0 1 2 3 4 5 6 7 8 9 10 Penn State Med School – Friday, 4 -Oct-2013
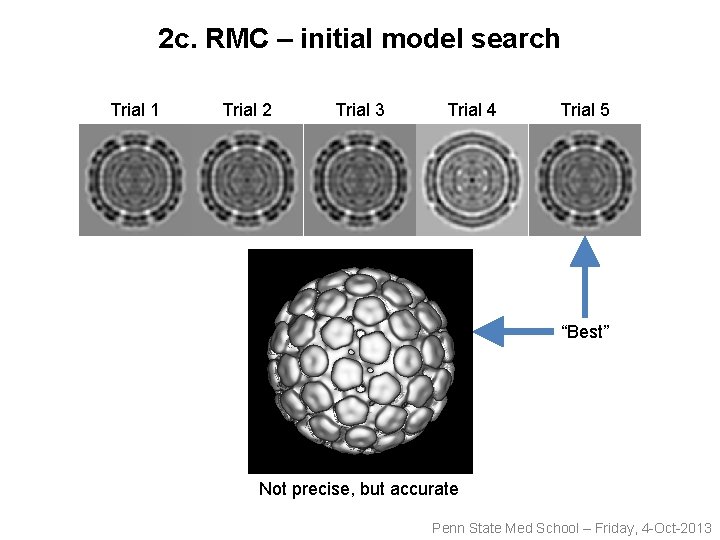
2 c. RMC – initial model search Trial 1 Trial 2 Trial 3 Trial 4 Trial 5 “Best” Not precise, but accurate Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem i. Initial orientation search – we have a model (RMC) – but no orientations auto mode search Uses PPFT ii. Refinement – improve orientations – improve resolution auto mode refine Uses POR global refine po 2 r mode global local refine po 2 r mode local Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem PPFT POR Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem i. Initial orientation search (PPFT) vs. • I usually run 10 iterations to ensure the model is decent and particle orientations have a chance to sample their correct values • Use coarse angle steps – want a quick answer, rough but accurate e. g. ppft delta_theta 2 ppft bin_factor 2 Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem i. Initial orientation search (PPFT) Projection from RMC map auto. . . ppft ppft ppft ppft mode search annulus_high annulus_low bin_factor ctfmode delta_theta filter_factor_1 input_mode pftrad_hi pftrad_lo pftrad_step resolution_high resolution_low temperature_factor 110 33 2 0. 1 2 112 1 1 30 124. 2 0 Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem i. Initial orientation search (PPFT) delta map itr mode estres angle undamp time cpu nptles ntot nmg defocus ------ ------ ------ --- ----# cycle 1. Start with RMC model (5), 5 iterations 1 s(2) 18. 48 2. 00 15. 60 13. 49 35 8 2639 2645 5 2. 77 -3. 20 2 s(2) 13. 07 2. 00 11. 56 10. 36 35 8 2638 2645 5 2. 77 -3. 20 3 s(2) 12. 58 2. 00 11. 17 10. 05 36 8 2638 2645 5 2. 77 -3. 20 4 s(2) 12. 64 2. 00 11. 22 10. 09 36 8 2638 2645 5 2. 77 -3. 20 5 s(2) 13. 59 2. 00 11. 97 10. 69 36 8 2639 2645 5 2. 77 -3. 20 RMC iter 1 iter 2 iter 3 iter 4 iter 5 150 p 2639 p 2638 p 2639 p Penn State Med School – Friday, 4 -Oct-2013
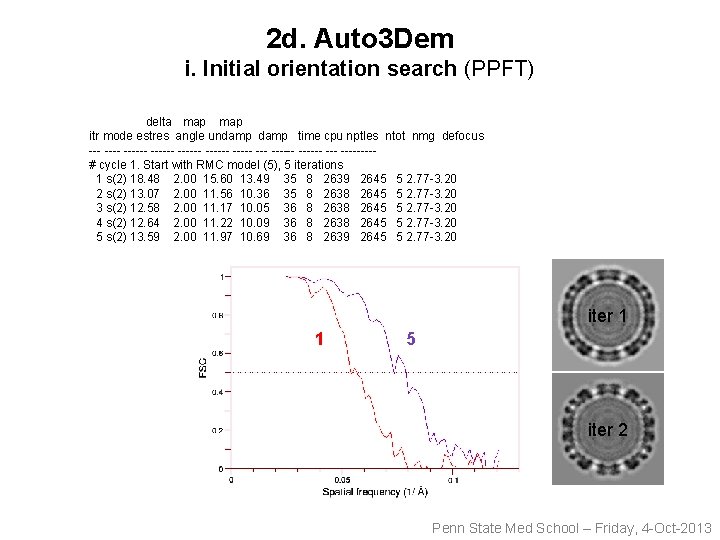
2 d. Auto 3 Dem i. Initial orientation search (PPFT) delta map itr mode estres angle undamp time cpu nptles ntot nmg defocus ------ ------ ------ --- ----# cycle 1. Start with RMC model (5), 5 iterations 1 s(2) 18. 48 2. 00 15. 60 13. 49 35 8 2639 2645 5 2. 77 -3. 20 2 s(2) 13. 07 2. 00 11. 56 10. 36 35 8 2638 2645 5 2. 77 -3. 20 3 s(2) 12. 58 2. 00 11. 17 10. 05 36 8 2638 2645 5 2. 77 -3. 20 4 s(2) 12. 64 2. 00 11. 22 10. 09 36 8 2638 2645 5 2. 77 -3. 20 5 s(2) 13. 59 2. 00 11. 97 10. 69 36 8 2639 2645 5 2. 77 -3. 20 iter 1 1 5 iter 2 Penn State Med School – Friday, 4 -Oct-2013
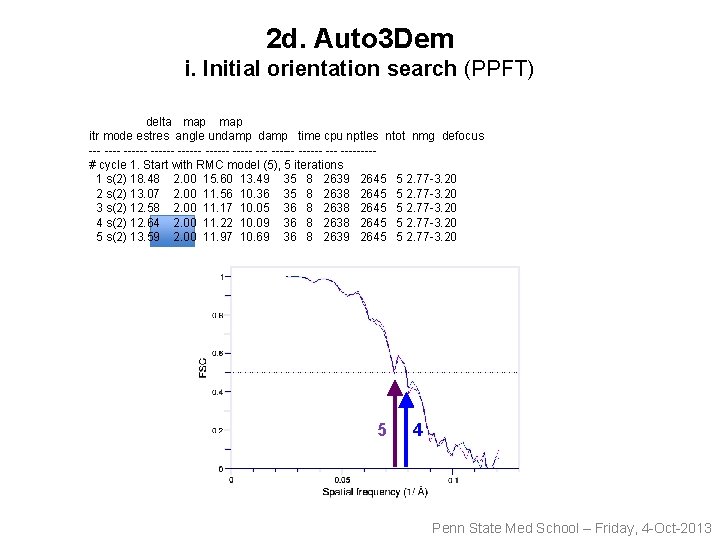
2 d. Auto 3 Dem i. Initial orientation search (PPFT) delta map itr mode estres angle undamp time cpu nptles ntot nmg defocus ------ ------ ------ --- ----# cycle 1. Start with RMC model (5), 5 iterations 1 s(2) 18. 48 2. 00 15. 60 13. 49 35 8 2639 2645 5 2. 77 -3. 20 2 s(2) 13. 07 2. 00 11. 56 10. 36 35 8 2638 2645 5 2. 77 -3. 20 3 s(2) 12. 58 2. 00 11. 17 10. 05 36 8 2638 2645 5 2. 77 -3. 20 4 s(2) 12. 64 2. 00 11. 22 10. 09 36 8 2638 2645 5 2. 77 -3. 20 5 s(2) 13. 59 2. 00 11. 97 10. 69 36 8 2639 2645 5 2. 77 -3. 20 5 4 Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem i. Initial orientation search (PPFT) RMC iter 5 Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem ii. Global orientation refine (POR mode global) • Better model, starting orientations for all particles • POR/global generally does a better job than PPFT • But…very slow • After PPFT, do 1 iteration of POR/global, coarse steps auto mode refine. . . # Iteration parameters auto iter_start auto niter. . . po 2 r ctfmode po 2 r dangle po 2 r dcenter po 2 r gangle po 2 r mode po 2 r nangle po 2 r ncenter po 2 r res_max po 2 r res_min po 2 r tempfac 6 1 2 2 3 3 global 4 4 12. 65 124. 2 0 delta map itr mode estres angle undamp time cpu nptles ntot nmg defocus ------ ------ ------ --- ----# cycle 1. Start with RMC model (5), 5 iterations 1 s(2) 18. 48 2. 00 15. 60 13. 49 35 8 2639 2645 5 2. 77 -3. 20 2 s(2) 13. 07 2. 00 11. 56 10. 36 35 8 2638 2645 5 2. 77 -3. 20 3 s(2) 12. 58 2. 00 11. 17 10. 05 36 8 2638 2645 5 2. 77 -3. 20 4 s(2) 12. 64 2. 00 11. 22 10. 09 36 8 2638 2645 5 2. 77 -3. 20 5 s(2) 13. 59 2. 00 11. 97 10. 69 36 8 2639 2645 5 2. 77 -3. 20 # cycle 2. Global POR, 1 iteration 6 r(g) 11. 34 3. 00 10. 18 9. 24 1766 4 1984 2645 5 2. 77 -3. 20 auto score_fraction 0. 75 Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem ii. Global orientation refine (POR mode global) delta map itr mode estres angle undamp time cpu nptles ntot nmg defocus ------ ------ ------ --- ----# cycle 1. Start with RMC model (5), 5 iterations 1 s(2) 18. 48 2. 00 15. 60 13. 49 35 8 2639 2645 5 2. 77 -3. 20 2 s(2) 13. 07 2. 00 11. 56 10. 36 35 8 2638 2645 5 2. 77 -3. 20 3 s(2) 12. 58 2. 00 11. 17 10. 05 36 8 2638 2645 5 2. 77 -3. 20 4 s(2) 12. 64 2. 00 11. 22 10. 09 36 8 2638 2645 5 2. 77 -3. 20 5 s(2) 13. 59 2. 00 11. 97 10. 69 36 8 2639 2645 5 2. 77 -3. 20 # cycle 2. Global POR, 1 iteration 6 r(g) 11. 34 3. 00 10. 18 9. 24 1766 4 1984 2645 5 2. 77 -3. 20 iter 5 iter 6 2639 p 1766 p Penn State Med School – Friday, 4 -Oct-2013
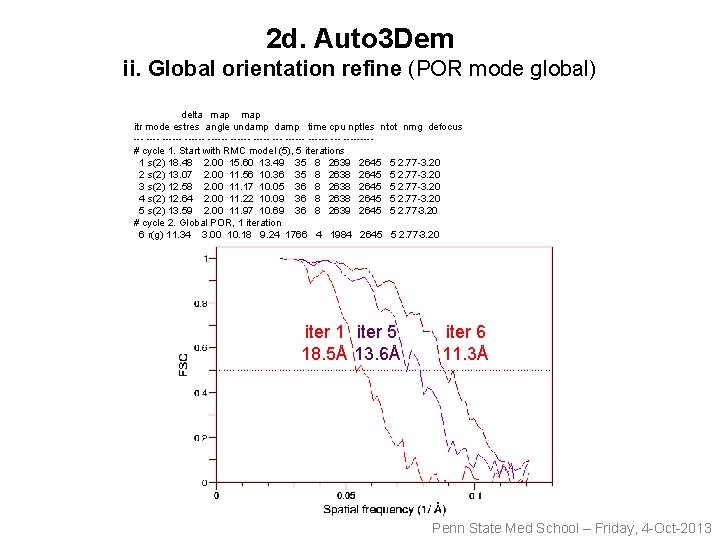
2 d. Auto 3 Dem ii. Global orientation refine (POR mode global) delta map itr mode estres angle undamp time cpu nptles ntot nmg defocus ------ ------ ------ --- ----# cycle 1. Start with RMC model (5), 5 iterations 1 s(2) 18. 48 2. 00 15. 60 13. 49 35 8 2639 2645 5 2. 77 -3. 20 2 s(2) 13. 07 2. 00 11. 56 10. 36 35 8 2638 2645 5 2. 77 -3. 20 3 s(2) 12. 58 2. 00 11. 17 10. 05 36 8 2638 2645 5 2. 77 -3. 20 4 s(2) 12. 64 2. 00 11. 22 10. 09 36 8 2638 2645 5 2. 77 -3. 20 5 s(2) 13. 59 2. 00 11. 97 10. 69 36 8 2639 2645 5 2. 77 -3. 20 # cycle 2. Global POR, 1 iteration 6 r(g) 11. 34 3. 00 10. 18 9. 24 1766 4 1984 2645 5 2. 77 -3. 20 iter 1 iter 5 18. 5Å 13. 6Å iter 6 11. 3Å Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem ii. Global orientation refine (POR mode global) delta map itr mode estres angle undamp time cpu nptles ntot nmg defocus ------ ------ ------ --- ----# cycle 1. Start with RMC model (5), 5 iterations 1 s(2) 18. 48 2. 00 15. 60 13. 49 35 8 2639 2645 5 2. 77 -3. 20 2 s(2) 13. 07 2. 00 11. 56 10. 36 35 8 2638 2645 5 2. 77 -3. 20 3 s(2) 12. 58 2. 00 11. 17 10. 05 36 8 2638 2645 5 2. 77 -3. 20 4 s(2) 12. 64 2. 00 11. 22 10. 09 36 8 2638 2645 5 2. 77 -3. 20 5 s(2) 13. 59 2. 00 11. 97 10. 69 36 8 2639 2645 5 2. 77 -3. 20 # cycle 2. Global POR, 1 iteration 6 r(g) 11. 34 3. 00 10. 18 9. 24 1766 4 1984 2645 5 2. 77 -3. 20 6 r(g) 11. 62 2. 00 10. 41 9. 43 5174 4 1984 2645 5 2. 77 -3. 20 • Iteration 6 repeated with a smaller angle (2° instead of 3°) • No significant change in resolution • Took 3 times longer to run (86 mins vs. 29 mins) • Don’t make your angle or center steps too small!! Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem ii. Local orientation refine (POR mode local) • Better model, better orientations • POR/local is faster by inspecting a local window of orientations • Do cycles of 5 iterations, reducing angle & center steps auto mode refine. . . # Iteration parameters auto iter_start auto niter. . . po 2 r ctfmode po 2 r dangle po 2 r dcenter po 2 r gangle po 2 r mode po 2 r nangle po 2 r ncenter po 2 r res_max po 2 r res_min po 2 r tempfac 7 5 2 2 3 3 local 4 4 11. 34 124. 2 0 delta map itr mode estres angle undamp time cpu nptles ntot nmg defocus ------ ------ ------ --- ----. . . # cycle 2. Global POR, 1 iteration 6 r(g) 11. 34 3. 00 10. 18 9. 24 1766 4 1984 2645 5 2. 77 -3. 20 # cycle 3. Local POR, 5 iterations 7 r(l) 10. 56 2. 00 9. 55 8. 72 70 4 1984 2645 5 2. 77 -3. 20 8 r(l) 10. 63 2. 00 9. 61 8. 77 38 4 1983 2645 5 2. 77 -3. 20 9 r(l) 10. 59 2. 00 9. 58 8. 74 38 4 1984 2645 5 2. 77 -3. 20 10 r(l) 10. 59 2. 00 9. 58 8. 74 37 4 1983 2645 5 2. 77 -3. 20 11 r(l) 10. 59 2. 00 9. 58 8. 74 38 4 1983 2645 5 2. 77 -3. 20 . . +8°. . . . +6°. . . . +4°. . . . +2°. . -8° -6° -4° -2° 0° +2° +4° +6° +8° (Φ). . -2°. . . . -4°. . . . -6°. . . . -8°. . (θ) Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem ii. Local orientation refine (POR mode local) delta map itr mode estres angle undamp time cpu nptles ntot nmg defocus ------ ------ ------ --- ----. . . # cycle 2. Global POR, 1 iteration 6 r(g) 11. 34 3. 00 10. 18 9. 24 1766 4 1984 2645 5 2. 77 -3. 20 # cycle 3. Local POR, 5 iterations 7 r(l) 10. 56 2. 00 9. 55 8. 72 70 4 1984 2645 5 2. 77 -3. 20 8 r(l) 10. 63 2. 00 9. 61 8. 77 38 4 1983 2645 5 2. 77 -3. 20 9 r(l) 10. 59 2. 00 9. 58 8. 74 38 4 1984 2645 5 2. 77 -3. 20 10 r(l) 10. 59 2. 00 9. 58 8. 74 37 4 1983 2645 5 2. 77 -3. 20 11 r(l) 10. 59 2. 00 9. 58 8. 74 38 4 1983 2645 5 2. 77 -3. 20 iter 6 iter 11 Penn State Med School – Friday, 4 -Oct-2013
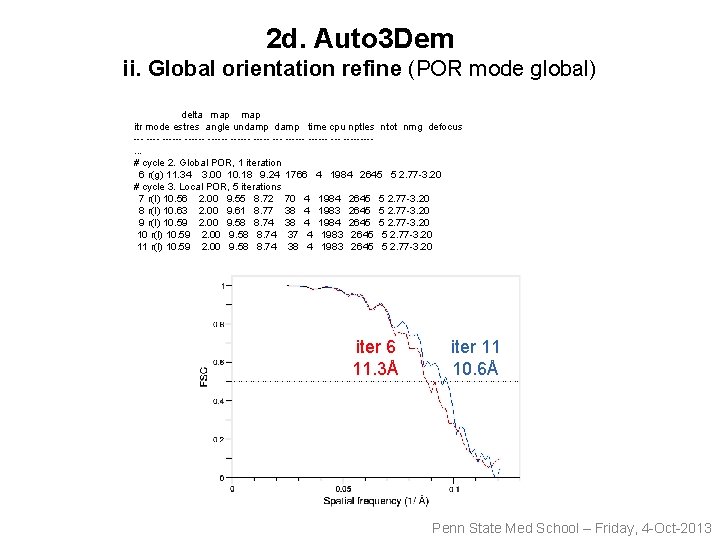
2 d. Auto 3 Dem ii. Global orientation refine (POR mode global) delta map itr mode estres angle undamp time cpu nptles ntot nmg defocus ------ ------ ------ --- ----. . . # cycle 2. Global POR, 1 iteration 6 r(g) 11. 34 3. 00 10. 18 9. 24 1766 4 1984 2645 5 2. 77 -3. 20 # cycle 3. Local POR, 5 iterations 7 r(l) 10. 56 2. 00 9. 55 8. 72 70 4 1984 2645 5 2. 77 -3. 20 8 r(l) 10. 63 2. 00 9. 61 8. 77 38 4 1983 2645 5 2. 77 -3. 20 9 r(l) 10. 59 2. 00 9. 58 8. 74 38 4 1984 2645 5 2. 77 -3. 20 10 r(l) 10. 59 2. 00 9. 58 8. 74 37 4 1983 2645 5 2. 77 -3. 20 11 r(l) 10. 59 2. 00 9. 58 8. 74 38 4 1983 2645 5 2. 77 -3. 20 iter 6 11. 3Å iter 11 10. 6Å Penn State Med School – Friday, 4 -Oct-2013
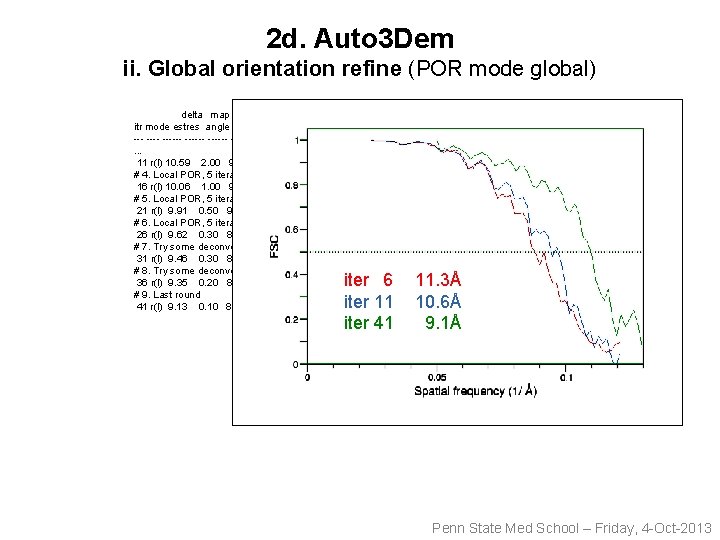
2 d. Auto 3 Dem ii. Global orientation refine (POR mode global) delta map itr mode estres angle undamp time cpu nptles ------ ------ ------ --- ----. . . 11 r(l) 10. 59 2. 00 9. 58 8. 74 38 4 1983 2645 # 4. Local POR, 5 iterations 16 r(l) 10. 06 1. 00 9. 14 8. 38 39 4 1983 2645 # 5. Local POR, 5 iterations 21 r(l) 9. 91 0. 50 9. 02 8. 27 40 4 1985 2645 # 6. Local POR, 5 iterations 26 r(l) 9. 62 0. 30 8. 77 8. 06 42 4 1983 2645 # 7. Try some deconvolution in P 3 DR 31 r(l) 9. 46 0. 30 8. 64 7. 95 45 4 1983 2645 # 8. Try some deconvolution in P 3 DR 36 r(l) 9. 35 0. 20 8. 55 7. 88 48 4 1983 2645 # 9. Last round 41 r(l) 9. 13 0. 10 8. 37 7. 72 63 4 1983 2645 ntot nmg defocus 5 2. 77 -3. 20 iter 562. 77 -3. 20 11. 3Å iter 11 10. 6Å 5 2. 77 -3. 20 iter 41 9. 1Å Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem ii. Global orientation refine (POR mode global) iter 11 iter 41 • Needs more particles: - reduce noise - increase resolution Penn State Med School – Friday, 4 -Oct-2013

2 d. Auto 3 Dem ii. Global orientation refine (POR mode global) • Previously bin-by-2 data (2. 76Å/pixel) • Un-binned data (1. 38Å/pixel) • FSC(0. 5) = 7. 0Å; 0. 26°; 11598 / 23248 p 50 µgraphs 100Å Penn State Med School – Friday, 4 -Oct-2013
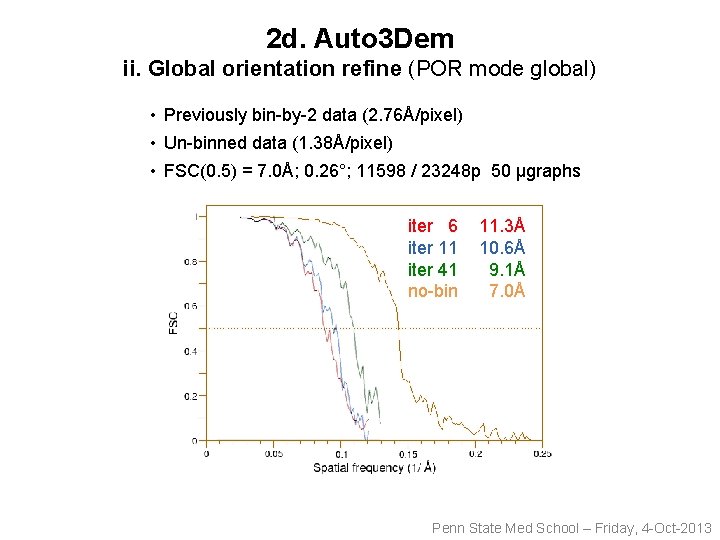
2 d. Auto 3 Dem ii. Global orientation refine (POR mode global) • Previously bin-by-2 data (2. 76Å/pixel) • Un-binned data (1. 38Å/pixel) • FSC(0. 5) = 7. 0Å; 0. 26°; 11598 / 23248 p 50 µgraphs iter 6 iter 11 iter 41 no-bin 11. 3Å 10. 6Å 9. 1Å 7. 0Å Penn State Med School – Friday, 4 -Oct-2013

2 d. Resolution FSC = Fourier Shell Correlation • Correlation between two ½-dataset density maps in Fourier (reciprocal) space DPR SSNR • Measure of consistency vs spatial frequency (spacing) • Consistency breaks down when common signal stops Problems • ½-dataset density maps inferior to full dataset map • Correlation limit – which? 0. 5, 0. 3, 0. 143, … • Consistency may be extended by systematic error including model bias “Gold Standard” (Henderson et al, Structure 20, 2012) • ½-datasets analysed independently (RMC onwards) • Correlation limit of 0. 143 Penn State Med School – Friday, 4 -Oct-2013

2 d. Resolution Over-fitting Solid – ½ datasets Black – “gold standard” Dashed – full dataset vs xtal map Grey – “classic” Penn State Med School – Friday, 4 -Oct-2013
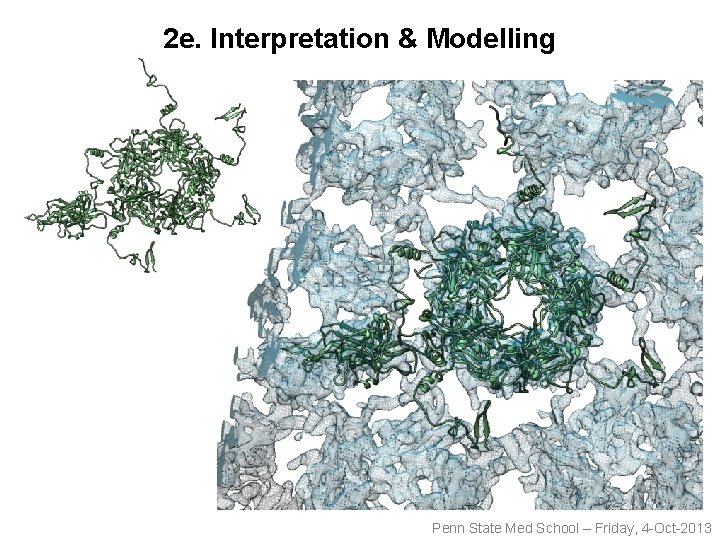
2 e. Interpretation & Modelling Penn State Med School – Friday, 4 -Oct-2013

Summary Recipe for RMC/Auto 3 Dem • RMC, check sections • Auto 3 Dem – search, 10 iterations: ppft delta_theta 2 (or 3) • Auto 3 Dem – refine (g), 1 iteration: po 2 r gangle 2 (or 3) • Auto 3 Dem – refine (l), 5 iterations: po 2 r dangle dcenter 1. 2 2– 5 • Match dangle and dcenter to current resolution eg: 300Å radius particle, 1° ≈ 1°*pi/180*300 5 Å at edge So at 20Å, refine at 1° angle steps 5 Å center steps (change to pixels) • Repeat refine (local) and adjust dangle and dcenter down to match new resolution • Use a realistic fraction of particles • Use phase flipping (ctfmode 2) especially in po 2 r • Use deconvolution (ctfmode 1) with modest tempfac in p 3 dr • Run p 3 dr “by hand” to optimize data limits & tempfac Penn State Med School – Friday, 4 -Oct-2013
- Slides: 45