CPAS Comparative Proteomics Analysis System Adam Rauch Lab

CPAS Comparative Proteomics Analysis System Adam Rauch Lab. Key Software adam@labkey. com

What Is CPAS? A proteomics analysis system that handles all data processing & management for high-throughput labs and core facilities

High-Throughput Proteomics Demands • Integrate a wide variety of hardware & software – – – Instruments MS/MS search engines Quantitation, validation, and other analytic tools Custom analysis tools Computing hardware, operating systems, databases IT infrastructure

High-Throughput Proteomics Demands • Integrate a wide variety of hardware & software • Rapidly analyze many large MS/MS runs – – Search, validate, quantify, store millions of peptides per day Re-analyze results, combine runs, experiment “in silico” Automation and repeatability Data analysis should not be the bottleneck

High-Throughput Proteomics Demands • Integrate a wide variety of hardware & software • Rapidly analyze many large MS/MS runs • Manage huge volume of data – Organized and easily accessible results – Analysis & experimental protocols, sample information – Ability to answer biologically interesting questions: • Compare runs & experiments to identify proteins of interest • Search experiments specific proteins • Query results linked to sample & experimental properties, rich protein annotations, custom protein annotation lists

High-Throughput Proteomics Demands • • Integrate a wide variety of hardware & software Rapidly analyze many large MS/MS runs Manage huge volume of data Collaborate easily while keeping data secure – – Colleagues at same institution Cross-institution collaborations and consortia Publish results publicly Appropriate and strong security

How CPAS Addresses These Demands • Integrate a wide variety of hardware & software – – – – CPAS is an integration platform Integrates with SEQUEST, Mascot, and X! Tandem Incorporates Trans-proteomic Pipeline (TPP) from ISB NCI ca. BIGTM silver level compliant Supports custom analytic tools (e. g. , Q 3 quantitation) Runs on all common server operating systems & hardware IT-friendly: LDAP, SASL, databases choice, simple config, etc. Exports results to Excel, various text formats

How CPAS Addresses These Demands • Integrate a wide variety of hardware & software • Rapidly analyze many large MS/MS runs – Fred Hutchinson CPAS pipeline has processed: • 67 thousand MS/MS fractions of 215 million spectra • Individual runs of 300 fractions and 2 million spectra • Millions of spectra per day on a regular basis

How CPAS Addresses These Demands • Integrate a wide variety of hardware & software • Rapidly analyze many large MS/MS runs • Manage large volumes of results – – – All data organized by lab, folder, and experiment Results are organized automatically by pipeline Provides easy reorganization Sophisticated cross-experiment query capabilities Fred Hutchinson: 31 thousand runs, 260 million peptides

How CPAS Addresses These Demands • • Integrate a wide variety of hardware & software Rapidly analyze many large MS/MS runs Manage large volumes of results Collaborate easily while keeping data secure – – CPAS system can be shared on intranet or Internet Access requires just a browser and proper credentials Keeps sensitive, unpublished scientific data secure Provides various publishing and export options

What Is CPAS? • Free, open source software connected to your… – – Instruments Search engine cluster Analytic pipeline Network file system • …that provides a single, easy-to-use interface for: – Submitting and monitoring pipeline jobs – Managing your data – Answering biologically interesting questions about results
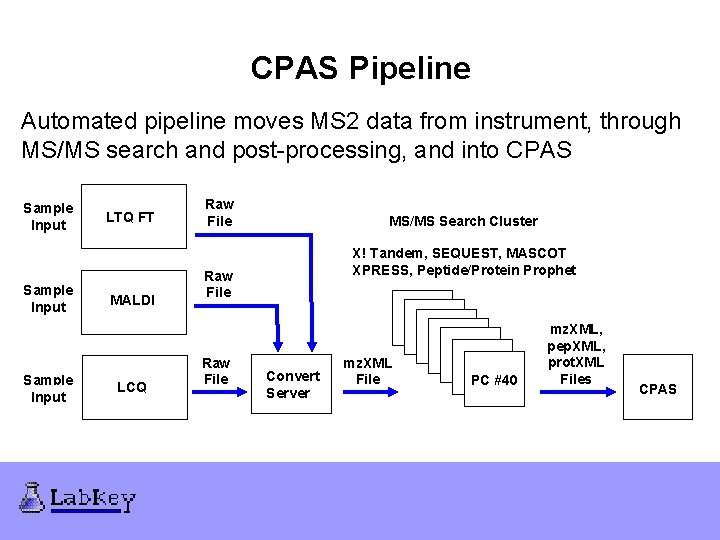
CPAS Pipeline Automated pipeline moves MS 2 data from instrument, through MS/MS search and post-processing, and into CPAS Sample Input LTQ FT MALDI LCQ Raw File MS/MS Search Cluster X! Tandem, SEQUEST, MASCOT XPRESS, Peptide/Protein Prophet Raw File Convert Server mz. XML File PC #40 mz. XML, pep. XML, prot. XML Files CPAS

Basic Analysis Features • Load results produced by Mascot, SEQUEST, X! Tandem • Inspect individual MS/MS spectra • Filter and sort results based on peptide and protein characteristics: – Search engine scores, Peptide. Prophet, delta mass, modifications – Sequence mass, sequence coverage, gene, Protein. Prophet • • Analyze peptide & protein quantitation Group results by protein or Protein. Prophet groups Customize columns, save favorite filters and views Export filtered results to Excel, TSV, DTA, PKL, AMT formats

Advanced Analysis Features • Filter groups of runs and compare peptides, proteins, Protein. Prophet, quantitation, etc • Analyze groups of runs based on sample properties • Search all experiments for a specific protein or gene name • Link results to protein annotations – Load protein knowledgebases: Tr. EMBL, Swiss-Prot – Gene Ontology: produce GO charts analyzing molecular function, cellular location, metabolic process – Custom protein annotation lists • Flexible, custom query capability – Join results to differ – Display exactly the data you care about

Experimental Annotations • Standards-based annotation of experiments • Data/experiment exchange format • See tutorial on http: //www. labkey. org

Demo

What Does “Apache 2. 0 Open Source” Mean? • • • The product is free All source code is available for your review You can modify and extend the product You can contribute changes back (or not) You can re-distribute source or product (modified or not) • Broad development community is emerging

Lab. Key Software, Inc. • Private consulting company created by FHCRC and team of software professionals – Formed to support, document, and extend the CPAS project to other functions and labs – Independent company to directly address other institutions’ needs and secure outside funding • Partnership: – Clients provide scientific leadership – Lab. Key focuses on software development • Lab. Key is available to customize, install, and support your pipeline, CPAS, and other Lab. Key applications – Business model ensures you get help & support when you need it

Resources • http: //www. labkey. org – CPAS Distribution & Support Site – Ask questions, contribute feedback – Peruse all the CPAS documentation & tutorials – Download the latest version (Lab. Key 2. 1) • Graphical installer for Windows installation • Well documented “manual” installation for Linux/Mac • http: //www. labkey. com – Lab. Key Software Inc. company web site • CPAS Paper – Rauch A, Bellew M, Eng J, et al. Computational Proteomics Analysis System (CPAS): An Extensible, Open-source Analytic System for Evaluating and Publishing Proteomic Data and High throughput Biological Experiments. J Proteome Res 2006; 5(1): 112 -121.

Next Steps • Visit our booth • Join our informal receptions here – 6: 30 – 9: 30 PM Mon, Tue, Wed • Install CPAS and give it a test drive – http: //www. labkey. org – USB key

Acknowledgements • • Fred Hutchinson Cancer Research Center National Cancer Institute Canary Foundation Gates Foundation Institute for Systems Biology Ron Beavis & The GPM Numerous developer contributors

Questions?
- Slides: 22