CPAS Comparative Proteomics Analysis System Adam Rauch Lab

CPAS Comparative Proteomics Analysis System Adam Rauch Lab. Key Software adam@labkey. com

Agenda • CPAS general overview • CPAS for tandem mass spectrometry • Hands on demo

What Is CPAS? • Easy-to-use, web-based system for managing project data and experimental results. Key goals: – Provide universal access to data and support collaboration – Handle high-throughput processing and analysis of results – Keep data secure • A scalable system running at FHCRC… – Analysis cluster pipeline with massive disk storage – Java application server running on Apache Tomcat – A separate SQL database server (MS SQL Server or Postgre. SQL) • …and many other institutions – Open source release in October, 2005 – Over 200 institutions have downloaded the system • A work in progress: features are being added every day

What Does CPAS Provide? • Project management and group communications – All data is organized in a hierarchy of projects & folders – Project administrators choose what type of data is shown in their project and who has permission to see it – Tools include wiki, message board, issues list, contact management • Storage and analysis of experimental data – Tandem mass spectrometry results – Sample and mouse tracking – Experimental annotations – Flow cytometry data – Clinical study data – Other data types will be added over time

CPAS Project Management • • Great way to organize and manage group efforts Separate projects for each lab, initiative, or group Project administrators determine layout & permissions Group communications tools include: – Message board • Users can contribute to threaded conversations • Messages can include formatted text and attachments • Used for meeting agendas & minutes, posting documents & announcements – Wiki • Simple way to add static, formatted text & documents to a page • Used for welcome messages, schedules, protocols, instructions – Issues list • Structured way to track tasks, issues, and decisions in a project • Can assign owners, priorities, and due dates to each issue • Keeps a permanent history of all discussion
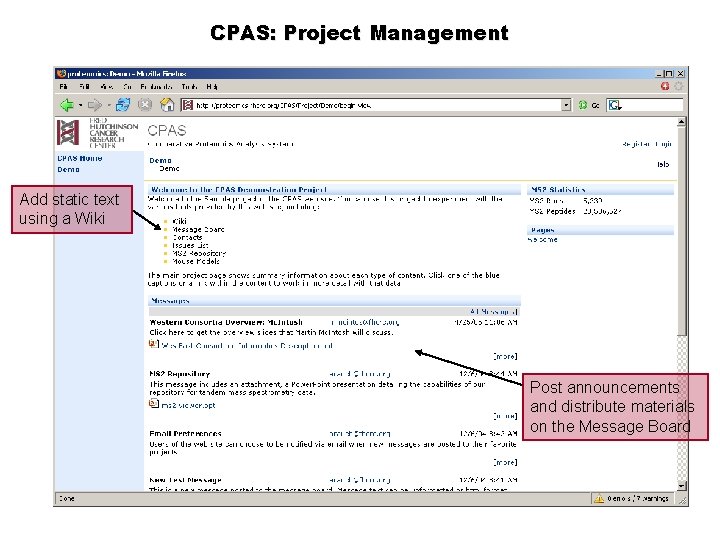
CPAS: Project Management Add static text using a Wiki Post announcements and distribute materials on the Message Board
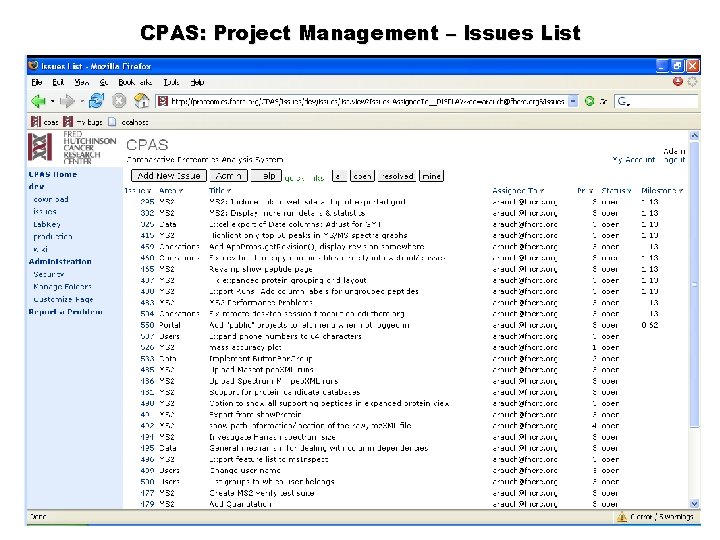
CPAS: Project Management – Issues List
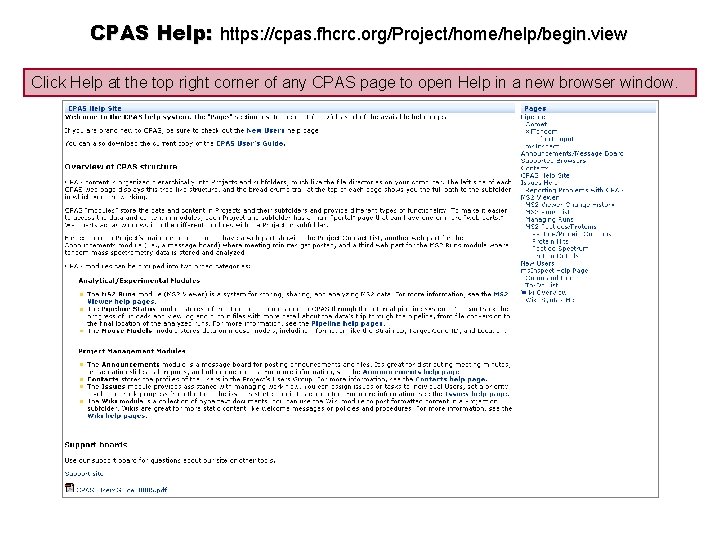
CPAS Help: https: //cpas. fhcrc. org/Project/home/help/begin. view Click Help at the top right corner of any CPAS page to open Help in a new browser window.

MS 2 Viewer
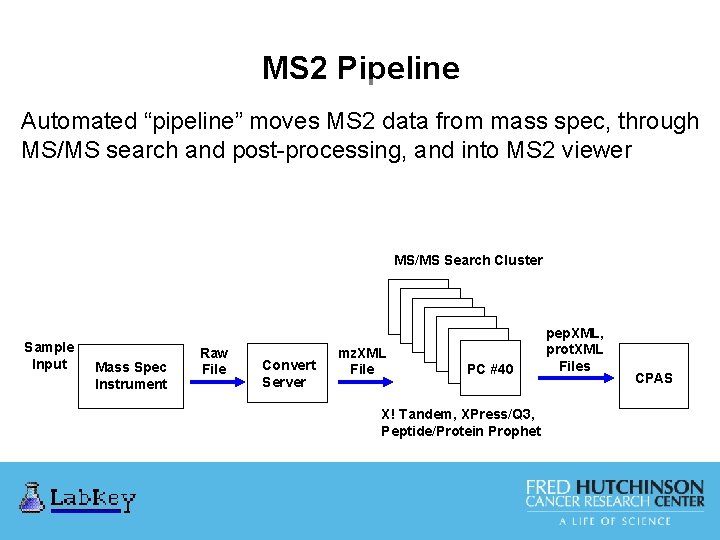
MS 2 Pipeline Automated “pipeline” moves MS 2 data from mass spec, through MS/MS search and post-processing, and into MS 2 viewer MS/MS Search Cluster Sample Input Mass Spec Instrument Raw File Convert Server mz. XML File PC #40 X! Tandem, XPress/Q 3, Peptide/Protein Prophet pep. XML, prot. XML Files CPAS

Key MS/MS Analysis Features • Load MS/MS results produced by many common search engines – X! Tandem, Mascot, SEQUEST, COMET • • Inspect individual MS/MS spectra Filter and sort results based on peptide and protein characteristics: – Search engine scores, Peptide. Prophet. TM, delta mass, modifications, etc. – Sequence mass, sequence coverage, gene name, Protein. Prophet. TM score, etc. • • • Group results by protein or Protein. Prophet groups Customize columns, save favorite filters and views Export filtered, sorted results to Excel, TSV, DTA, PKL formats Filter groups of runs and compare peptides/proteins between them Analyze quantitation of peptides & proteins (via XPRESS/Protein. Prophet) Link results to rich protein annotations & experimental annotations
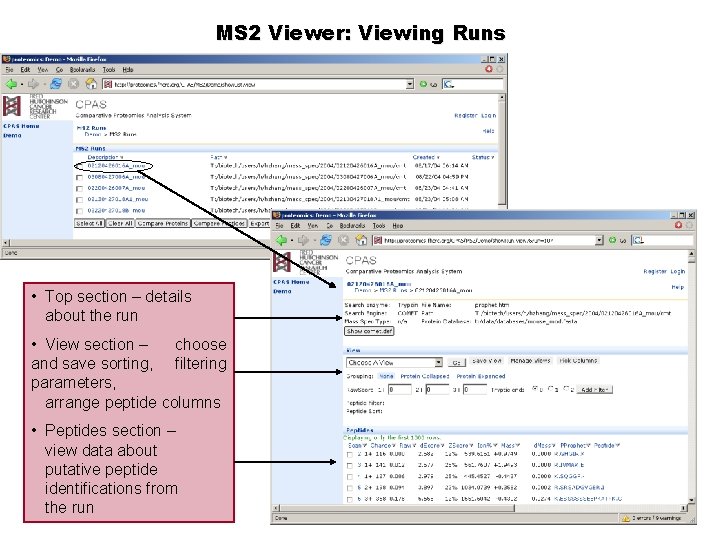
MS 2 Viewer: Viewing Runs • Top section – details about the run • View section – choose and save sorting, filtering parameters, arrange peptide columns • Peptides section – view data about putative peptide identifications from the run

MS 2 Viewer: Expanded Protein View Protein Hits Protein Details Individual MS/MS spectrum
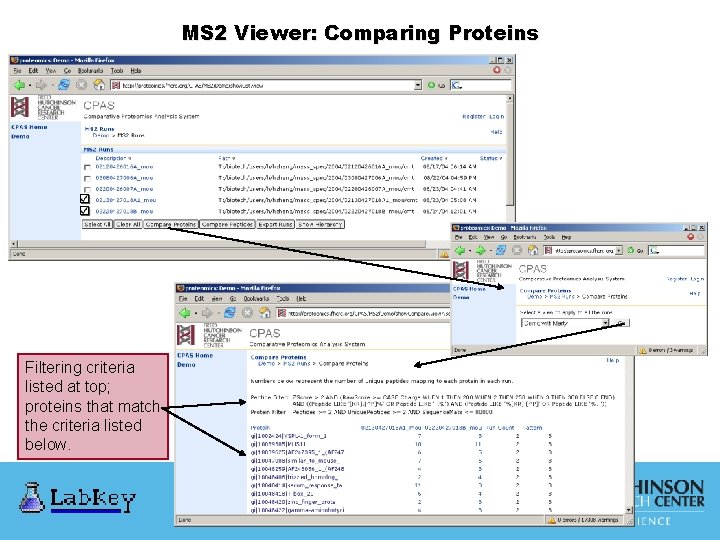
MS 2 Viewer: Comparing Proteins Filtering criteria listed at top; proteins that match the criteria listed below.

Experimental Annotations • Standards-based annotation of experiments • Data/experiment exchange format • See tutorial on http: //cpas. fhcrc. org

Protein Services • Links MS/MS results to database of protein sequences & annotations – Protein sequences are loaded from both FASTA files and annotated protein databases (e. g. , Uni. Prot) – Each sequence is stored once per organism and given a unique Seq. ID – All identifiers, descriptions, annotations, and references from all sources are linked to corresponding Seq. ID – Schema supports addition of new types of identifiers and annotations • This provides ability to: – Display and link to biologically relevant protein information – Compare results searched against different FASTA files (IPI vs. NCBI) – Generate charts summarizing GO metabolic function, cellular location, and molecular function of your results – Link new annotations to old results & regenerate FASTA files needed for re-analysis

MS 2 Viewer: Hands On
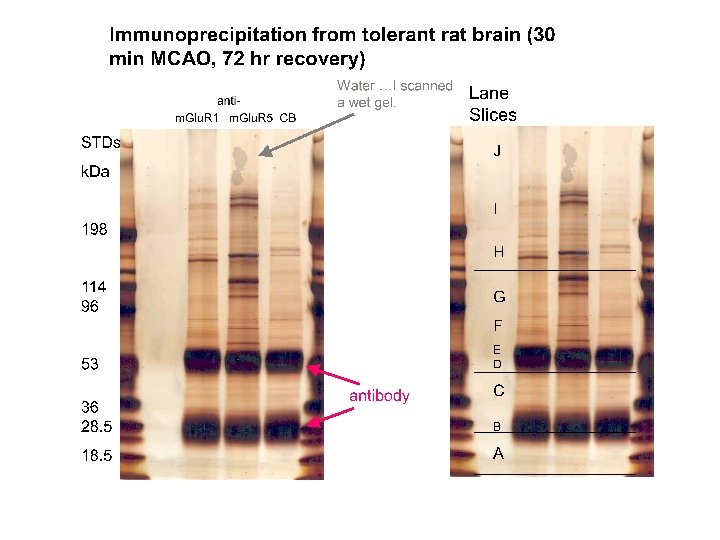
Immunoprecipitation

Scenario #1 • • • From “Proteomics Course” folder, click link for run 5 C Check Tandem Settings and Modifications details Filter peptides to Peptide. Prophet score >= 0. 9 Sort peptides by Peptide. Prophet score Group filtered list of peptides by protein Filter proteins to those with > 1 unique peptide hit Sort proteins by sequence mass Save this view Export list to Excel

Scenario #2 • Apply your saved view to all runs in an experiment – Peptides with Peptide. Prophet score >= 0. 9 – Proteins with more than one unique peptide hit • Show all qualifying proteins – Ordered by number of runs each appeared in – Showing number of unique peptides in each run that indicate that protein • Export results to Excel

Scenario #3 • • • Go to Analysis folder View experiment flow and raw files View Collapsed Protein. Prophet information Filter protein probability > 0. 9 Add protein ratio columns Expand peptides for LALBA_BOSMU and investigate
- Slides: 21