CRG GENCODE Andrea Tanzer Thomas Derrien Cambridge September

CRG GENCODE Andrea Tanzer Thomas Derrien Cambridge, September 2011

CRG contributions to GENCODE • Contribution to the pipeline – U 12 introns – Selenoproteins • Experimental verification • RGASP • Long non-coding RNAs
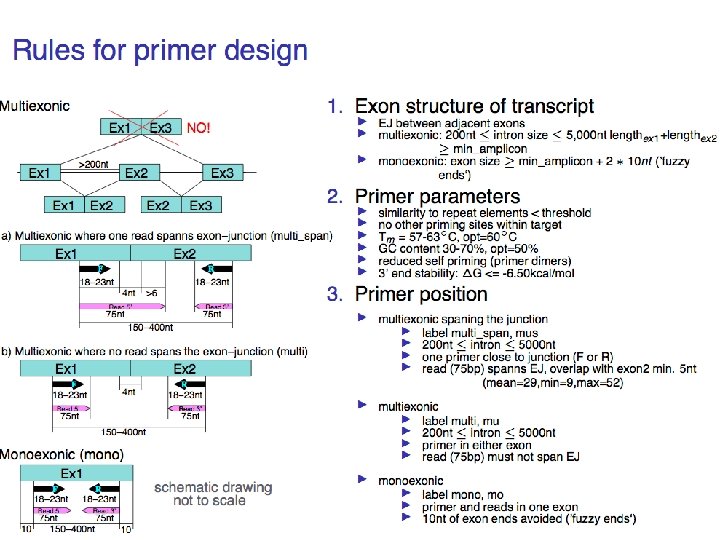
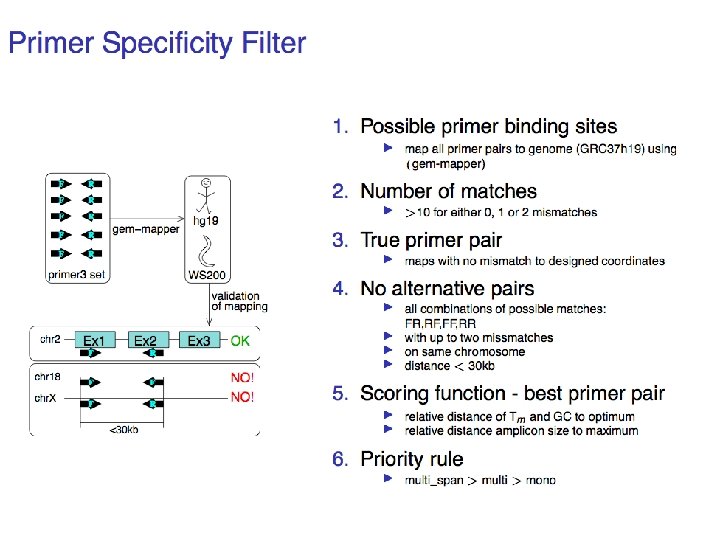

Experimental verification so far
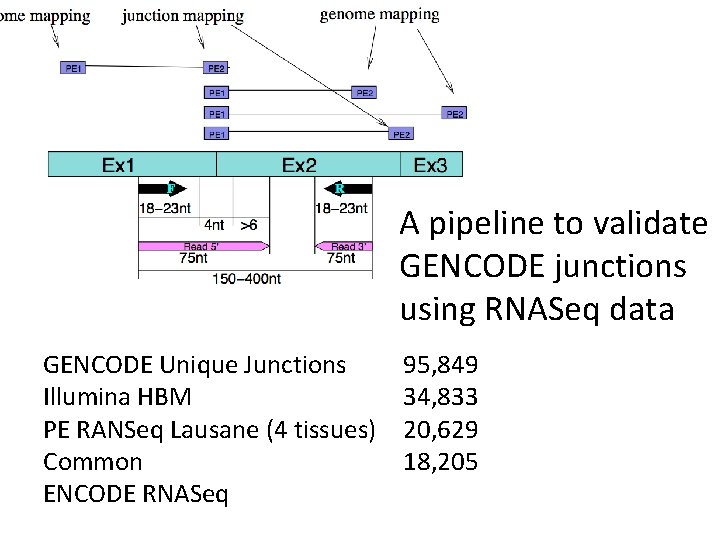
A pipeline to validate GENCODE junctions using RNASeq data GENCODE Unique Junctions Illumina HBM PE RANSeq Lausane (4 tissues) Common ENCODE RNASeq 95, 849 34, 833 20, 629 18, 205

RGASP • RGASP 1, 2. Hinxton • RGASP 3. Barcelona
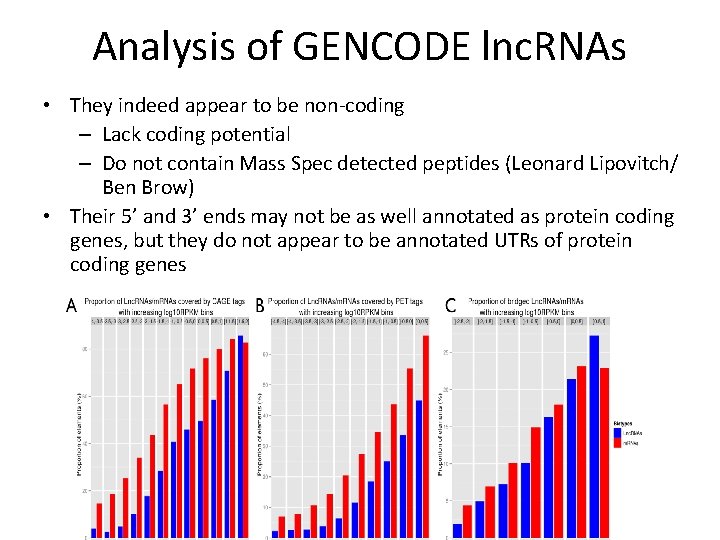
Analysis of GENCODE lnc. RNAs • They indeed appear to be non-coding – Lack coding potential – Do not contain Mass Spec detected peptides (Leonard Lipovitch/ Ben Brow) • Their 5’ and 3’ ends may not be as well annotated as protein coding genes, but they do not appear to be annotated UTRs of protein coding genes

Analysis of GENCODE lnc. RNAs • They are regulated by similar transcriptional, epigenetic and splicing pathways as proteincoding genes. • They are under clear negative evolutionary selection, but their exonic sequence is evolving more rapidly than coding m. RNAs. • They are expressed throughout organs and tissues in a highly specific way, but at uniformly lower levels than coding m. RNAs. • They display strong positive correlation with neighbouring protein-coding genes.
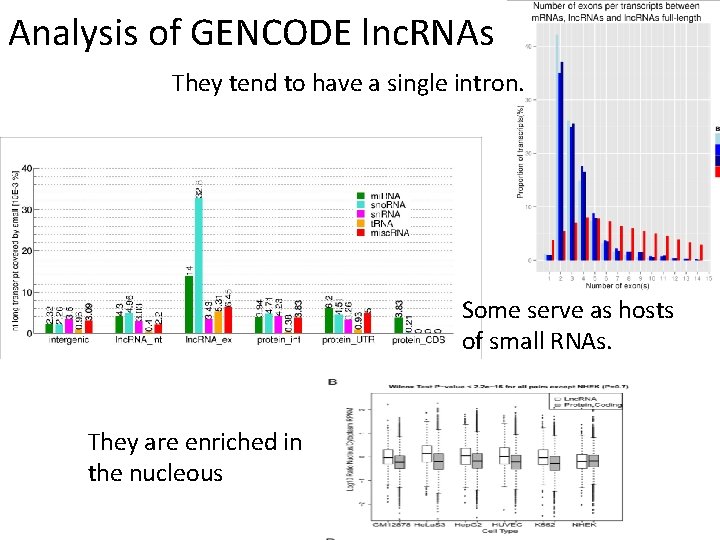
Analysis of GENCODE lnc. RNAs They tend to have a single intron. Some serve as hosts of small RNAs. They are enriched in the nucleous
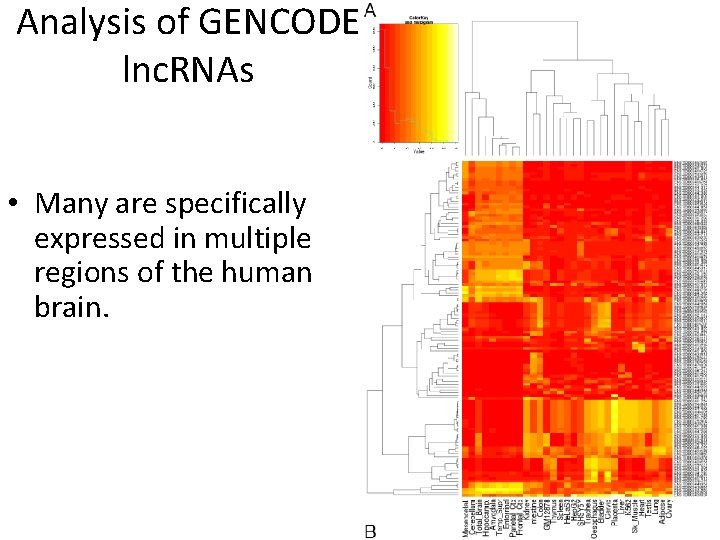
Analysis of GENCODE lnc. RNAs • Many are specifically expressed in multiple regions of the human brain.
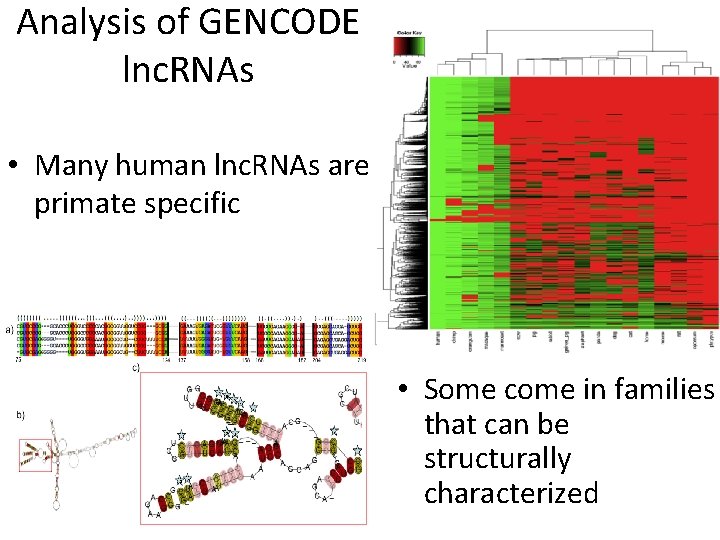
Analysis of GENCODE lnc. RNAs • Many human lnc. RNAs are primate specific • Some come in families that can be structurally characterized

Future plans. 2012

1. Ongoing work on characterization of lnc. RNAs • Development of methods to map lnc. RNAs to other genomes (mouse) • Collaborate with Havana on their chategorization and delineation • Pseudogenes of lnc. RNAs? 2. Ongoing work on experimental validation of GENCODE models • focus on lnc. RNA 5’ and 3’ ends?

3. From RNASeq to GENCODE • Assess methods to build gene models from RNASeq • Develop a pipeline (computational experimenta) to infer full length RNA sequences from RNASeq data – Cufflinks models and related plus experimental verification (RT-PCR, 5’ 3’ RACE) – Use RNASeq to nucleate computational gene modelling (using geneid) – Use RNASeq computational derived models (contigs, transcripts) to design exon capture microarrays followed by Pac. Bio sequencing. – Etc.
- Slides: 15