Convolutional LSTM Networks for Subcellular Localization of Proteins

Convolutional LSTM Networks for Subcellular Localization of Proteins Søren Kaae Sønderby, Casper Kaae Sønderby, Henrik Nielsen*, and Ole Winther *Center for Biological Sequence Analysis Department of Systems Biology Technical University of Denmark 1
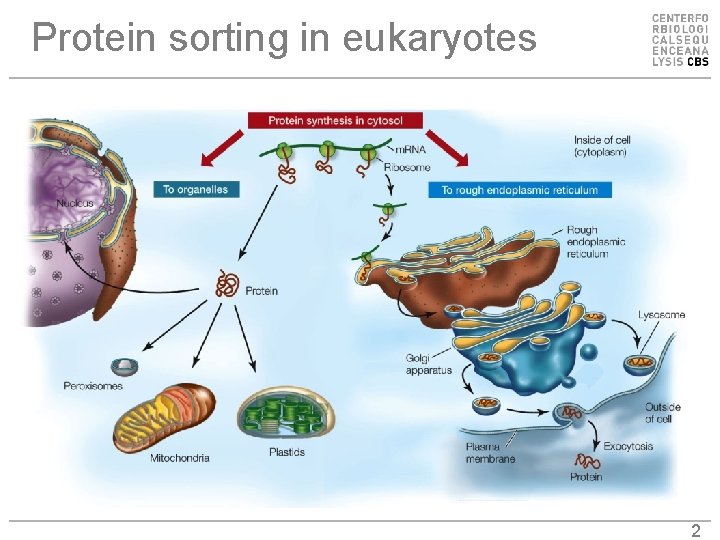
Protein sorting in eukaryotes 2
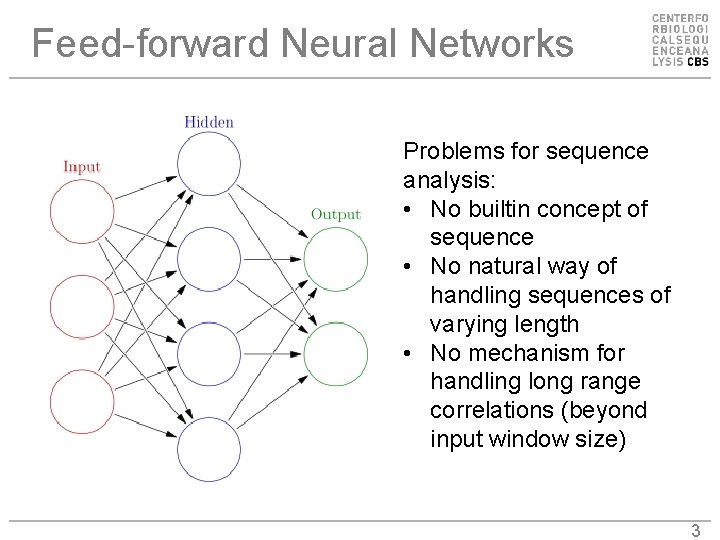
Feed-forward Neural Networks Problems for sequence analysis: • No builtin concept of sequence • No natural way of handling sequences of varying length • No mechanism for handling long range correlations (beyond input window size) 3
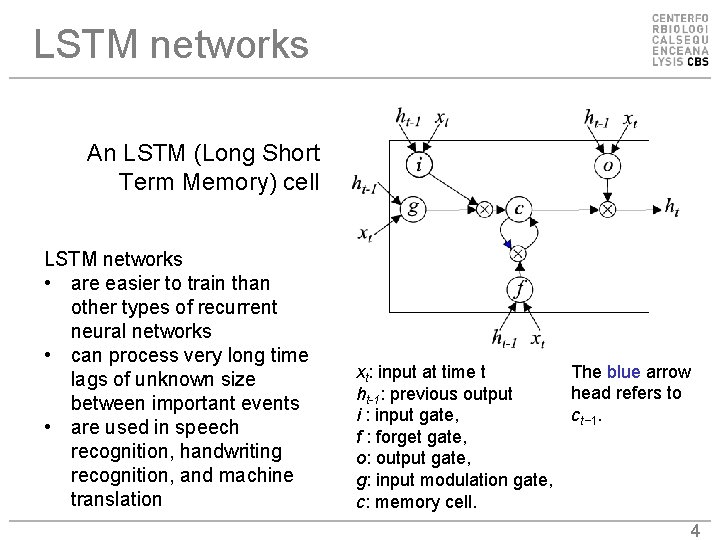
LSTM networks An LSTM (Long Short Term Memory) cell LSTM networks • are easier to train than other types of recurrent neural networks • can process very long time lags of unknown size between important events • are used in speech recognition, handwriting recognition, and machine translation xt: input at time t ht-1: previous output i : input gate, f : forget gate, o: output gate, g: input modulation gate, c: memory cell. The blue arrow head refers to ct− 1. 4
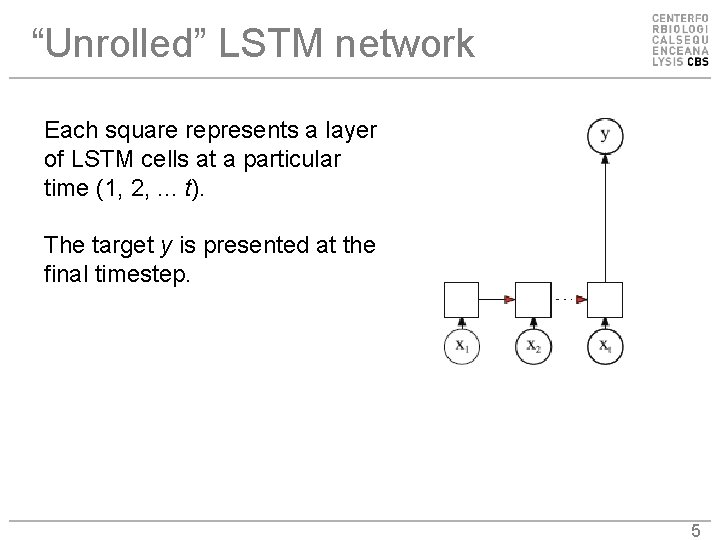
“Unrolled” LSTM network Each square represents a layer of LSTM cells at a particular time (1, 2, . . . t). The target y is presented at the final timestep. 5
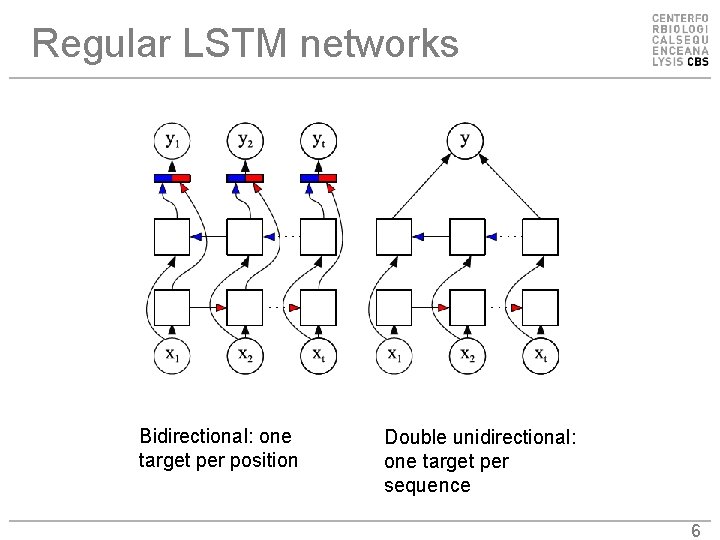
Regular LSTM networks Bidirectional: one target per position Double unidirectional: one target per sequence 6
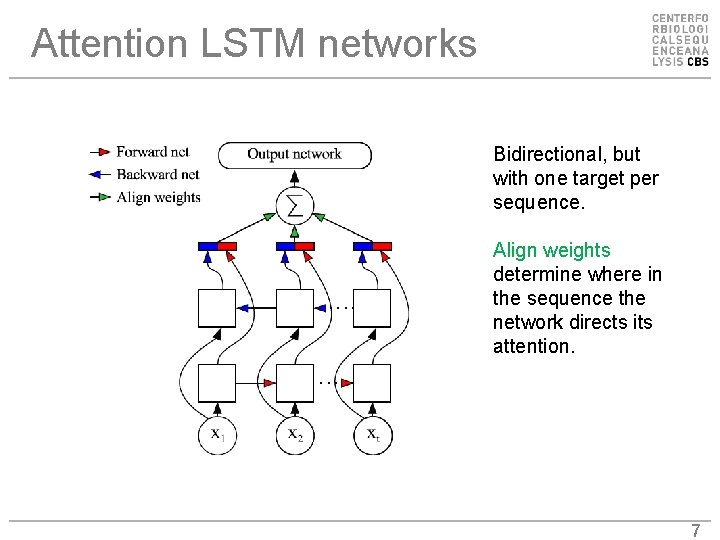
Attention LSTM networks Bidirectional, but with one target per sequence. Align weights determine where in the sequence the network directs its attention. 7
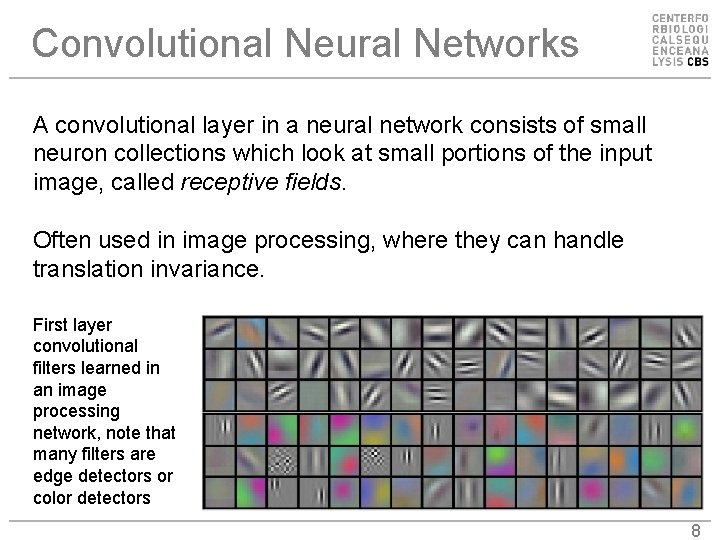
Convolutional Neural Networks A convolutional layer in a neural network consists of small neuron collections which look at small portions of the input image, called receptive fields. Often used in image processing, where they can handle translation invariance. First layer convolutional filters learned in an image processing network, note that many filters are edge detectors or color detectors 8
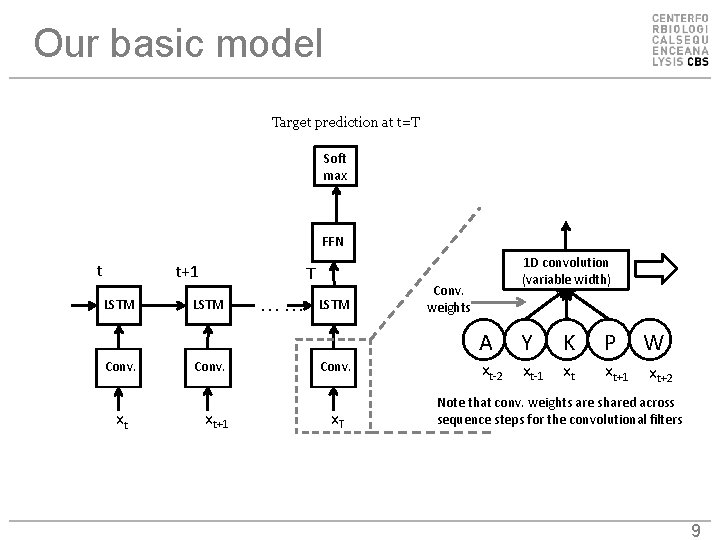
Our basic model Target prediction at t=T Soft max FFN t t+1 LSTM Conv. xt T LSTM Conv. xt+1 …… LSTM Conv. x. T 1 D convolution (variable width) Conv. weights A Y K P xt-2 xt-1 xt xt+1 W xt+2 Note that conv. weights are shared across sequence steps for the convolutional filters 9
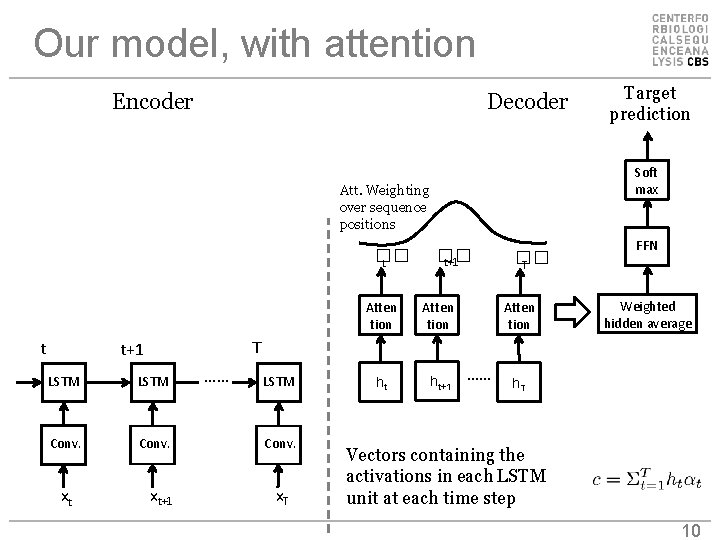
Our model, with attention Encoder Decoder Soft max Att. Weighting over sequence positions �� t t �� t+1 Atten tion ht ht+1 �� T Atten tion FFN Weighted hidden average T t+1 LSTM Conv. xt Target prediction xt+1 …… LSTM Conv. x. T …… h. T Vectors containing the activations in each LSTM unit at each time step 10

Our model, specifications – Input encoding: Sparse, BLOSUM 80, HSDM and profile (R 1× 80) – Conv. filter sizes: 1, 3, 5, 9, 15, 21 (10 of each) – LSTM layer: 1× 200 units – Fully connected FFN layer: 1× 200 units – Attention model: Wa (R 200× 400), va (R 1× 200) 11
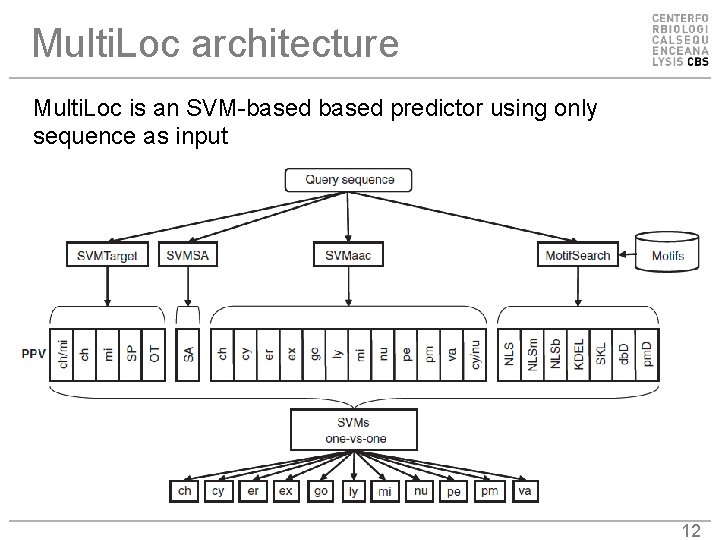
Multi. Loc architecture Multi. Loc is an SVM-based predictor using only sequence as input 12
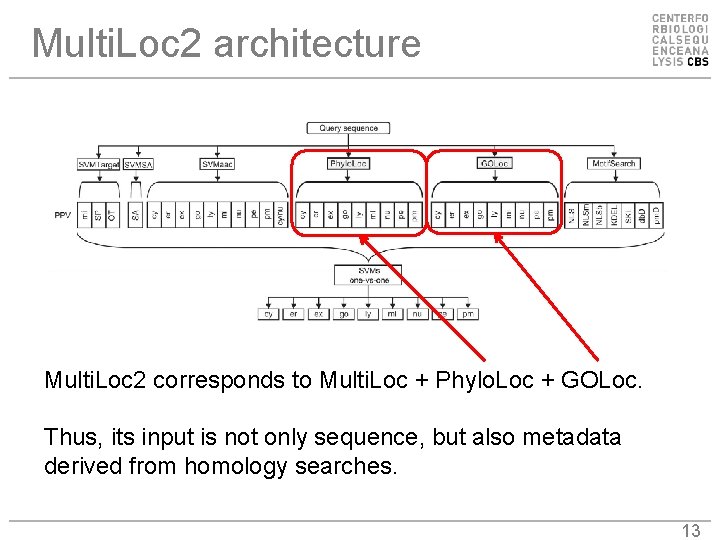
Multi. Loc 2 architecture Multi. Loc 2 corresponds to Multi. Loc + Phylo. Loc + GOLoc. Thus, its input is not only sequence, but also metadata derived from homology searches. 13
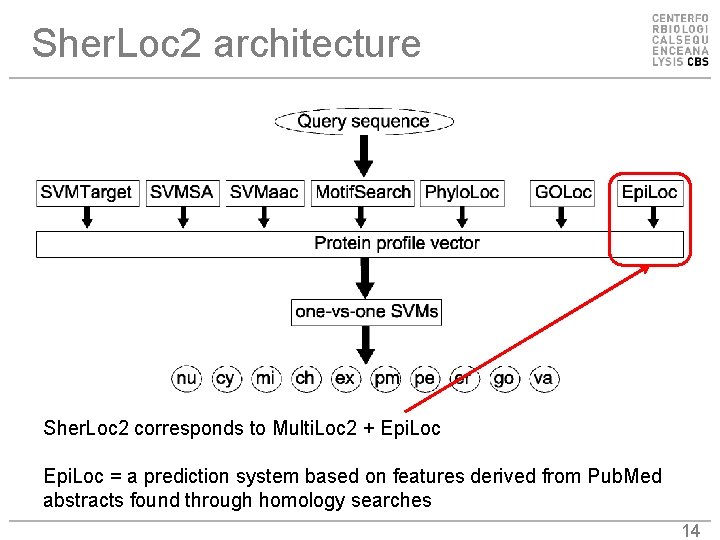
Sher. Loc 2 architecture Sher. Loc 2 corresponds to Multi. Loc 2 + Epi. Loc = a prediction system based on features derived from Pub. Med abstracts found through homology searches 14
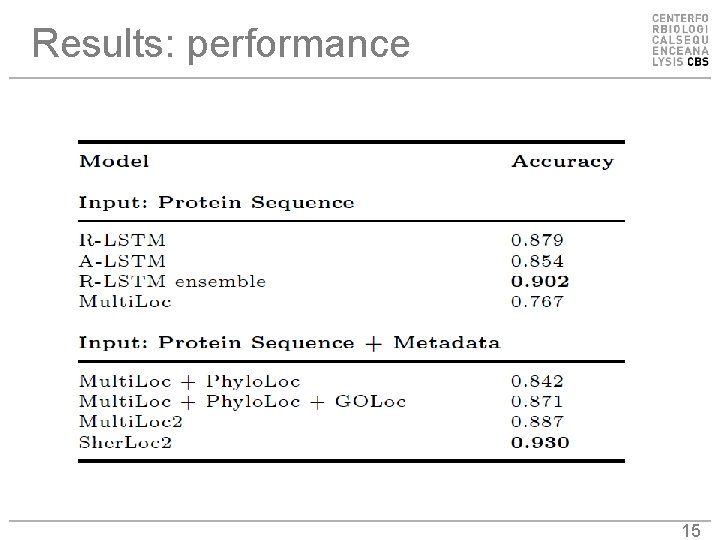
Results: performance 15
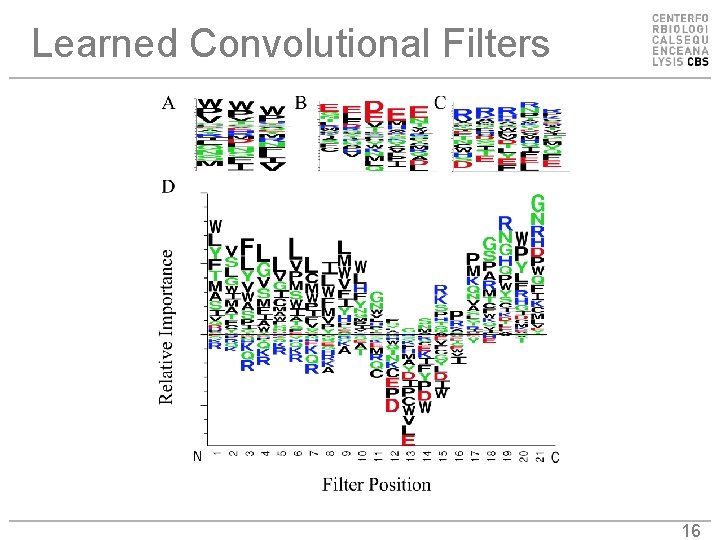
Learned Convolutional Filters 16
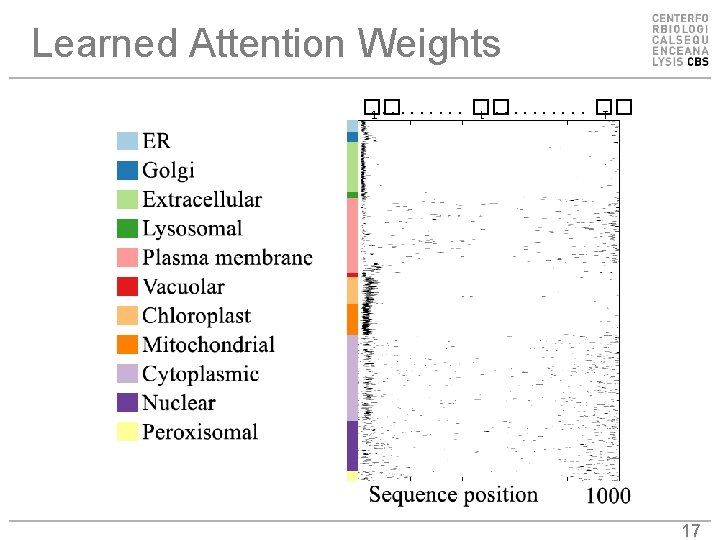
Learned Attention Weights �� 1. . �� t. . �� T 17
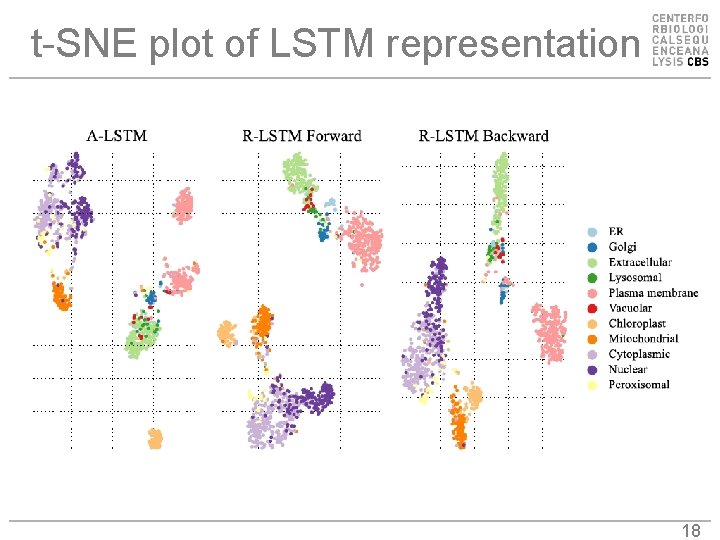
t-SNE plot of LSTM representation 18

Contributions 1. We show that LSTM networks combined with convolutions are efficient for predicting subcellular localization of proteins from sequence. 2. We show that convolutional filters can be used for amino acid sequence analysis and introduce a visualization technique. 3. We investigate an attention mechanism that lets us visualize where the LSTM network focuses. 4. We show that the LSTM network effectively extracts a fixed length representation of variable length proteins. 19

Acknowledgments Thanks to: • Søren & Casper Kaae Sønderby, for doing the actual implementation and training • Ole Winther for supervising Søren & Casper • Søren Brunak for introducing me to the world of neural networks • The organizers for accepting our paper • You for listening! 20
- Slides: 20