Converting DNA Sequence file formats with Bio Python
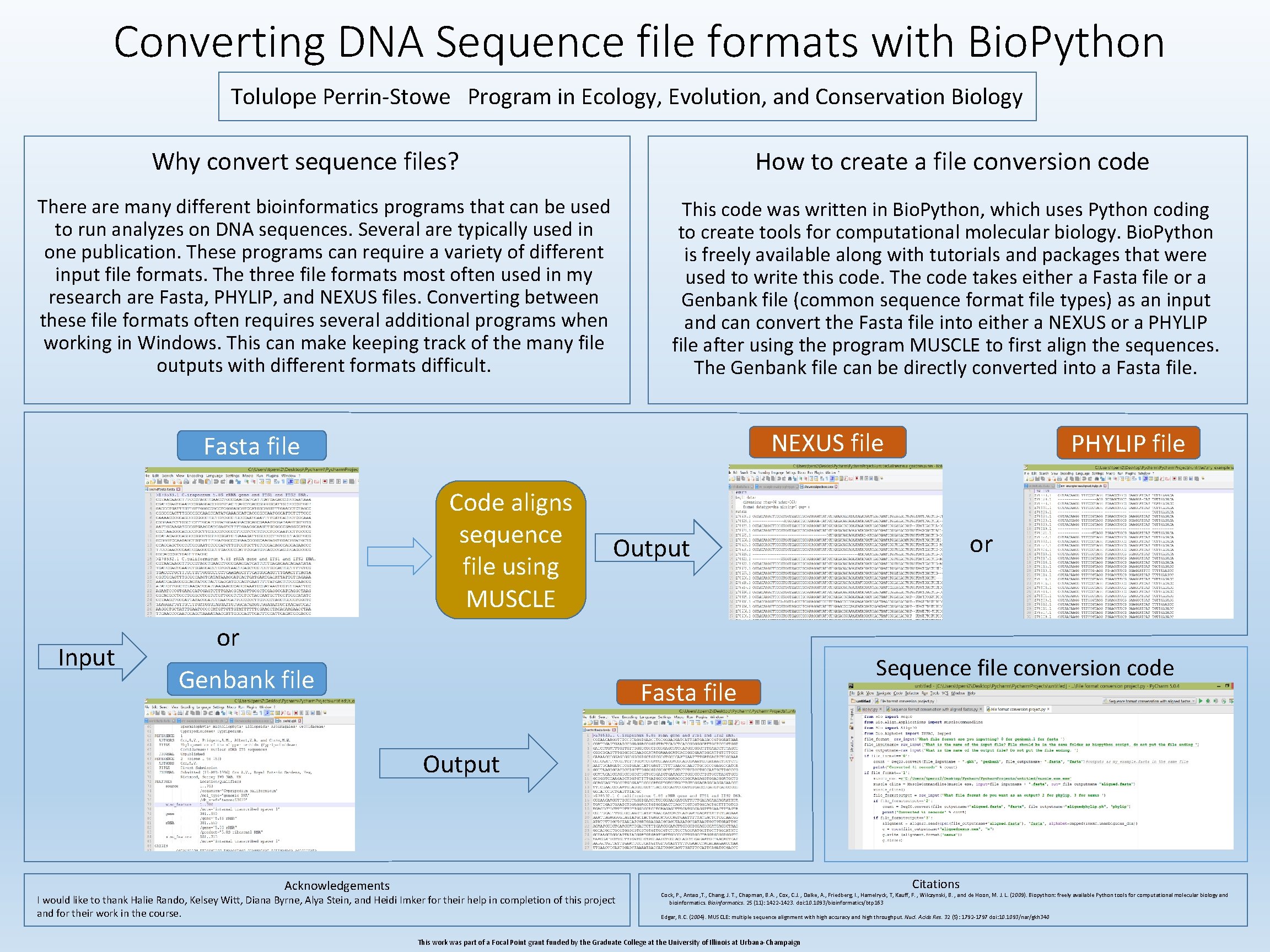
Converting DNA Sequence file formats with Bio. Python Tolulope Perrin-Stowe Program in Ecology, Evolution, and Conservation Biology How to create a file conversion code Why convert sequence files? There are many different bioinformatics programs that can be used to run analyzes on DNA sequences. Several are typically used in one publication. These programs can require a variety of different input file formats. The three file formats most often used in my research are Fasta, PHYLIP, and NEXUS files. Converting between these file formats often requires several additional programs when working in Windows. This can make keeping track of the many file outputs with different formats difficult. This code was written in Bio. Python, which uses Python coding to create tools for computational molecular biology. Bio. Python is freely available along with tutorials and packages that were used to write this code. The code takes either a Fasta file or a Genbank file (common sequence format file types) as an input and can convert the Fasta file into either a NEXUS or a PHYLIP file after using the program MUSCLE to first align the sequences. The Genbank file can be directly converted into a Fasta file. NEXUS file Fasta file Code aligns sequence file using MUSCLE Input PHYLIP file or Output or Genbank file Fasta file Sequence file conversion code Output Acknowledgements Citations I would like to thank Halie Rando, Kelsey Witt, Diana Byrne, Alya Stein, and Heidi Imker for their help in completion of this project and for their work in the course. Cock, P. , Antao , T. , Chang, J. T. , Chapman, B. A. , Cox, C. J. , Dalke, A. , Friedberg, I. , Hamelryck, T, Kauff, F. , Wilczynski, B. , and de Hoon, M. J. L. (2009). Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics. 25 (11): 1422 -1423. doi: 10. 1093/bioinformatics/btp 163 Edgar, R. C. (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucl. Acids Res. 32 (5): 1792 -1797 doi: 10. 1093/nar/gkh 340 This work was part of a Focal Point grant funded by the Graduate College at the University of Illinois at Urbana-Champaign
- Slides: 1