CRAM referencebased compression format developed by Vadim Zalunin

CRAM: reference-based compression format developed by Vadim Zalunin EBI is an Outstation of the European Molecular Biology Laboratory.

Data horror EMBL-EBI 10 petabytes SRA ~1 petabytes Over 2 million DVDs or 2. 5 km Complete Genomics 0. 5 TB for a single file
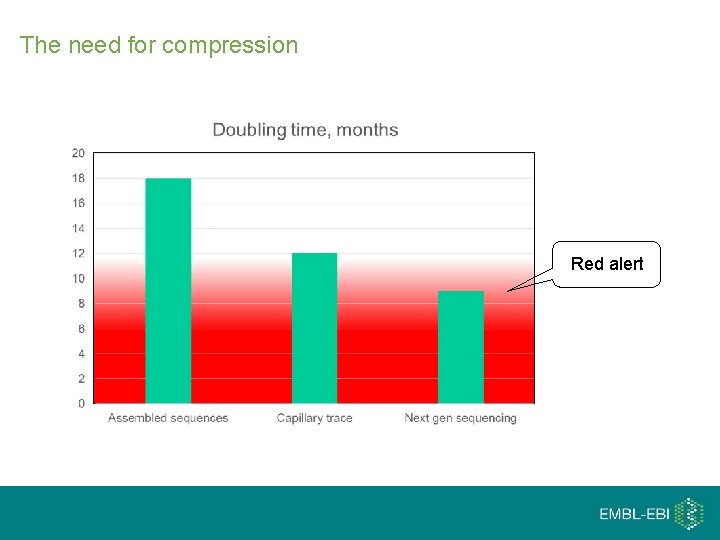
The need for compression Red alert
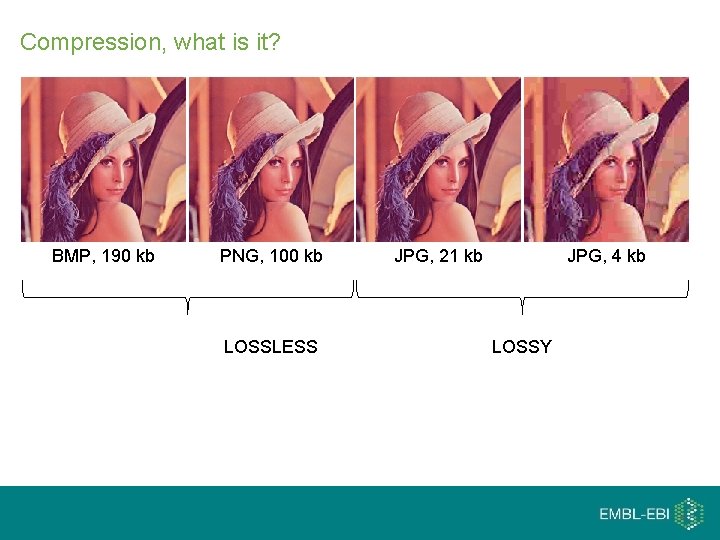
Compression, what is it? BMP, 190 kb PNG, 100 kb LOSSLESS JPG, 21 kb JPG, 4 kb LOSSY
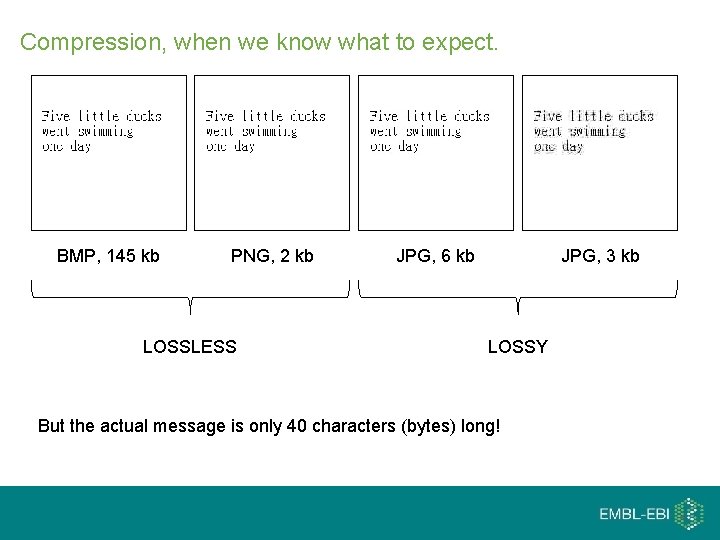
Compression, when we know what to expect. BMP, 145 kb PNG, 2 kb LOSSLESS JPG, 6 kb JPG, 3 kb LOSSY But the actual message is only 40 characters (bytes) long!

Compression at it’s best compress IMAGE, 145 kb "Five little ducks went swimming one day" uncompress TEXT, 40 b ~3500 times more efficient IMAGE, 145 kb
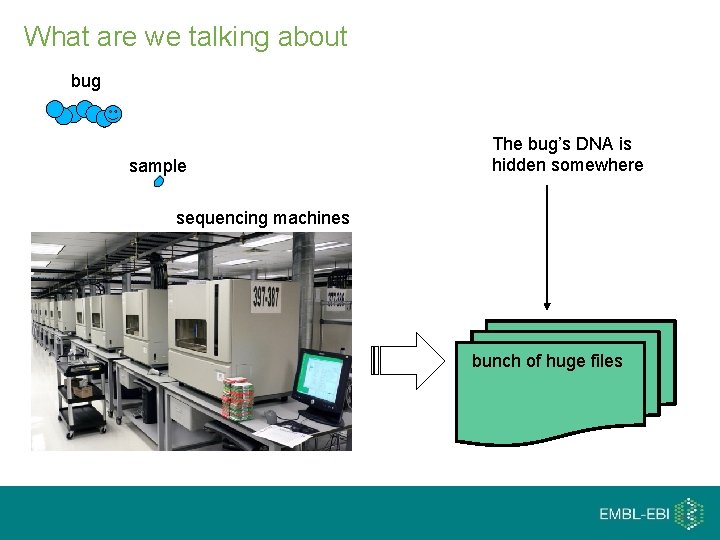
What are we talking about bug sample The bug’s DNA is hidden somewhere sequencing machines bunch of huge files
Looking closer at the data bunch of huge files It boils down to a long list of reads: read 1 read 2 read 3 …. . read bizzilion Each read represents a short nucleotide sequence from the genome. Additional information may be attached to it, for example error estimates.

What is a Read? @SRR 081241. 20758946 CCAGATCCTGGCCCTAAACAGGTGGTAAGGAGAGAGTG… + IDCEFFGGHHGGGHIGIHGFEFCFFDDGFFGIIHHIGIHHFI… An excerpt from of a FASTQ file.

What is a Read? read name @SRR 081241. 20758946 CCAGATCCTGGCCCTAAACAGGTGGTAAGGAGAGAGTG… + IDCEFFGGHHGGGHIGIHGFEFCFFDDGFFGIIHHIGIHHFI… An excerpt from of a FASTQ file.
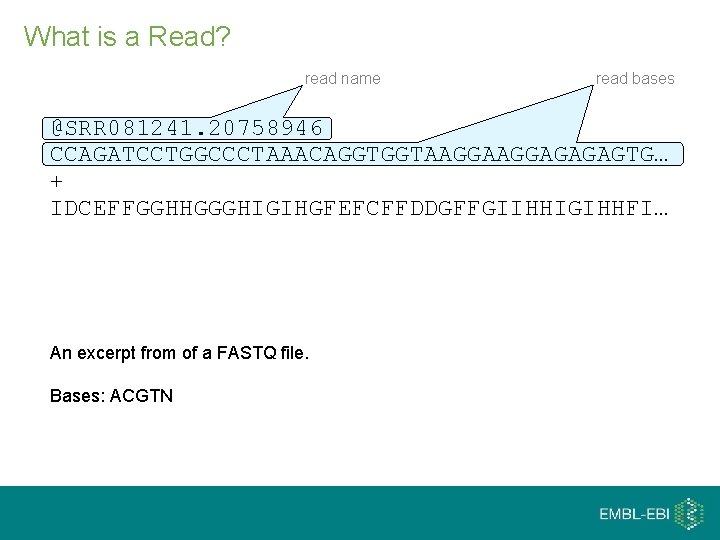
What is a Read? read name read bases @SRR 081241. 20758946 CCAGATCCTGGCCCTAAACAGGTGGTAAGGAGAGAGTG… + IDCEFFGGHHGGGHIGIHGFEFCFFDDGFFGIIHHIGIHHFI… An excerpt from of a FASTQ file. Bases: ACGTN

What is a Read? read name read bases @SRR 081241. 20758946 CCAGATCCTGGCCCTAAACAGGTGGTAAGGAGAGAGTG… + IDCEFFGGHHGGGHIGIHGFEFCFFDDGFFGIIHHIGIHHFI… read quality scores An excerpt from of a FASTQ file. Bases: ACGTN Quality scores: from ‘!’ (ASCII 33) to ‘~’ (ASCII 126)

What is quality score? Then quality score is phred quality score encoded as ASCII symbols 33 -126. Basically: higher scores are better, so ‘!’ is bad, ‘I’ is good.
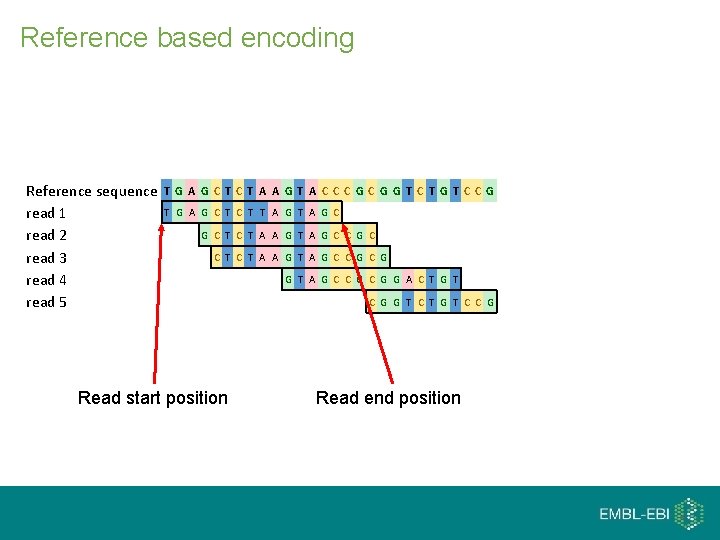
Reference based encoding Reference sequence read 1 read 2 read 3 read 4 read 5 T G A G C T A A G T A C CC G G T C T G T C C G T G A G C T T A G C G C T C T A A G T A G C C G C G Read start position G T A G C C G G A C T G T C G G T C T G T C C G Read end position
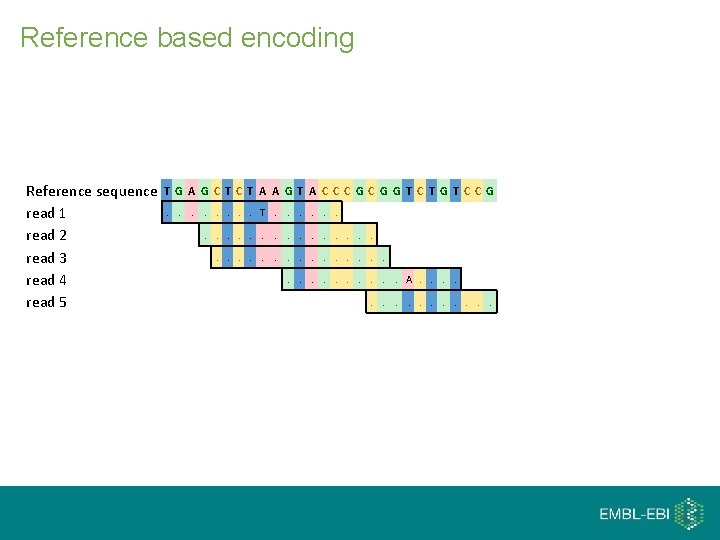
Reference based encoding Reference sequence read 1 read 2 read 3 read 4 read 5 T G A G C T A A G T A C CC G G T C T G T C C G. . . . T. . . A. . . .
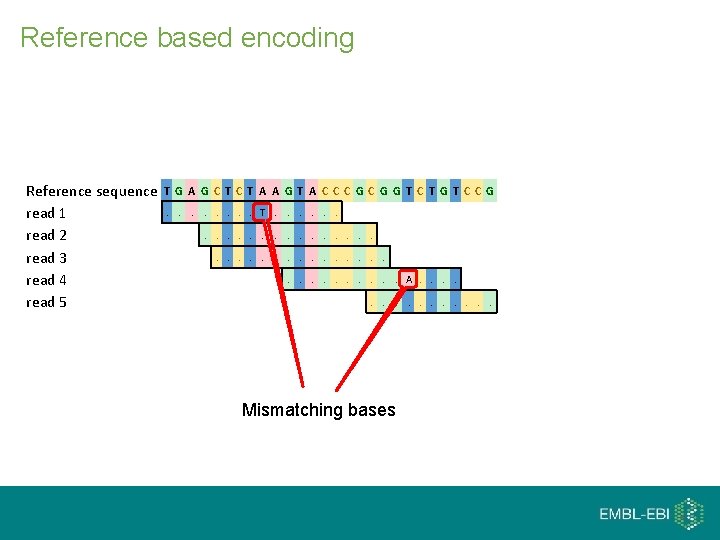
Reference based encoding Reference sequence read 1 read 2 read 3 read 4 read 5 T G A G C T A A G T A C CC G G T C T G T C C G. . . . T. . . A. . . . Mismatching bases

Lossy quality scores Approach 1 Quality scores are usually values from 0 to 39. tal zon i r o h For example: preserve only quality scores for mismatching bases. vertic Approach 2 Let’s treat quality scores using alignment information. al Let’s shrink them, so that they are from 0 to 7 now.

Comparison study: 1 K Genomes exomes BAM compress CRAM uncompress BAM
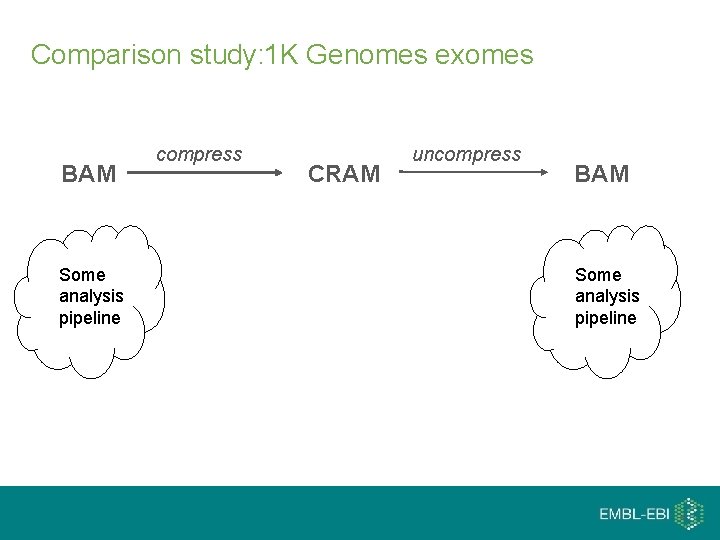
Comparison study: 1 K Genomes exomes BAM Some analysis pipeline compress CRAM uncompress BAM Some analysis pipeline
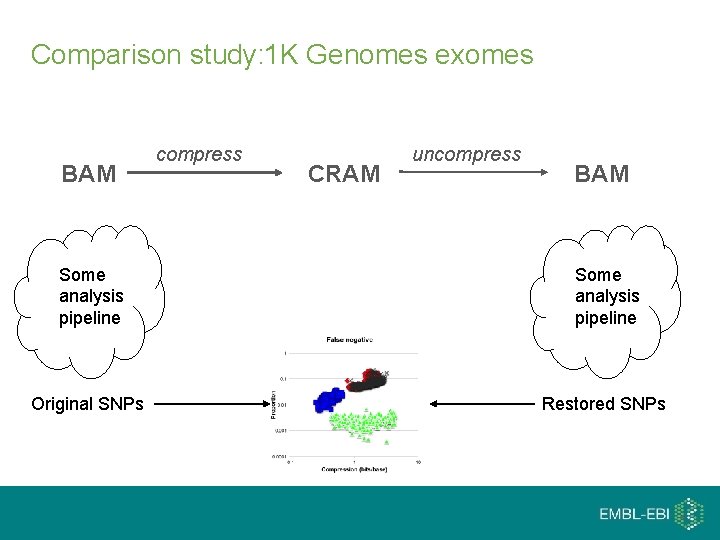
Comparison study: 1 K Genomes exomes BAM compress CRAM uncompress BAM Some analysis pipeline Original SNPs Restored SNPs
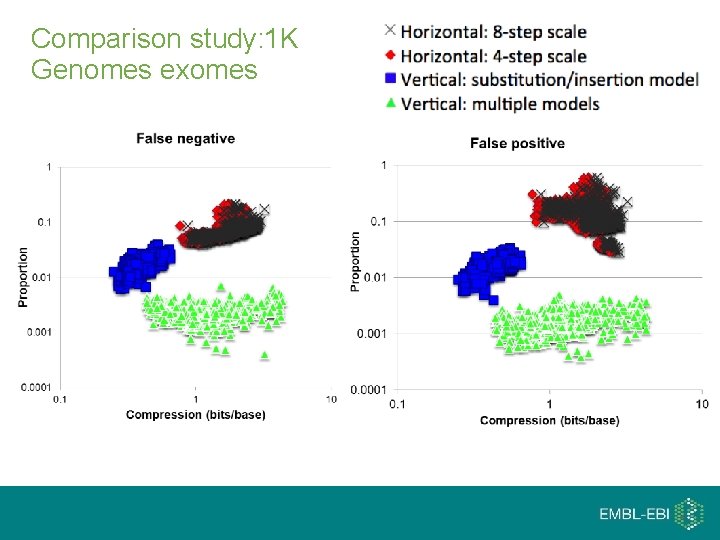
Comparison study: 1 K Genomes exomes
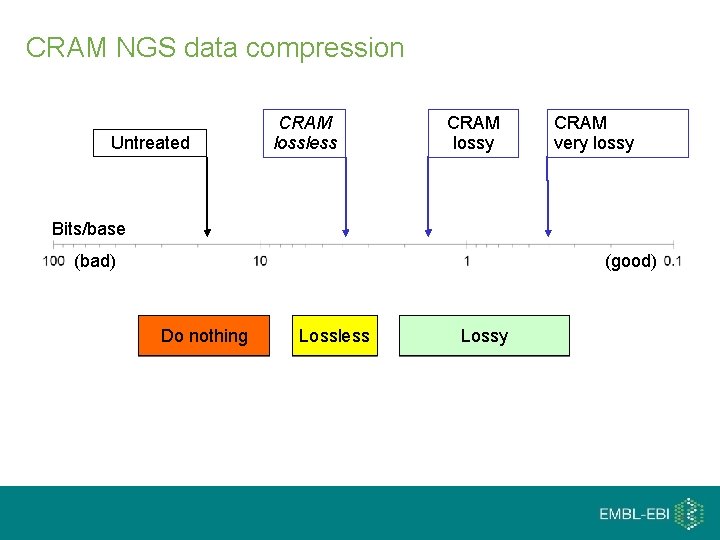
CRAM NGS data compression Untreated CRAM lossless CRAM lossy CRAM very lossy Bits/base (bad) (good) Do nothing Lossless Lossy
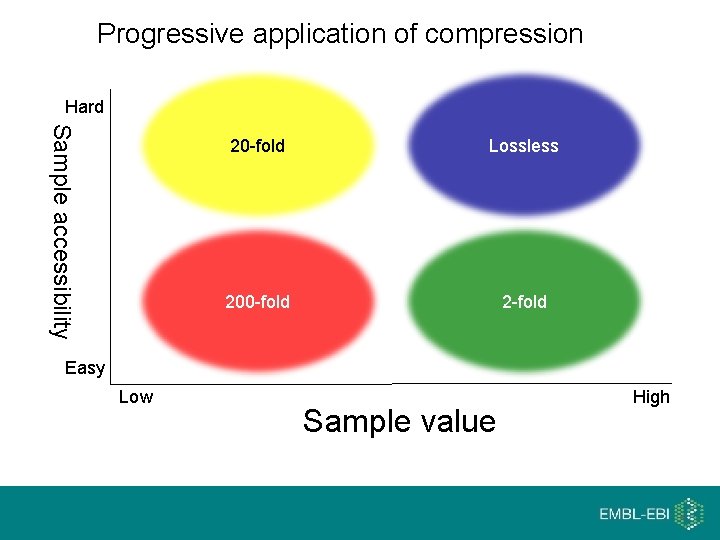
Progressive application of compression Hard Sample accessibility 20 -fold Lossless 200 -fold 2 -fold Easy Low Sample value High

References More information: http: //www. ebi. ac. uk/ena/about/cram_toolkit Mailing list: http: //listserver. ebi. ac. uk/mailman/listinfo/cram-dev Publications: Fritz, M. H. Leinonen, R. , et al. (2011) Efficient storage of high throughput DNA sequencing data using reference-based compression. Genome Res. 21 (5), 734 -40 Cochrane G. , Cook C. E. and Birney E. (2012) The future of DNA sequence archiving. Gigascience 1
- Slides: 24