Crossgenome Assembly Scaffolding using Crossspecies Synteny Zemin Ning

Cross_genome: Assembly Scaffolding using Cross-species Synteny Zemin Ning High Performance Assembly
Can synteny help? And How? Scaffolding Contig gap closure
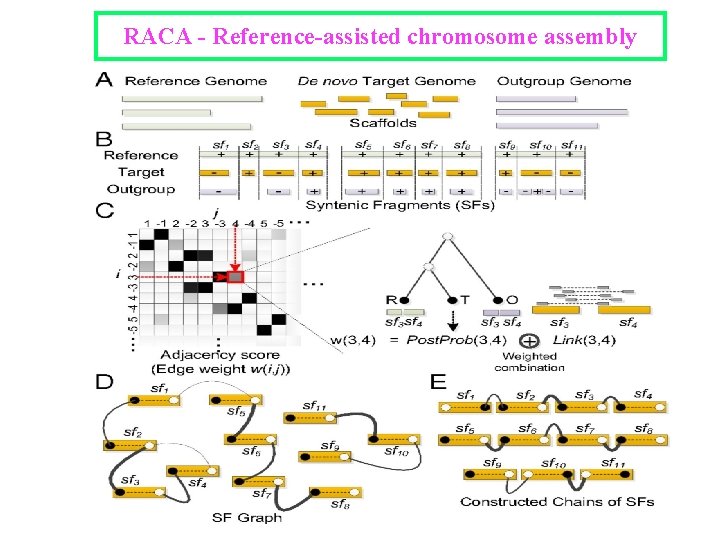
RACA - Reference-assisted chromosome assembly
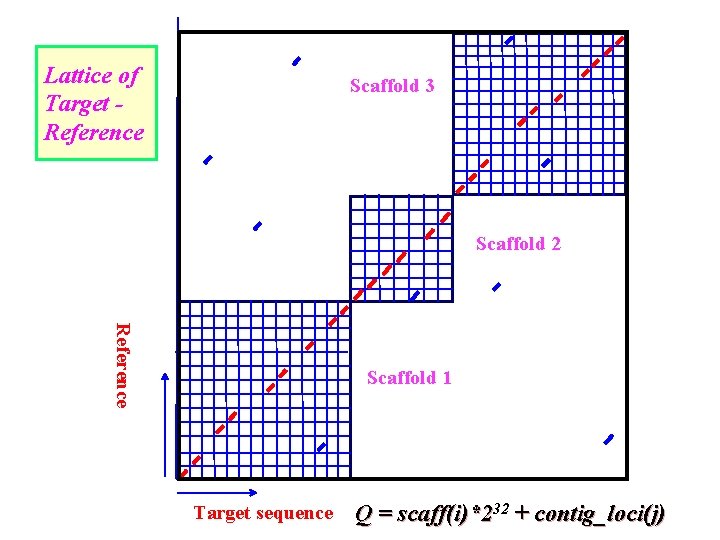
Lattice of Target Reference Scaffold 3 Scaffold 2 Reference Scaffold 1 Target sequence Q = scaff(i)*232 + contig_loci(j)
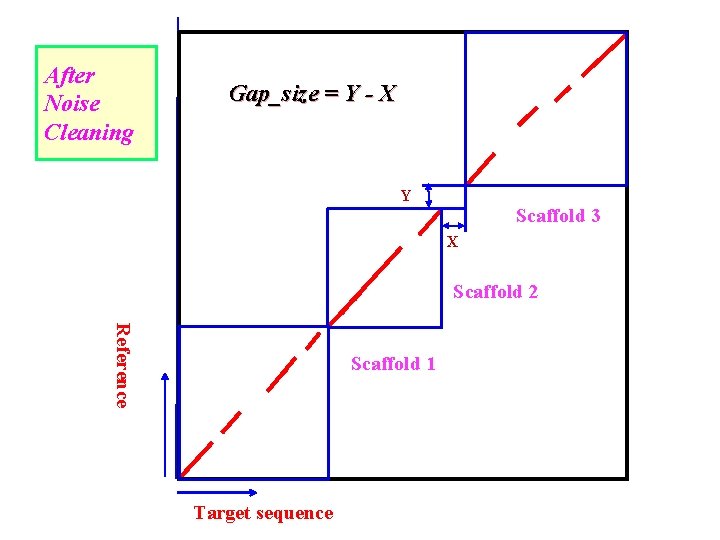
After Noise Cleaning Gap_size = Y - X Y Scaffold 3 X Scaffold 2 Reference Scaffold 1 Target sequence
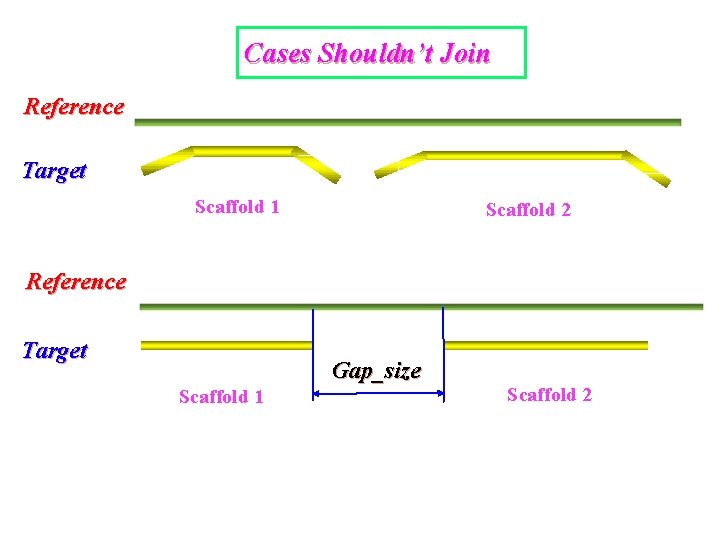
Cases Shouldn’t Join Reference Target Scaffold 1 Scaffold 2 Reference Target Gap_size Scaffold 1 Scaffold 2
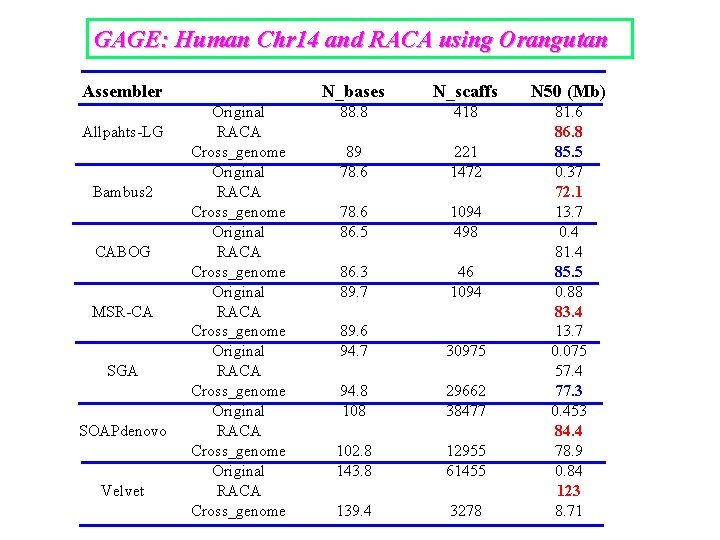
GAGE: Human Chr 14 and RACA using Orangutan Assembler Allpahts-LG Bambus 2 CABOG MSR-CA SGA SOAPdenovo Velvet Original RACA Cross_genome Original RACA Cross_genome N_bases N_scaffs N 50 (Mb) 88. 8 418 89 78. 6 221 1472 78. 6 86. 5 1094 498 86. 3 89. 7 46 1094 89. 6 94. 7 30975 94. 8 108 29662 38477 102. 8 143. 8 12955 61455 139. 4 3278 81. 6 86. 8 85. 5 0. 37 72. 1 13. 7 0. 4 81. 4 85. 5 0. 88 83. 4 13. 7 0. 075 57. 4 77. 3 0. 453 84. 4 78. 9 0. 84 123 8. 71

Scaffold N 50 for Other Genome Assemblies Original Cross_g References Panda 1. 3 Mb 25 Mb Dog, Human Tibetan Antelope 2. 6 Mb 42 Mb Tasmanian Devil 1. 8 Mb 6. 8 Mb Cattle, Dog, Human Opossum Availability ftp: //ftp. sanger. ac. uk/pub/users/zn 1/merge/cross_genome/
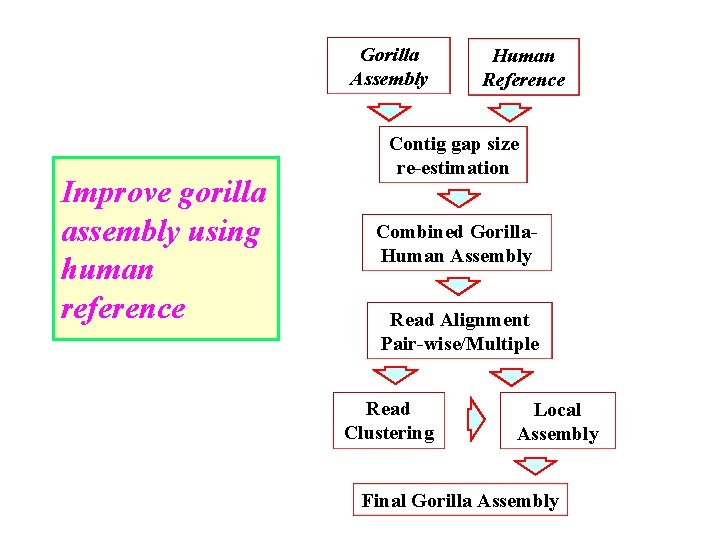
Gorilla Assembly Improve gorilla assembly using human reference Human Reference Contig gap size re-estimation Combined Gorilla. Human Assembly Read Alignment Pair-wise/Multiple Read Clustering Local Assembly Final Gorilla Assembly
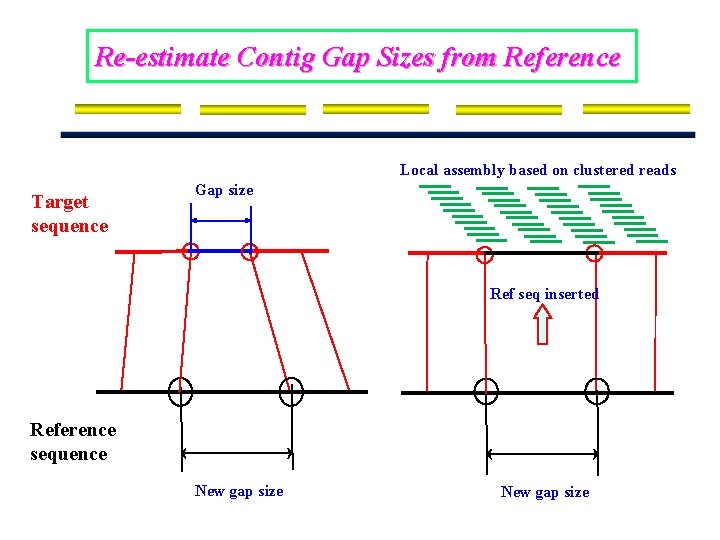
Re-estimate Contig Gap Sizes from Reference Local assembly based on clustered reads Target sequence Gap size Ref seq inserted Reference sequence New gap size
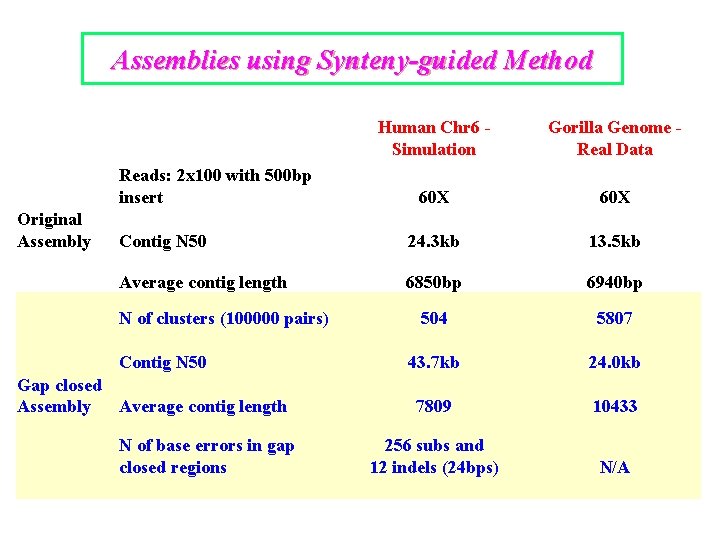
Assemblies using Synteny-guided Method Human Chr 6 Simulation Gorilla Genome Real Data 60 X Contig N 50 24. 3 kb 13. 5 kb Average contig length 6850 bp 6940 bp 504 5807 43. 7 kb 24. 0 kb 7809 10433 256 subs and 12 indels (24 bps) N/A Reads: 2 x 100 with 500 bp insert Original Assembly N of clusters (100000 pairs) Contig N 50 Gap closed Assembly Average contig length N of base errors in gap closed regions

Gorilla - Merge with other De novo Assemblies Original assembly (dev 5) Merge with Fermi* Merge with Masurca+ 13. 5 kb 30. 2 kb 53. 1 kb Average length 6850 12577 18768 Largest contig 215 kb 391. 2 kb 448. 8 kb 0 182661 257167 Contig N 50 N of gaps closed *Fermi assembler: https: //github. com/lh 3/fermi/ +Masurca assembler: http: //www. genome. umd. edu/masurca. html
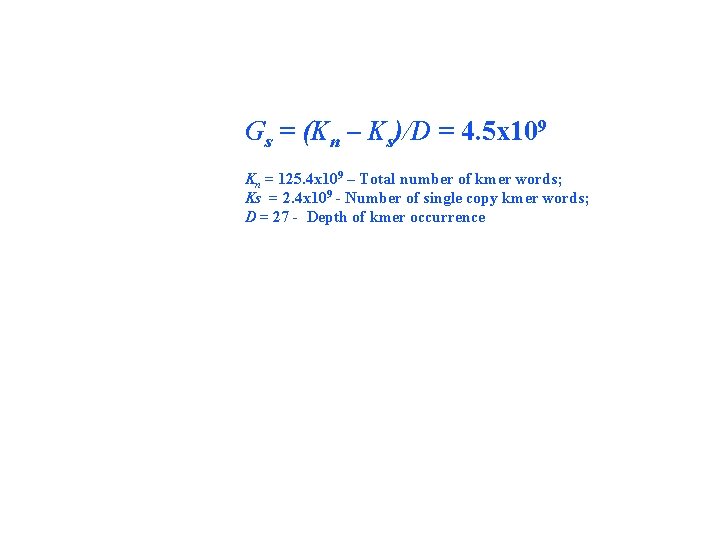
Gs = (Kn – Ks)/D = 4. 5 x 109 Kn = 125. 4 x 109 – Total number of kmer words; Ks = 2. 4 x 109 - Number of single copy kmer words; D = 27 - Depth of kmer occurrence

Original Contig (query) against New Assembly after Contig Break
Alignment Inconsistency

Original Contig (query) against New Assembly after Contig Break
Alignment Inconsistency
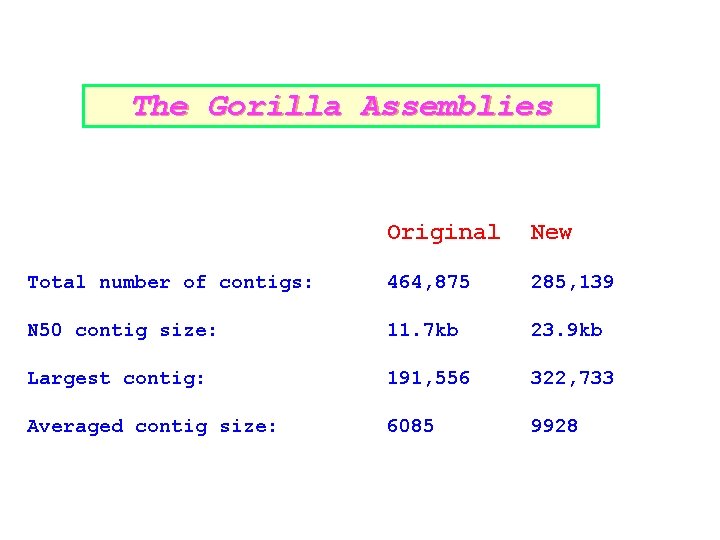
The Gorilla Assemblies Original New Total number of contigs: 464, 875 285, 139 N 50 contig size: 11. 7 kb 23. 9 kb Largest contig: 191, 556 322, 733 Averaged contig size: 6085 9928

Acknowledgements: q q q Hanness Ponstingl Frank Liu – Nanjing University of Information Technology (NUIT) Yan Li – (NUIT) Gorilla genome sequencing data BGI – Panda and Tibetan Antelope assemblies
- Slides: 19