Correlating m RNA and protein abundance via genomic


Correlating m. RNA and protein abundance via genomic and proteomic characteristics Dov Greenbaum Gerstein Lab Thesis Seminar April 21, 2004

outline Why analyze m. RNA and protein correlations Background Disparate Data Sources Correlating m. RNA and Protein Results Other analyses Formalism – comparing genome, transcriptome and proteome in terms of broad categories New Data Sets Analysis via Broad Categories Analysis of factors affecting correlations Another reason to expect correlations Expression and Protein Interactions
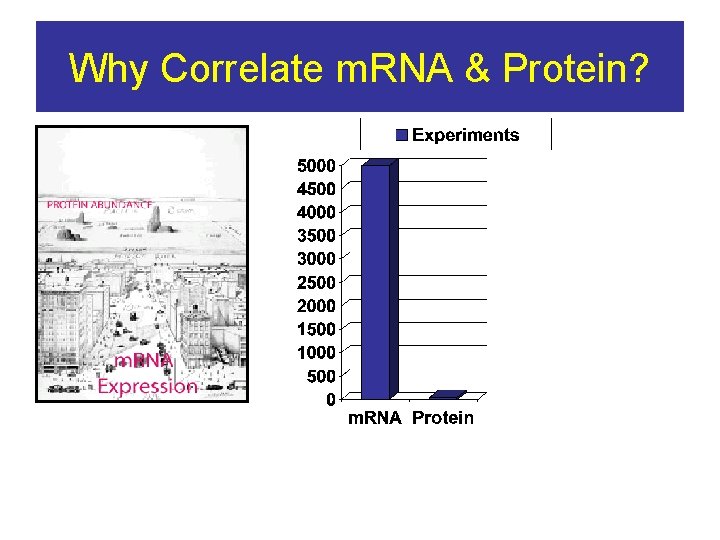
Why Correlate m. RNA & Protein?

Both m. RNA and Protein Levels are necessary for complete analysis Shown mathematically in Hatzimanikatis et al Biotechnology 1999 Combinations of RNA and protein detection approaches have recently aided in the identification of biomarkers in cancer Hegde et al Current Opinion in Biotech 2003
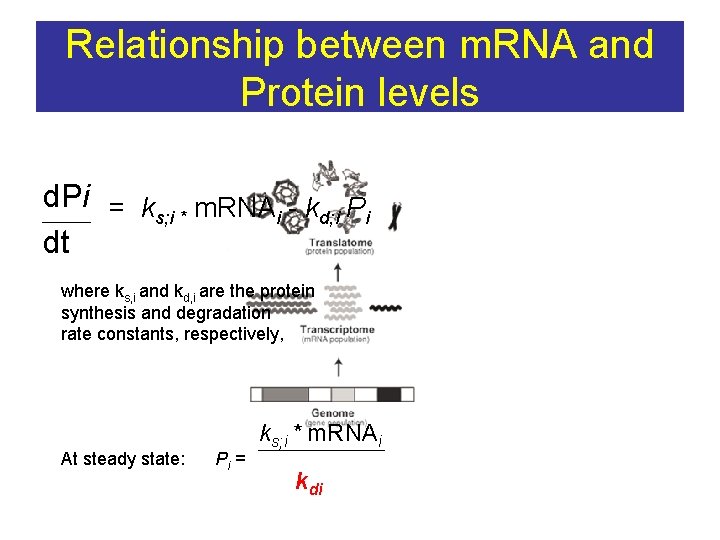
Relationship between m. RNA and Protein levels d. Pi = k m. RNA - k P s; i * i d; i i dt where ks, i and kd, i are the protein synthesis and degradation rate constants, respectively, At steady state: Pi = ks; i * m. RNAi kdi
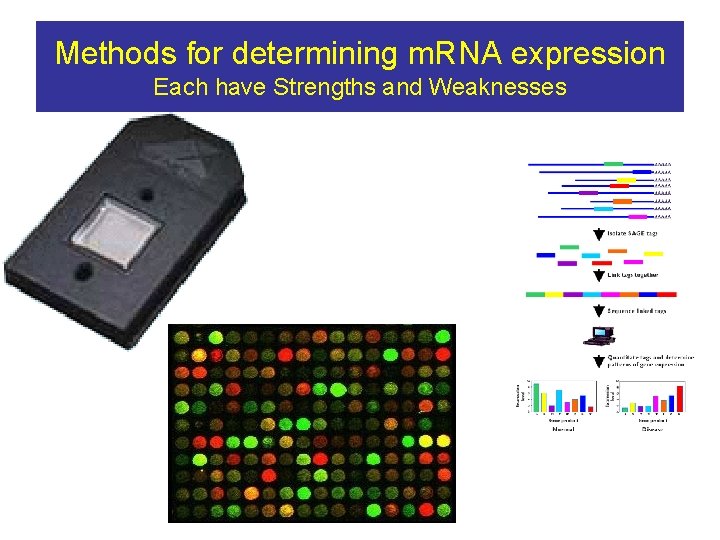
Methods for determining m. RNA expression Each have Strengths and Weaknesses
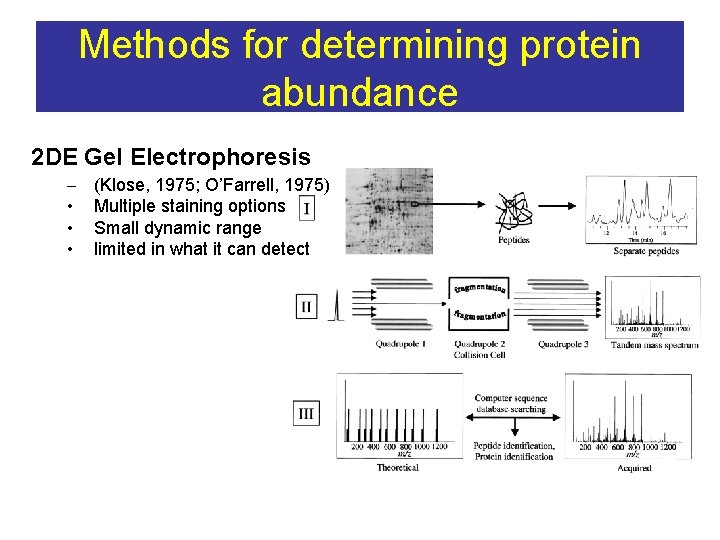
Methods for determining protein abundance 2 DE Gel Electrophoresis – • • • (Klose, 1975; O’Farrell, 1975) Multiple staining options Small dynamic range limited in what it can detect
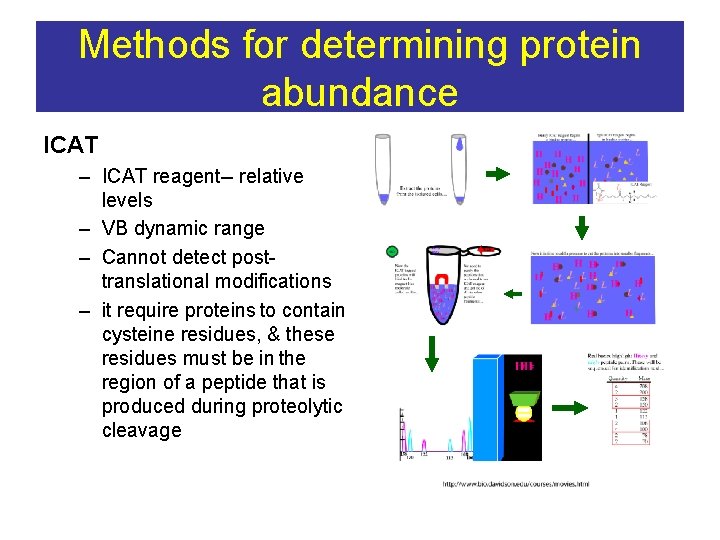
Methods for determining protein abundance ICAT – ICAT reagent-- relative levels – VB dynamic range – Cannot detect posttranslational modifications – it require proteins to contain cysteine residues, & these residues must be in the region of a peptide that is produced during proteolytic cleavage

Mud. Pit Really only HT that can detect PT modifications
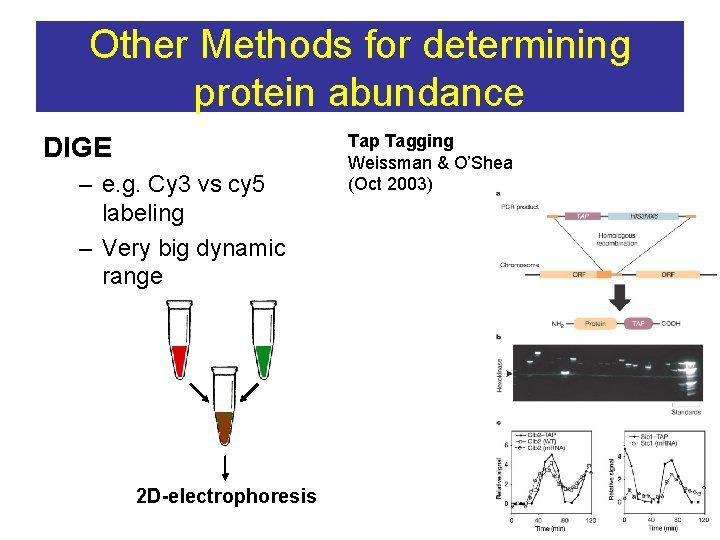
Other Methods for determining protein abundance DIGE – e. g. Cy 3 vs cy 5 labeling – Very big dynamic range 2 D-electrophoresis Tap Tagging Weissman & O’Shea (Oct 2003)
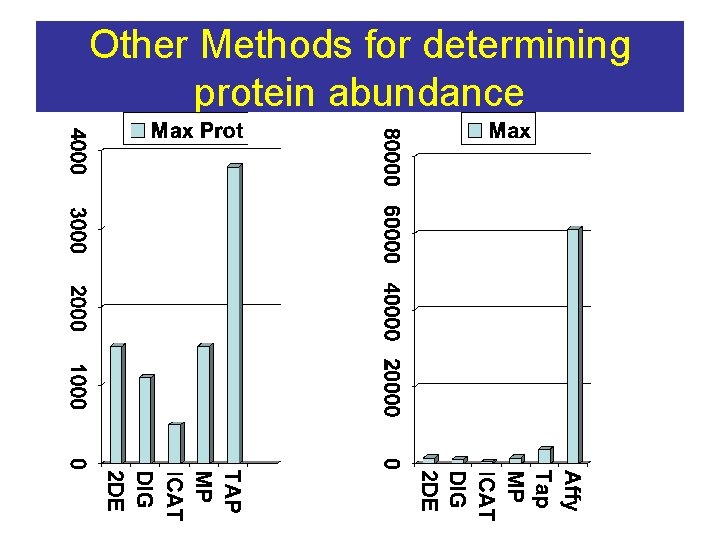
Other Methods for determining protein abundance
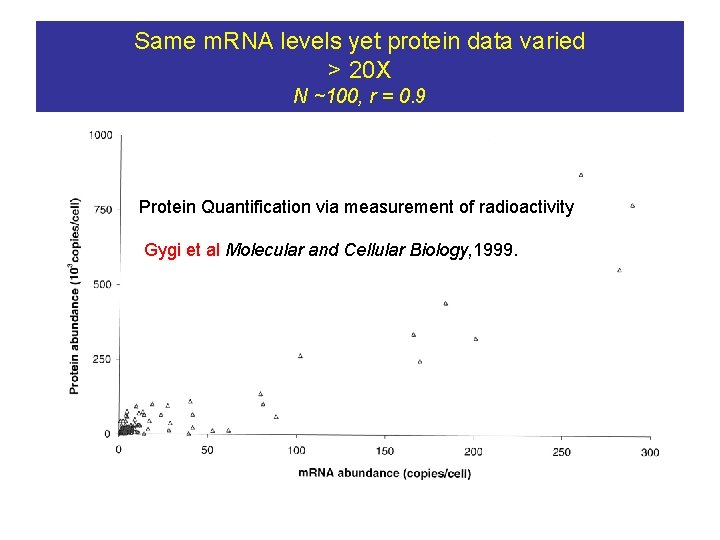
Same m. RNA levels yet protein data varied > 20 X N ~100, r = 0. 9 Protein Quantification via measurement of radioactivity Gygi et al Molecular and Cellular Biology, 1999.
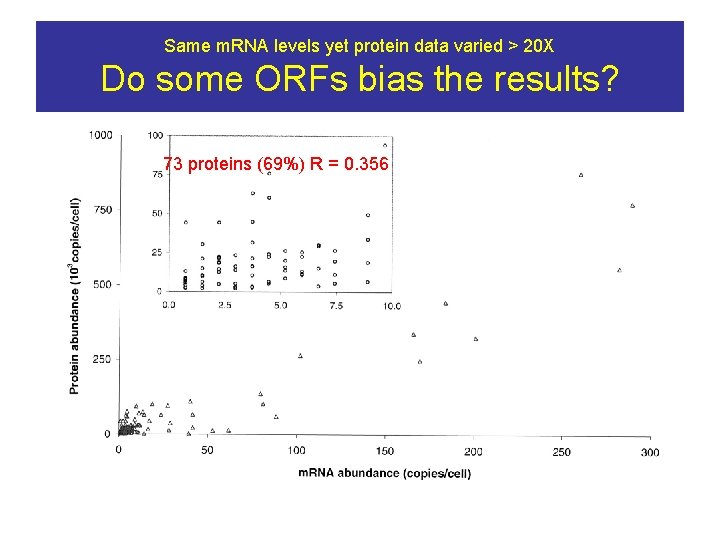
Same m. RNA levels yet protein data varied > 20 X Do some ORFs bias the results? 73 proteins (69%) R = 0. 356
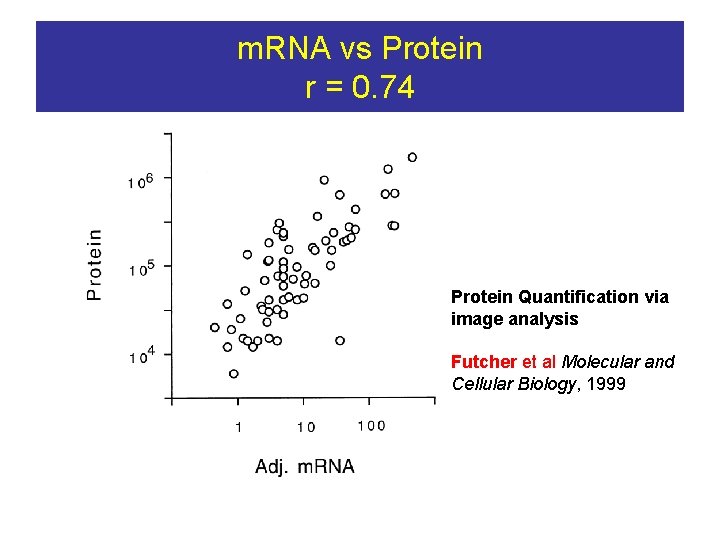
m. RNA vs Protein r = 0. 74 Protein Quantification via image analysis Futcher et al Molecular and Cellular Biology, 1999

Jury is out… Gygi et al: “This study revealed that transcript levels provide little predictive value with respect to the extent of protein expression. ” Futcher et al: “there is a good correlation between protein abundance and m. RNA abundance for the proteins that we have studied”.
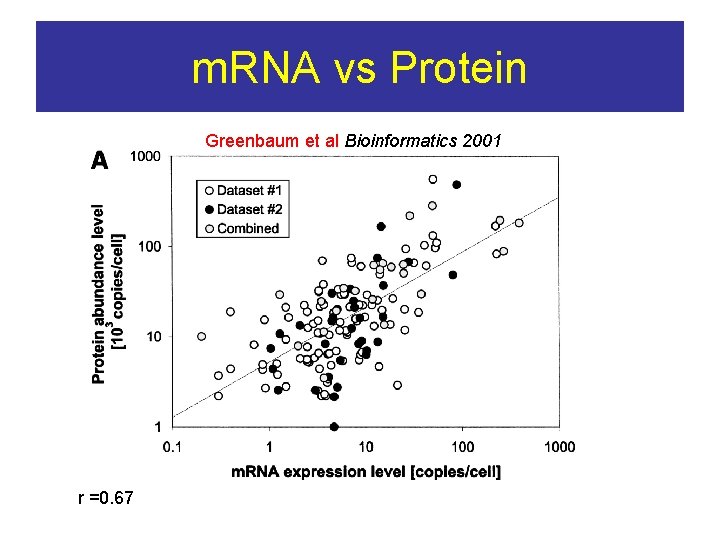
m. RNA vs Protein Greenbaum et al Bioinformatics 2001 r =0. 67
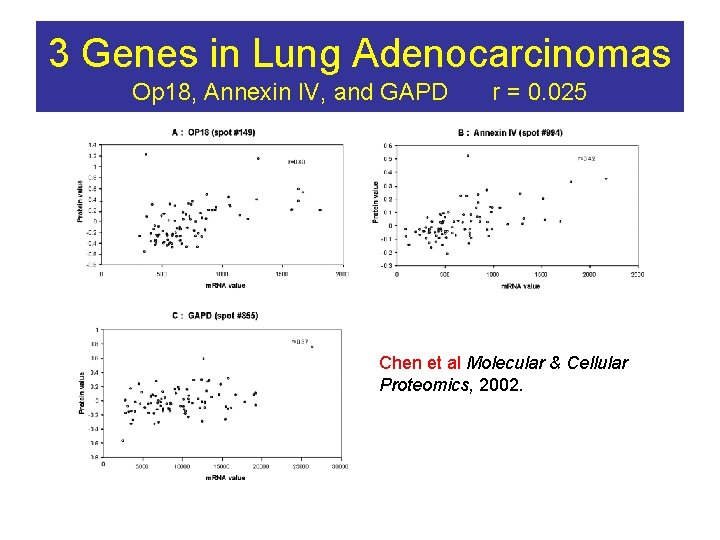
3 Genes in Lung Adenocarcinomas Op 18, Annexin IV, and GAPD r = 0. 025 Chen et al Molecular & Cellular Proteomics, 2002.
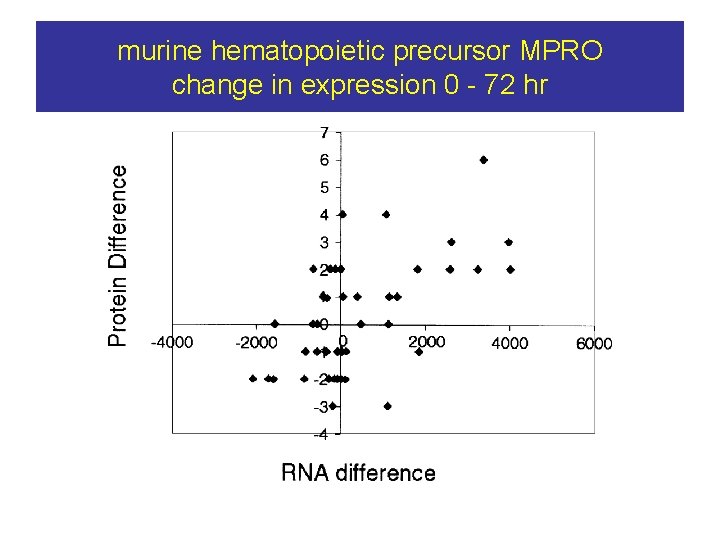
murine hematopoietic precursor MPRO change in expression 0 - 72 hr
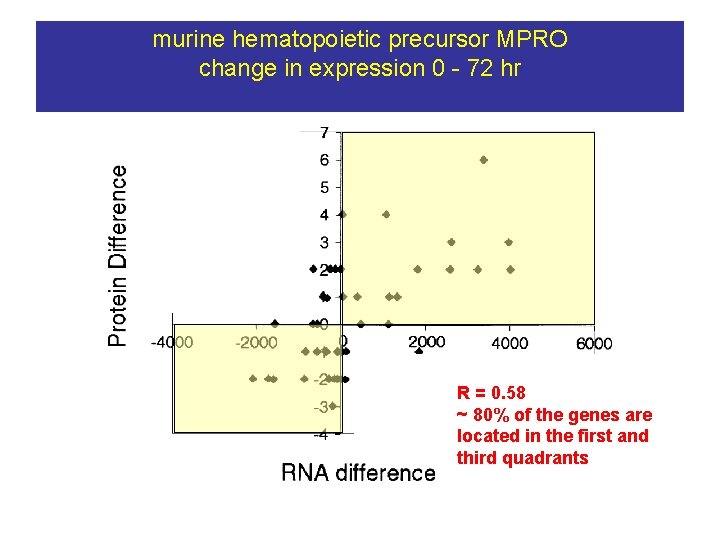
murine hematopoietic precursor MPRO change in expression 0 - 72 hr R = 0. 58 ~ 80% of the genes are located in the first and third quadrants
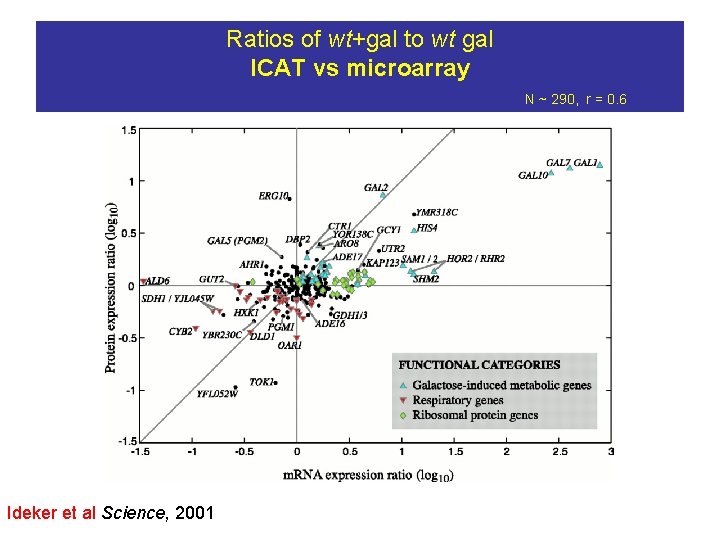
Ratios of wt+gal to wt gal ICAT vs microarray N ~ 290, r = 0. 6 Ideker et al Science, 2001
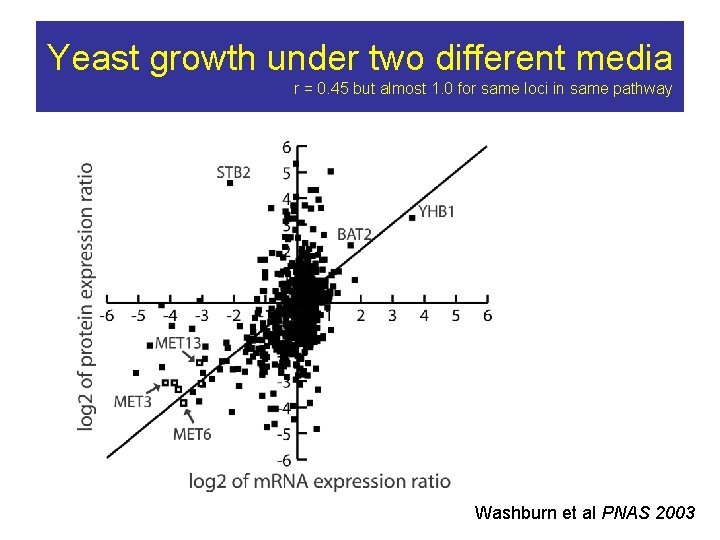
Yeast growth under two different media r = 0. 45 but almost 1. 0 for same loci in same pathway Washburn et al PNAS 2003

Integrating multiple sources of Information The challenge for computational biology is to provide methodologies for transforming high-throughput heterogeneous data sets into biological insights about the underlying mechanisms. Although highthroughput assays provide a global picture, the details are often noisy, hence conclusions should be supported by several types of observations. Integration of data from assays that examine cellular systems from different viewpoints (for instance, gene viewpoints expression and protein-protein interactions) can lead to a more coherent reconstruction and reduce the effects of noise Nir Friedman Science 2004
![Sources of Data set m. RNA expression Protein abundance Annotation Description Size [ORFs] Reference Sources of Data set m. RNA expression Protein abundance Annotation Description Size [ORFs] Reference](http://slidetodoc.com/presentation_image/38c9106b7fe683133a5bcc228abb4140/image-24.jpg)
Sources of Data set m. RNA expression Protein abundance Annotation Description Size [ORFs] Reference Young Gene chip profiles yeast cells with mutations that affect transcription 5455 Holstege et al. (1998) Church Gene chip profiles of yeast cells under four different conditions 6263 Roth et al. (1998) Samson Comparing gene chip profiles for yeast cells subjected to alkylating agent 6090 Jelinsky et al. (1998) SAGE Yeast cells during vegetative growth 3778 Velculescu et al. (1997) Reference expression Scaling and integrating the m. RNA expression set into one data source 6249 - 2 -DE #1 Measurement of yeast protein abundance by two-dimensional (2 D) gel electrophoresis and mass spectrometry 156 2 -DE #2 Similar to 2 -DE set #1 Transposon Large-scale fusions of yeast genes with lac. Z by transposon insertion Reference abundance Scaling and integrating the 2 -DE data sets into one data source Annotated Localization Subcellular localizations of yeast proteins 2133 (6280) Drawid et al. (2000) Transmem-brane segments Predicted transmembrane and soluble proteins in yeast 2710 (6280) Gerstein (1998) MIPS functions Functional categories for yeast ORFs 3519 (6194) Mewes et al. (2000) GOR secondary structure Predicted secondary structure for yeast ORFs 71 1410 181 6280 Gygi et al. (1999) Futcher et al. (1999) Ross-Macdonald et al. (1999) - Gerstein (1998)
Reference m. RNA Sets Young Church Samson SAGE

Fitting Protein Data Original Set
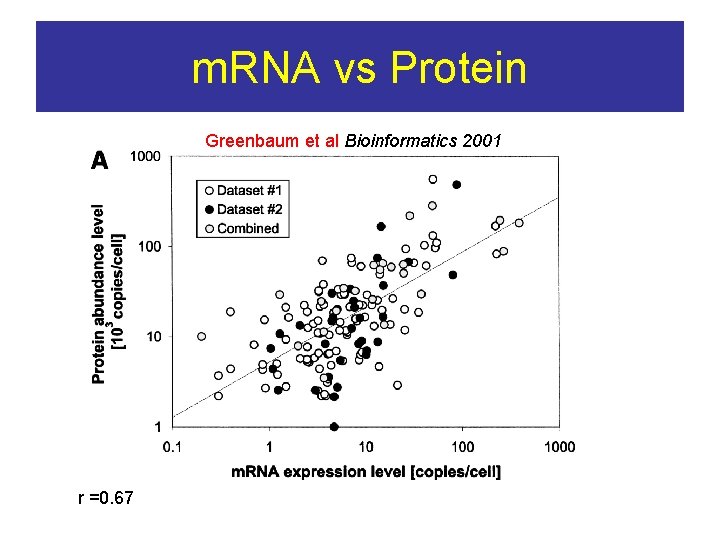
m. RNA vs Protein Greenbaum et al Bioinformatics 2001 r =0. 67
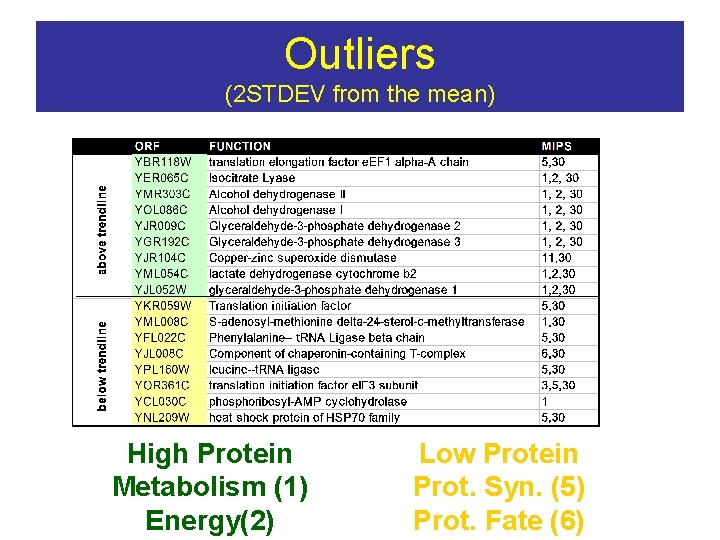
Outliers (2 STDEV from the mean) High Protein Metabolism (1) Energy(2) Low Protein Prot. Syn. (5) Prot. Fate (6)
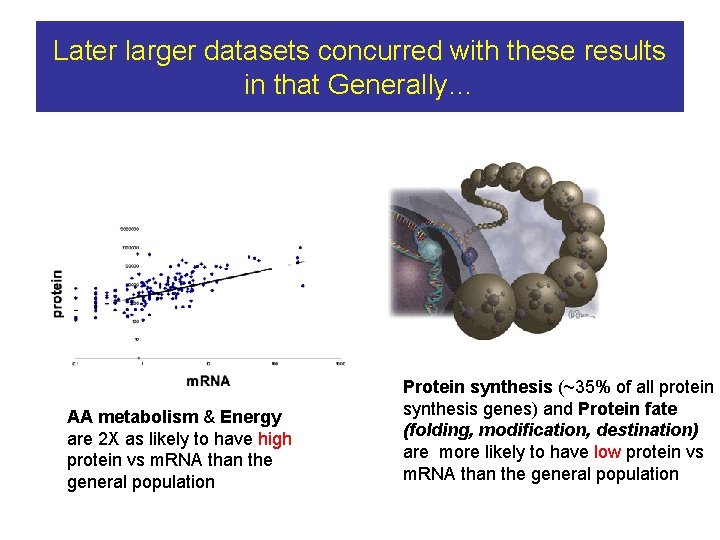
Later larger datasets concurred with these results in that Generally… AA metabolism & Energy are 2 X as likely to have high protein vs m. RNA than the general population Protein synthesis (~35% of all protein synthesis genes) and Protein fate (folding, modification, destination) are more likely to have low protein vs m. RNA than the general population

Non-Outliers Generally… Tight Regulation by the cell Only 3% of transcription associated genes (n = 441) have significantly uncorrelated m. RNA and protein levels (2 STDEV from trendline) Transcription Assoc. genes are 25% of the essential genes in yeast. Essential Genes as a group have higher correlations than the general yeast population 7% of Cell Cycle associated genes (n = 432) have significant non-correlation

Quick Summary • Why correlate m. RNA and protein levels? • Merged Disparate Data Sets – Distinct but complimentary • Global Correlations • Outliers are interesting: – Metabolism & Energy Relatively high protein levels – Protein Synthesis & Protein Fate low protein levels
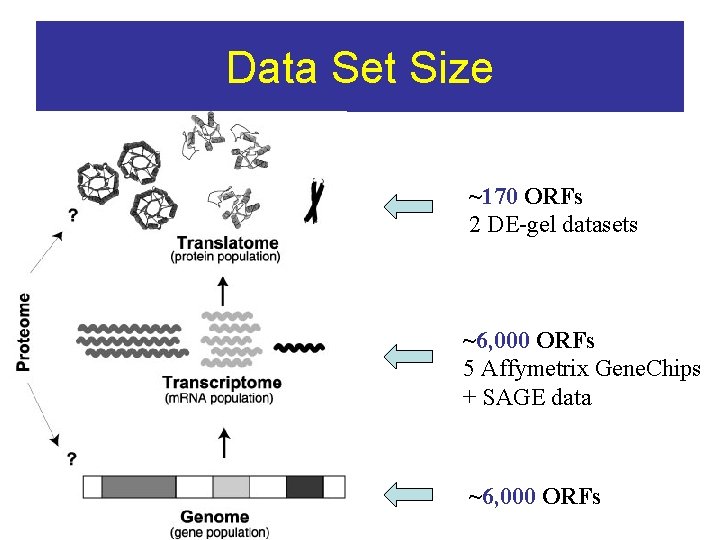
Data Set Size ~170 ORFs 2 DE-gel datasets ~6, 000 ORFs 5 Affymetrix Gene. Chips + SAGE data ~6, 000 ORFs
![Enrichments (Feature, [v, S], [w, G]) = (F, [v, S]) - (F, [w, G]) Enrichments (Feature, [v, S], [w, G]) = (F, [v, S]) - (F, [w, G])](http://slidetodoc.com/presentation_image/38c9106b7fe683133a5bcc228abb4140/image-33.jpg)
Enrichments (Feature, [v, S], [w, G]) = (F, [v, S]) - (F, [w, G]) V & W are weights (expression level) of Sets S & G
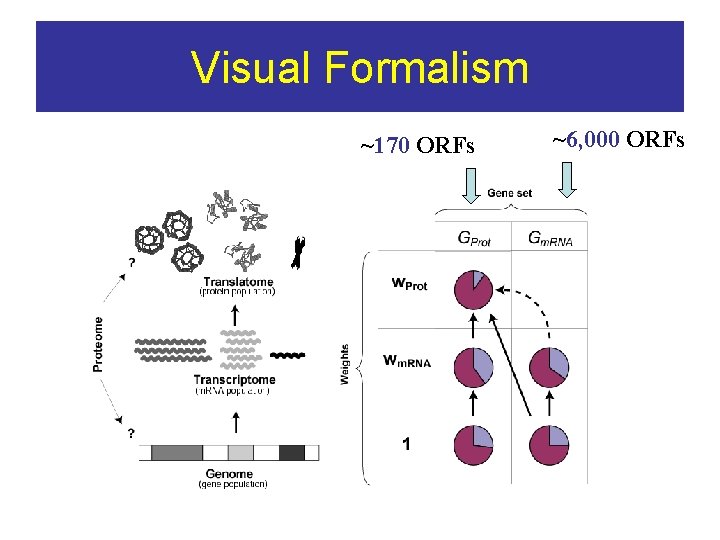
Visual Formalism ~170 ORFs ~6, 000 ORFs
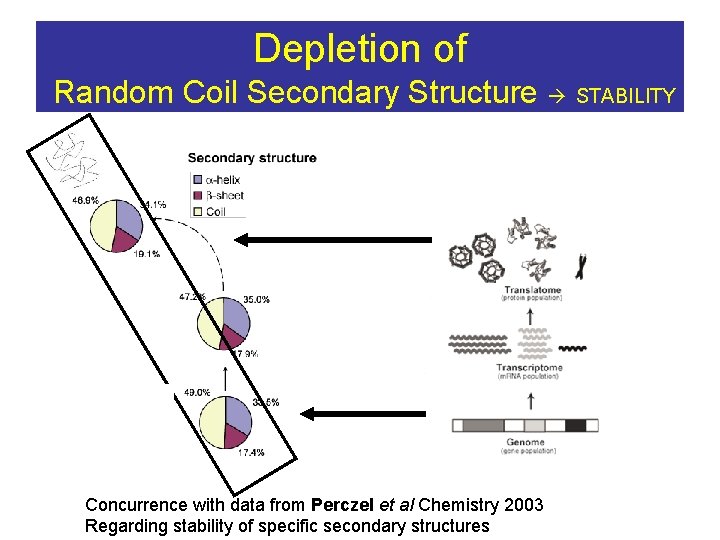
Depletion of Random Coil Secondary Structure STABILITY Concurrence with data from Perczel et al Chemistry 2003 Regarding stability of specific secondary structures

Enrichment of Amino Acids STABILITY Alanine’s, Glycines, Valines result in more compact structures More compact = more stable (i. e. thermophilic enzymes tend to be very compact)
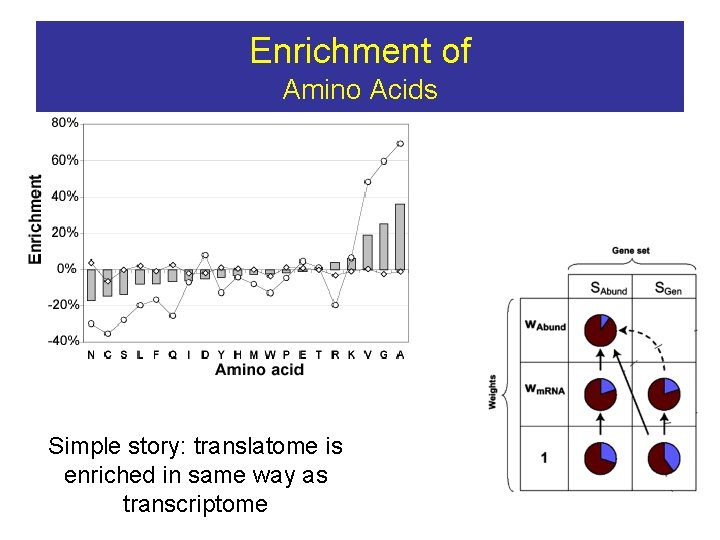
Enrichment of Amino Acids Simple story: translatome is enriched in same way as transcriptome

Enrichment of Molecular Weights/Biomass Abundant proteins are smaller = reduces cost Effect of transcription yeast cell favors the expression of shorter ORFs over longer ones (as opposed to long lightweight ORFs – see MW of aa) This selection is happening, for the most part at the transcriptome level -------------------------------------------------Neg Correlation between ORF length and m. RNA expression Jansen & Gerstein 2000 (And to a lesser degree with Protein Abundance)

Enrichment of Molecular Weights/Biomass Abundant proteins are smaller = reduces cost Effect of transcription CONCURS with experimental results from Akashi, Genetics 2003 See also: Akashi, Genetics 1996 & Moriyama and Powell, NAR 1998 hypothesize that this trend exists in S. cerevisiae, D. melanogaster and E. coli. (although probably not in C. elegans)
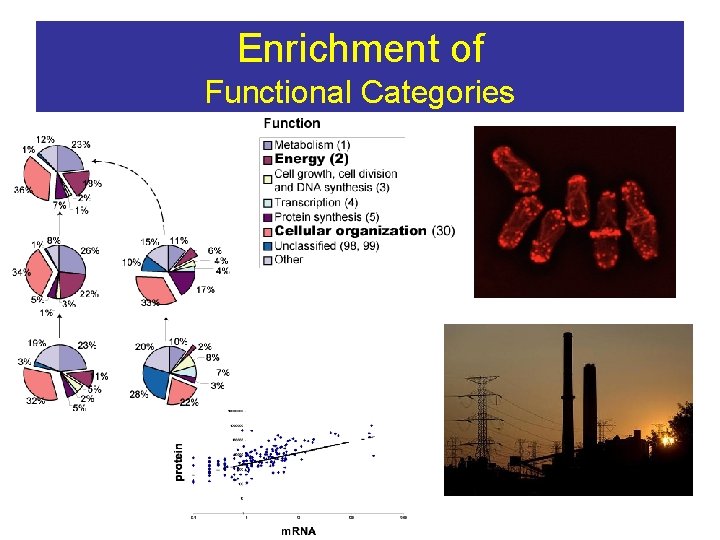
Enrichment of Functional Categories
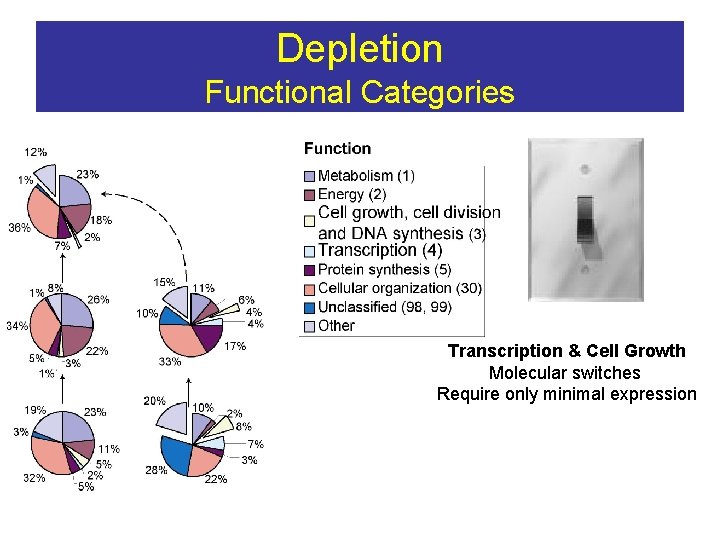
Depletion Functional Categories Transcription & Cell Growth Molecular switches Require only minimal expression
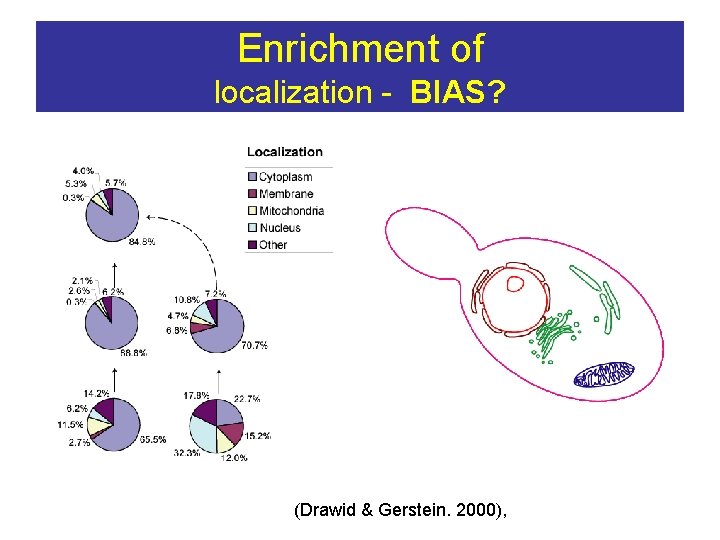
Enrichment of localization - BIAS? (Drawid & Gerstein. 2000),

Review Formalism Different gene sets b/c of limited data Enrichments concur with experimental results
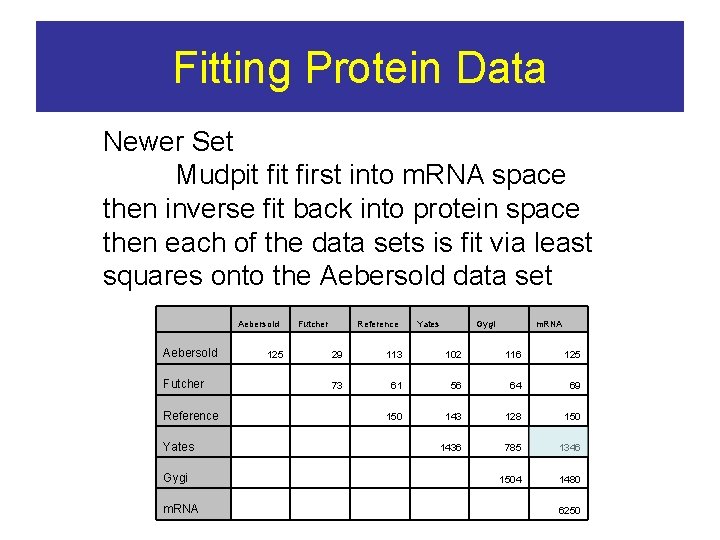
Fitting Protein Data Newer Set Mudpit first into m. RNA space then inverse fit back into protein space then each of the data sets is fit via least squares onto the Aebersold data set Aebersold Futcher 125 Reference Yates Gygi m. RNA 29 113 102 116 125 73 61 56 64 69 150 143 128 150 1436 785 1346 1504 1480 Futcher Reference Yates Gygi m. RNA 6250
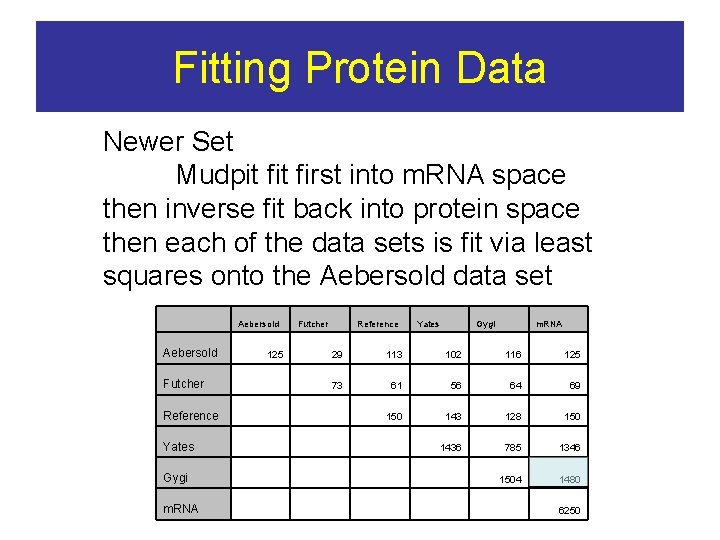
Fitting Protein Data Newer Set Mudpit first into m. RNA space then inverse fit back into protein space then each of the data sets is fit via least squares onto the Aebersold data set Aebersold Futcher 125 Reference Yates Gygi m. RNA 29 113 102 116 125 73 61 56 64 69 150 143 128 150 1436 785 1346 1504 1480 Futcher Reference Yates Gygi m. RNA 6250
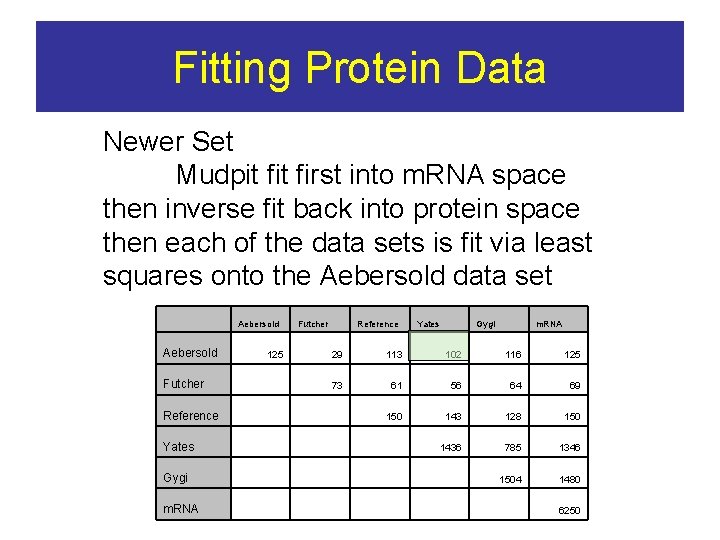
Fitting Protein Data Newer Set Mudpit first into m. RNA space then inverse fit back into protein space then each of the data sets is fit via least squares onto the Aebersold data set Aebersold Futcher 125 Reference Yates Gygi m. RNA 29 113 102 116 125 73 61 56 64 69 150 143 128 150 1436 785 1346 1504 1480 Futcher Reference Yates Gygi m. RNA 6250
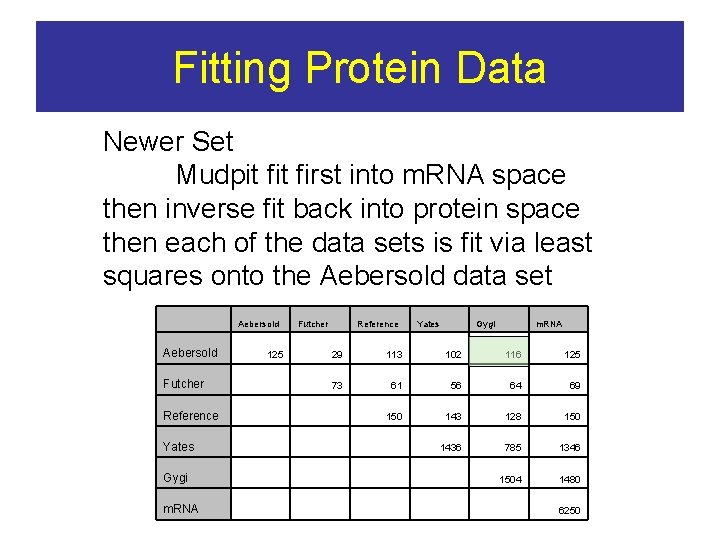
Fitting Protein Data Newer Set Mudpit first into m. RNA space then inverse fit back into protein space then each of the data sets is fit via least squares onto the Aebersold data set Aebersold Futcher 125 Reference Yates Gygi m. RNA 29 113 102 116 125 73 61 56 64 69 150 143 128 150 1436 785 1346 1504 1480 Futcher Reference Yates Gygi m. RNA 6250
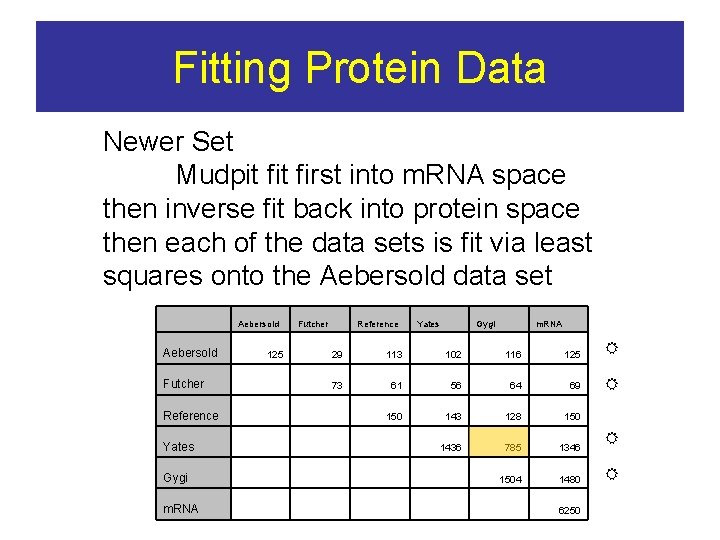
Fitting Protein Data Newer Set Mudpit first into m. RNA space then inverse fit back into protein space then each of the data sets is fit via least squares onto the Aebersold data set Aebersold Futcher 125 Reference Yates Gygi m. RNA 29 113 102 116 125 73 61 56 64 69 150 143 128 150 1436 785 1346 1504 1480 Futcher Reference Yates Gygi m. RNA 6250
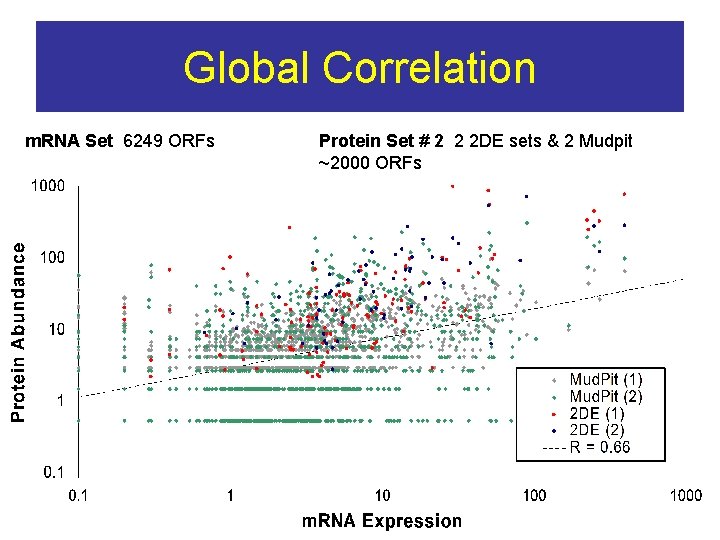
Global Correlation m. RNA Set 6249 ORFs Protein Set # 2 2 2 DE sets & 2 Mudpit ~2000 ORFs
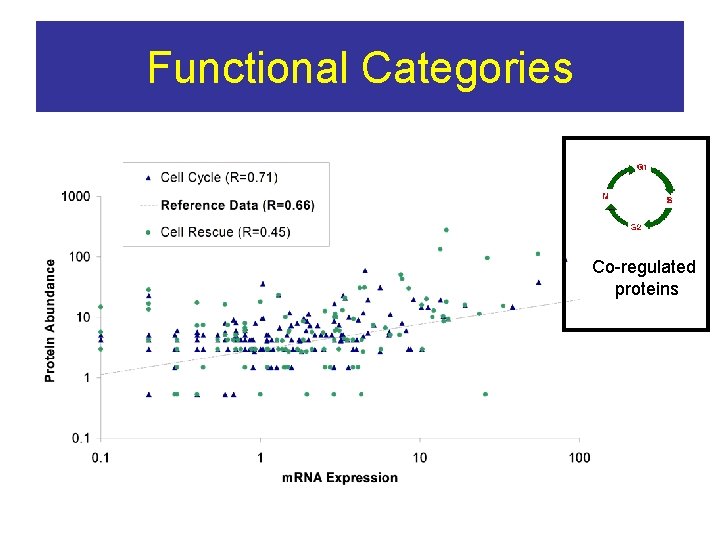
Functional Categories Co-regulated proteins
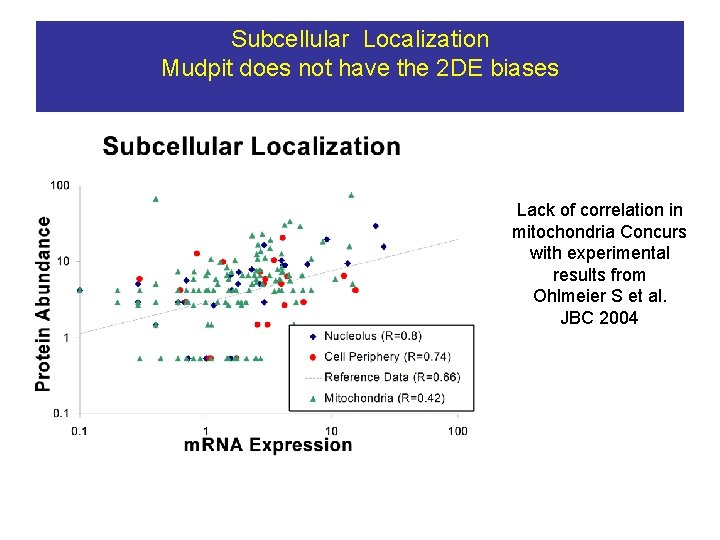
Subcellular Localization Mudpit does not have the 2 DE biases Lack of correlation in mitochondria Concurs with experimental results from Ohlmeier S et al. JBC 2004
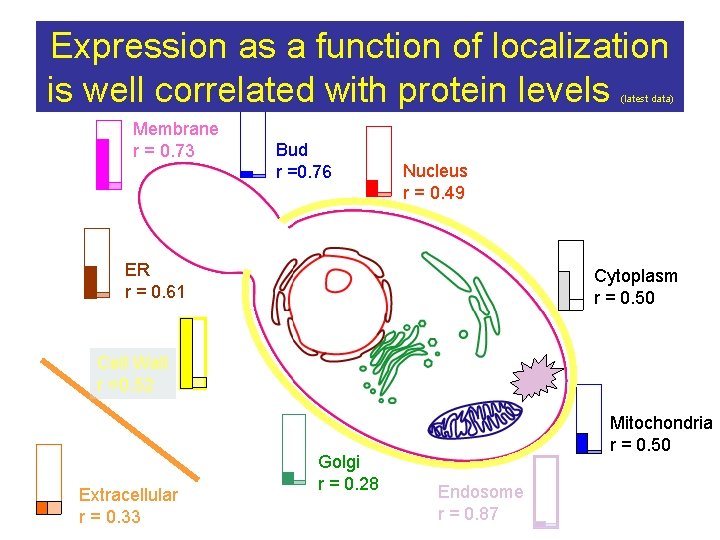
Expression as a function of localization is well correlated with protein levels (latest data) Membrane r = 0. 73 Bud r =0. 76 r global = 0. 46 Nucleus r = 0. 49 P M ER r = 0. 61 Cytoplasm r = 0. 50 Cell Wall r =0. 52 Extracellular r = 0. 33 Golgi r = 0. 28 Mitochondria r = 0. 50 Endosome r = 0. 87

Why would we not find strong correlations? Post translational modifications Protein degradation Error and Bias

Ribosomal Occupancy Arava et al. (2003) Proc. Natl. Acad. Sci. USA Our results concurred with experimental findings by Brown and Herschlag’s groups: Moreover: m. RNAs not associated with any polysomes have even less of a correlation r = 0. 2 v. strong translational control
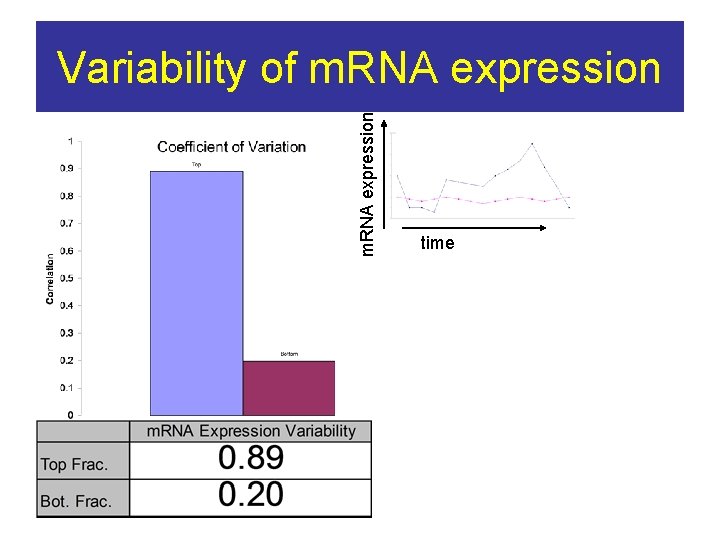
m. RNA expression Variability of m. RNA expression time
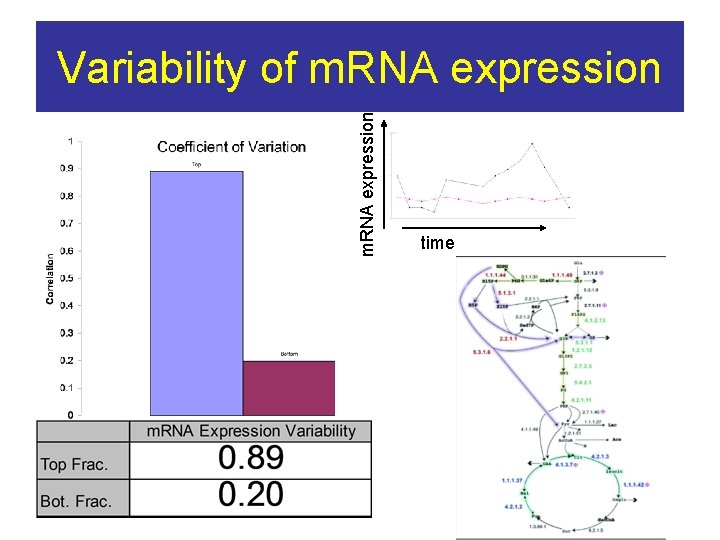
m. RNA expression Variability of m. RNA expression time
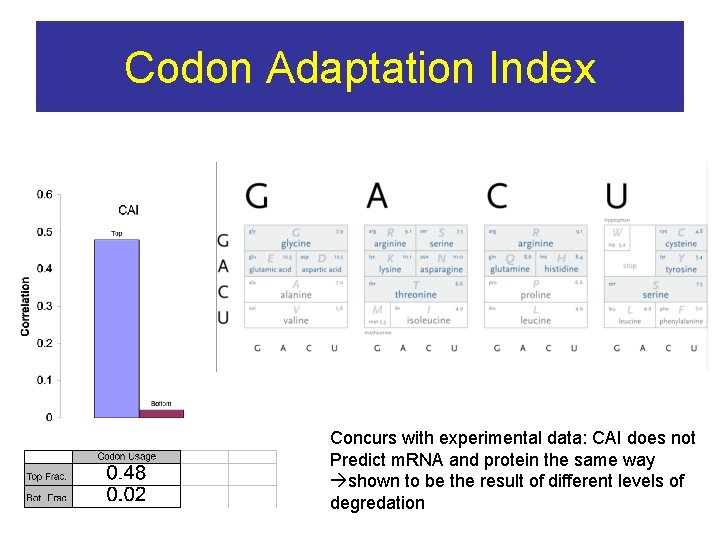
Codon Adaptation Index Concurs with experimental data: CAI does not Predict m. RNA and protein the same way shown to be the result of different levels of degredation

Another summary Newer, larger data set Looking at Broad Catagories I Post translational modifications? where we expect PT control --> low r. Where we don’t expect --> high r Occupancy Variability II Protein Degradation? CAI III Experimental Error? next section

Expression and interactions Types of protein-protein interactions – Protein complexes • For example: proteasome, ribosome – Aggregated interactions • Yeast two-hybrid (Y 2 H) • Genetic/physical interactions from MIPS
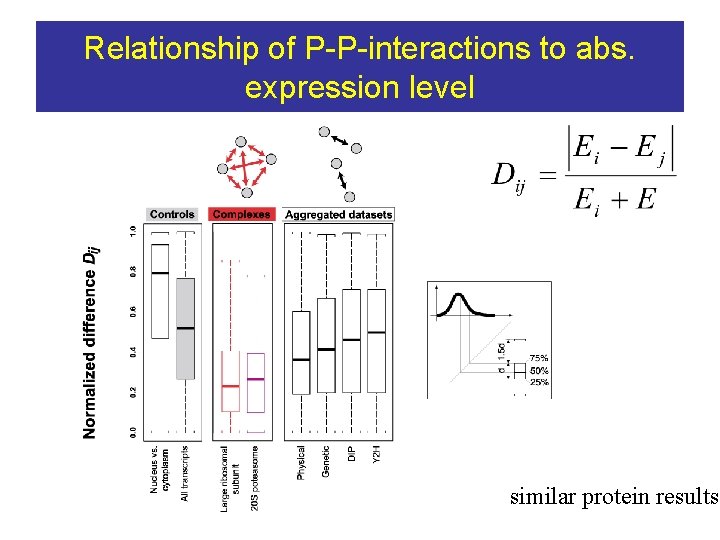
Relationship of P-P-interactions to abs. expression level similar protein results
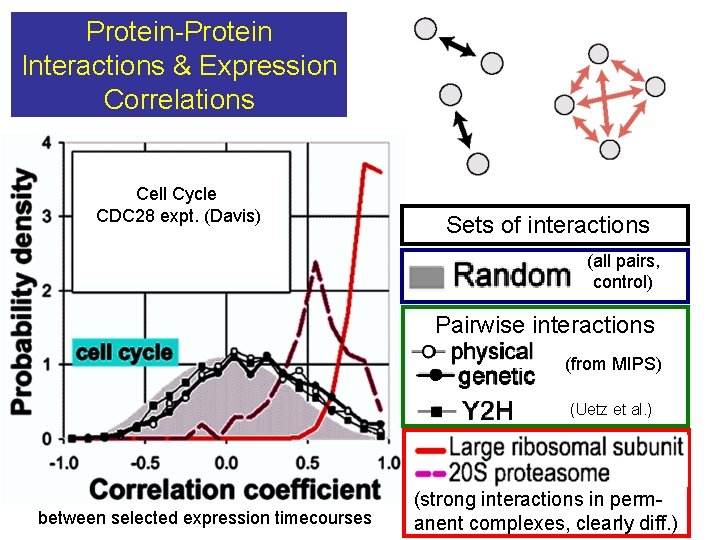
Protein-Protein Interactions & Expression Correlations Cell Cycle CDC 28 expt. (Davis) Sets of interactions (all pairs, control) Pairwise interactions (from MIPS) (Uetz et al. ) between selected expression timecourses (strong interactions in perm- anent complexes, clearly diff. )
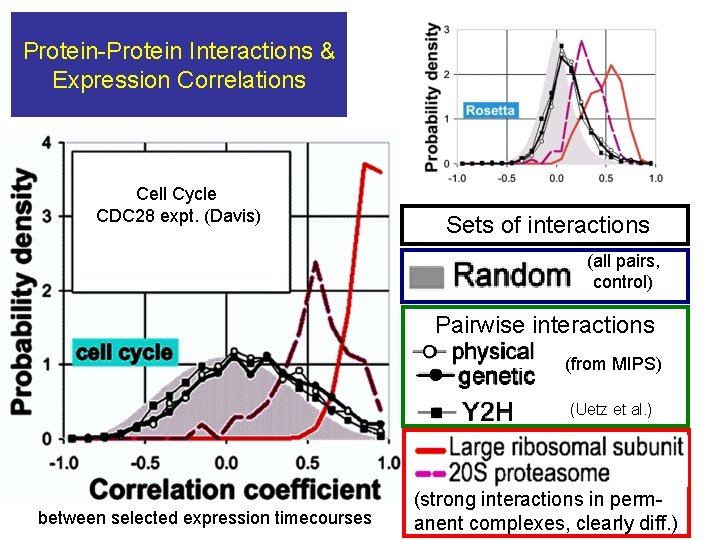
Protein-Protein Interactions & Expression Correlations Cell Cycle CDC 28 expt. (Davis) Sets of interactions (all pairs, control) Pairwise interactions (from MIPS) (Uetz et al. ) between selected expression timecourses (strong interactions in perm- anent complexes, clearly diff. )
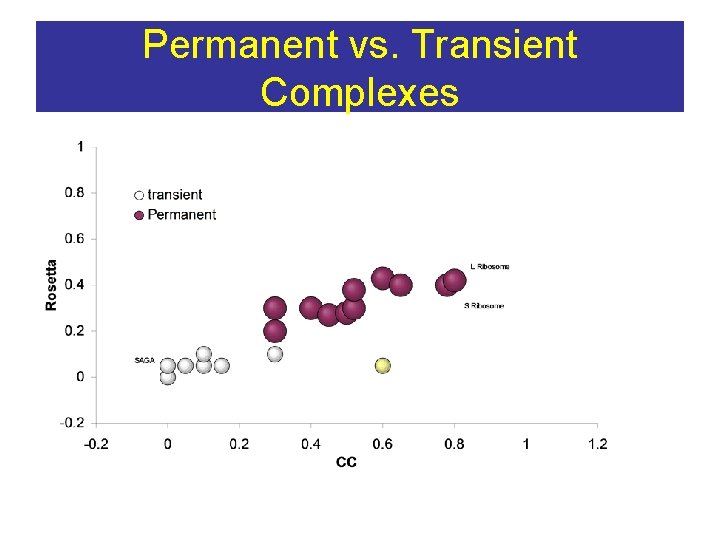
Permanent vs. Transient Complexes
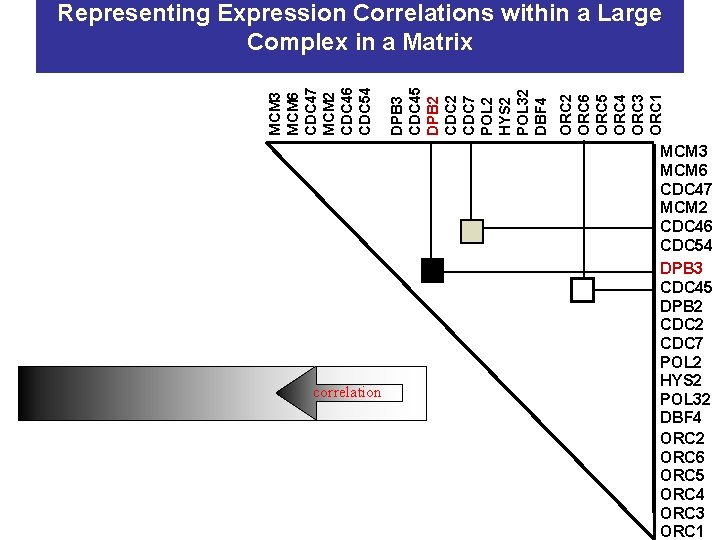
correlation ORC 2 ORC 6 ORC 5 ORC 4 ORC 3 ORC 1 DPB 3 CDC 45 DPB 2 CDC 7 POL 2 HYS 2 POL 32 DBF 4 MCM 3 MCM 6 CDC 47 MCM 2 CDC 46 CDC 54 Representing Expression Correlations within a Large Complex in a Matrix MCM 3 MCM 6 CDC 47 MCM 2 CDC 46 CDC 54 DPB 3 CDC 45 DPB 2 CDC 7 POL 2 HYS 2 POL 32 DBF 4 ORC 2 ORC 6 ORC 5 ORC 4 ORC 3 ORC 1

Permanent? Transient? correlation

L 7/L 12 Cell degrades all excess riboosmal proteins, except L 7 & L 12 correlation
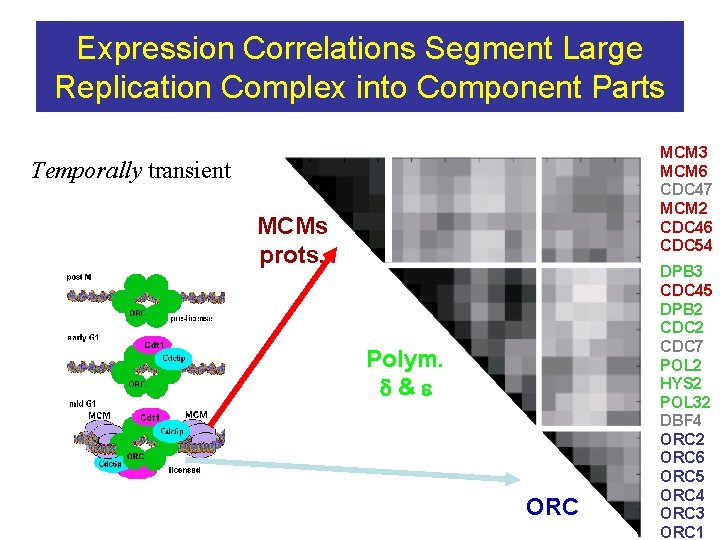
Expression Correlations Segment Large Replication Complex into Component Parts MCM 3 MCM 6 CDC 47 MCM 2 CDC 46 CDC 54 Temporally transient MCMs prots. Polym. d&e ORC DPB 3 CDC 45 DPB 2 CDC 7 POL 2 HYS 2 POL 32 DBF 4 ORC 2 ORC 6 ORC 5 ORC 4 ORC 3 ORC 1

Proteasome No distinction visible between components Proteasome Overall. 43 20 S. 50 19 S. 51 indicative of the possibility that the two components are really one? Division is an artifact of their discovery—M Hochstrasser
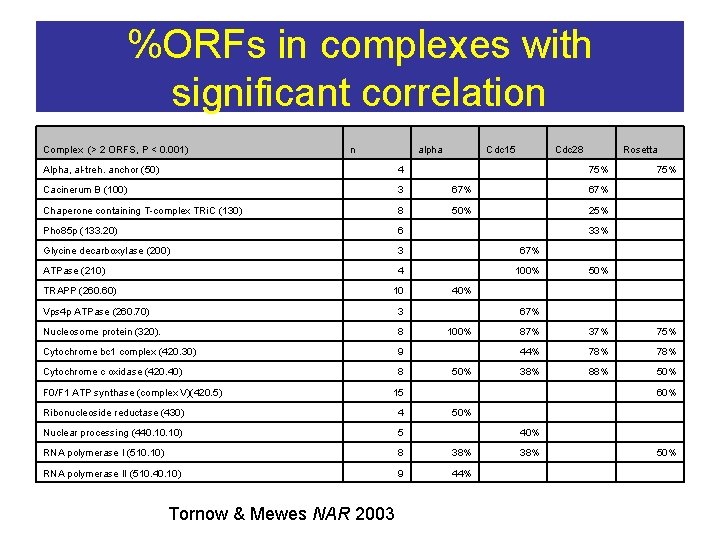
%ORFs in complexes with significant correlation Complex (> 2 ORFS, P < 0. 001) n alpha Cdc 15 Cdc 28 Rosetta 75% Alpha, al-treh. anchor (50) 4 Cacinerum B (100) 3 67% Chaperone containing T-complex TRi. C (130) 8 50% 25% Pho 85 p (133. 20) 6 33% Glycine decarboxylase (200) 3 67% ATPase (210) 4 100% TRAPP (260. 60) 10 Vps 4 p ATPase (260. 70) 3 Nucleosome protein (320). 8 Cytochrome bc 1 complex (420. 30) 9 Cytochrome c oxidase (420. 40) 8 F 0/F 1 ATP synthase (complex V)(420. 5) 15 40% 67% 100% 50% Ribonucleoside reductase (430) 4 Nuclear processing (440. 10) 5 RNA polymerase I (510. 10) 8 38% RNA polymerase II (510. 40. 10) 9 44% Tornow & Mewes NAR 2003 50% 75% 50% 87% 37% 75% 44% 78% 38% 88% 50% 40% 38% 60% 50%
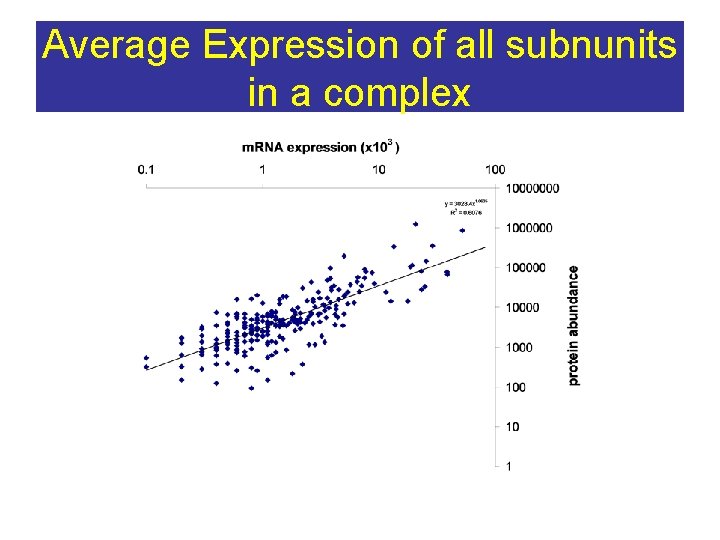
Average Expression of all subnunits in a complex

PP INT Summary Complexes broad catagories minimize noise – Permanent complexes show strong co-expression Posttranscriptional regulation functions at a whole complex level (Washburn et al PNAS 2003) – Transient complexes have weaker co-expression Aggregated BINARY interactions (Y 2 H, physical, genetic) Weak co-expression similar to transient complexes --noisy data? ERROR ? minimized in larger groups

Global Summary m. RNA expression is related to protein abundance Broad categories minimize noise that prevents us from seeing this correlation Integrating various genomic data is integral to an analysis Biologically relevant results can be seen when looking at m. RNA and protein populations

Future Research Further indepth analysis into protein degredation Integrate new Tap Tagging data into protein abundance ref set More intensive modeling of the relationship between m. RNA and protein
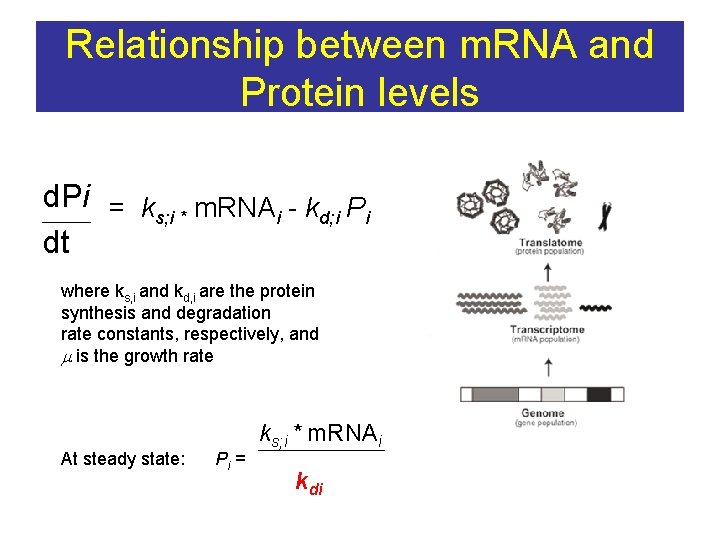
Relationship between m. RNA and Protein levels d. Pi = k m. RNA - k P s; i * i d; i i dt where ks, i and kd, i are the protein synthesis and degradation rate constants, respectively, and m is the growth rate At steady state: Pi = ks; i * m. RNAi kdi

N end rule PEST?

Results of protein degredation Significantly higher correlation for fast decaying proteins Not for slow decay high decay rate is indicative of greater cellular control over level e. g. proteins with half lives of days – cell can’t tightly control Results are same for m. RNA degredation -half lives have been quantified

Acknowledgments Gerstein Lab This work Ronald Jansen (MSKCC) Yuval Kluger (NYU) Other Projects Haiyuan Yu Hedi Hegyi Jimmy Lin Rajdeep Das Jiang Qian Nick Luscombe Entire Gerstein Lab Weissman Lab Zheng Lian Keck (HHMI Biopolymer Laboratory and W. M. Keck Foundation Biotechnology Resource Laboratory) Christopher Colangelo Ken Williams Thesis Committee Mark Gerstein Sherman Weissman Kevin White Genetics Department SABRINA

Liana
- Slides: 78