control of metabolic reactions making ve DGo reactions

control of metabolic reactions • making +ve DGo‘ reactions happen • points of control: DGo‘ and equilibrium • multi-active enzymes: enzyme complexes and multiple active sites

reactions with +ve DGo‘ can occur by: • coupling with a reaction with –ve DGo‘ • –ve physiological DG due to cellular low ratio [products]/[reactants]
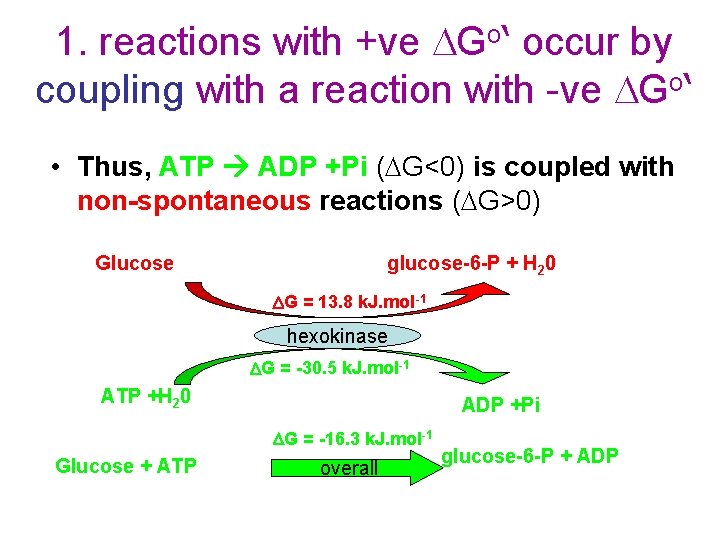
1. reactions with +ve DGo‘ occur by coupling with a reaction with -ve DGo‘ • Thus, ATP ADP +Pi (DG<0) is coupled with non-spontaneous reactions (DG>0) Glucose glucose-6 -P + H 20 DG = 13. 8 k. J. mol-1 hexokinase DG = -30. 5 k. J. mol-1 ATP +H 20 ADP +Pi DG = -16. 3 k. J. mol-1 Glucose + ATP overall glucose-6 -P + ADP

2. Recall: DGo' of a reaction may be positive, and DG negative, depending on cellular concentrations of reactants and products. Many reactions for which DGo' is positive are spontaneous in vivo because other reactions cause [products] or [substrate]. any [products] or [substrate] that moves the reaction away from equilibrium ratio causes reaction to proceed spontaneously forward to restore equilibrium
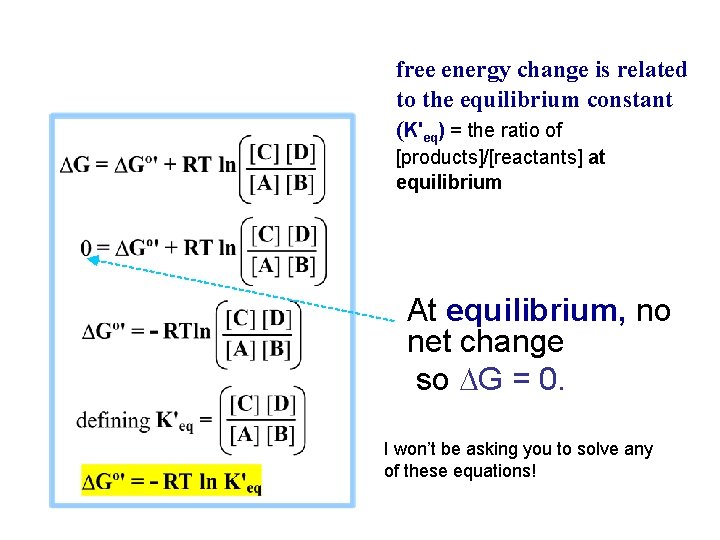
free energy change is related to the equilibrium constant (K'eq) = the ratio of [products]/[reactants] at equilibrium At equilibrium, no net change so DG = 0. I won’t be asking you to solve any of these equations!

many reactions are near equilibrium • then DG ~0 (no net change in free energy) • easily reversed by changing ratio of [products]/[substrate] as don’t need to overcome high DG For A+B ↔C+D product A+B C+D substrate A+B C+D • enzymes that catalyse such reactions act to restore equilibrium • rate regulated by [products]/[reactants]

Implication: a reaction near equilibrium may have +ve DGo' but be spontaneous in the cell -ve DG because other reactions cause [products] or [substrate].

Other reactions are FAR from equilibrium • enzyme rate is too slow to allow products to build to equilibrium concentration • [substrate] builds up in excess of Keq DG <<<0 (highly negative) • not affected by D [substrate] (saturated) • essentially irreversible • rate controlled by changing activity of enzyme (eg allosteric interactions) • reactions with DG <<0 are often sites of regulation

reactions with DG <<0 are often sites of regulation 1. Often occur early as a “committed step” in metabolic pathways (eg Acetyl. Co. A carboxylase) 2. most metabolic pathways are irreversible 1. ≥ 1 step with -ve DG required to drive: eg PDH, pyruvate carboxylase) 2. one way street: return by a different street 3. catabolic and anabolic pathways are separate independent control (eg glycolysis and gluconeogenesis (eg pyruvate carboxylase) use different enzymes)

Pyruvate dehydrogenase a pretty, pink multi-enzyme complex ‘gatekeeper’ to entry to citric acid cycle http: //www. brookscole. com/chemistry_d/templates/student_resources/shared_resources/animations/pdc. html
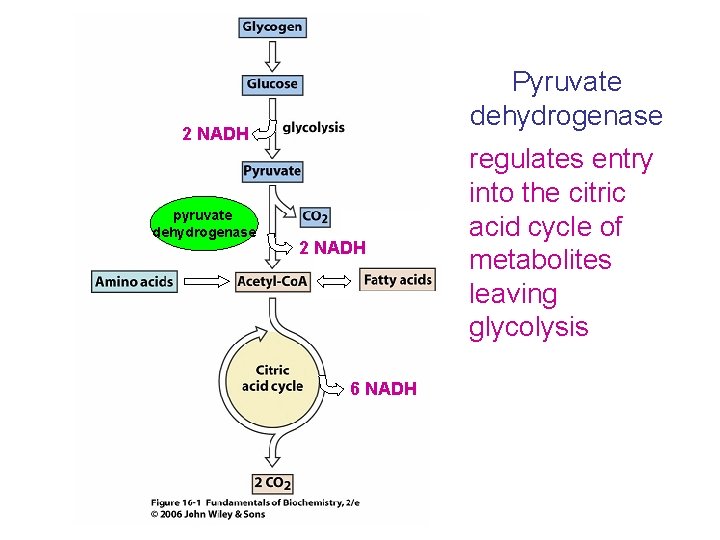
Pyruvate dehydrogenase 2 NADH pyruvate dehydrogenase 2 NADH 6 NADH regulates entry into the citric acid cycle of metabolites leaving glycolysis

Summary 1. structure of PDH complex 3 enzymes (E 1, E 2, E 3) 2. reactions of PDH complex 5 reactions, 5 cofactors 3. mechanism of PDH complex lipoamide swinging arm 4. regulation of PDH complex de/phosphorylation of E 1 product inhibition of E 2 and E 3 Excellent animation of PDH reactions if you can access it: (not examinable, but might help understanding!) http: //www. brookscole. com/chemistry_d/templates/student_resources/shared_resources/animations/pdc. html
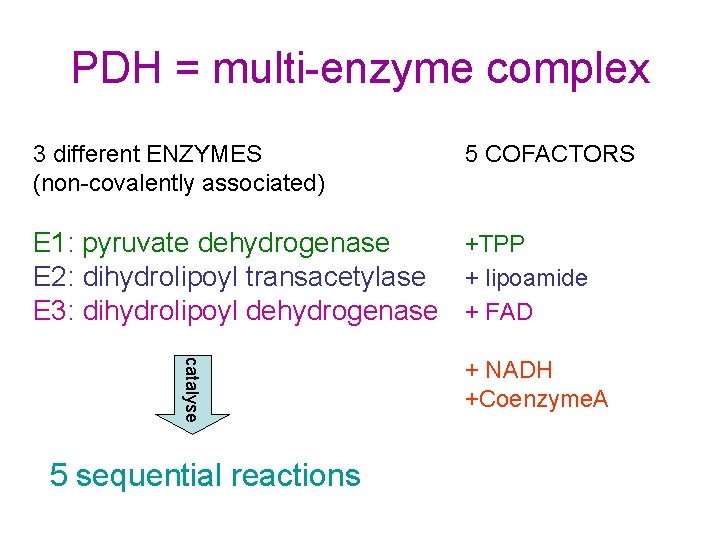
PDH = multi-enzyme complex 3 different ENZYMES (non-covalently associated) 5 COFACTORS E 1: pyruvate dehydrogenase +TPP E 2: dihydrolipoyl transacetylase + lipoamide E 3: dihydrolipoyl dehydrogenase + FAD catalyse 5 sequential reactions + NADH +Coenzyme. A
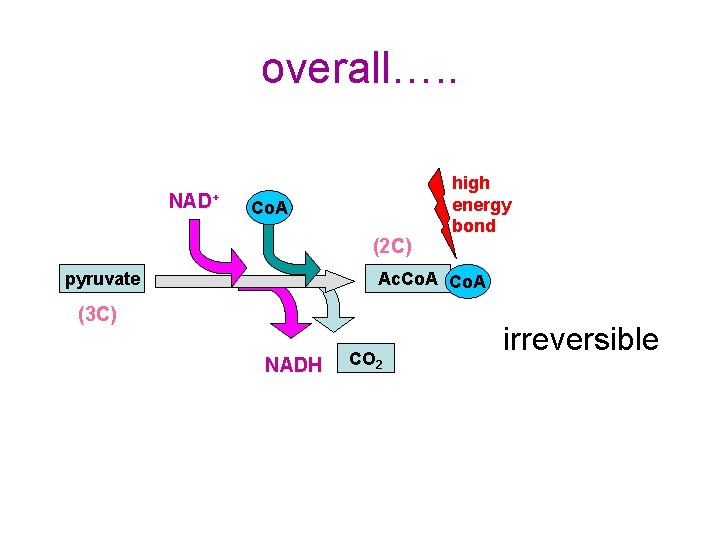
overall…. . NAD+ Co. A (2 C) pyruvate high energy bond Ac. Co. A (3 C) NADH CO 2 irreversible
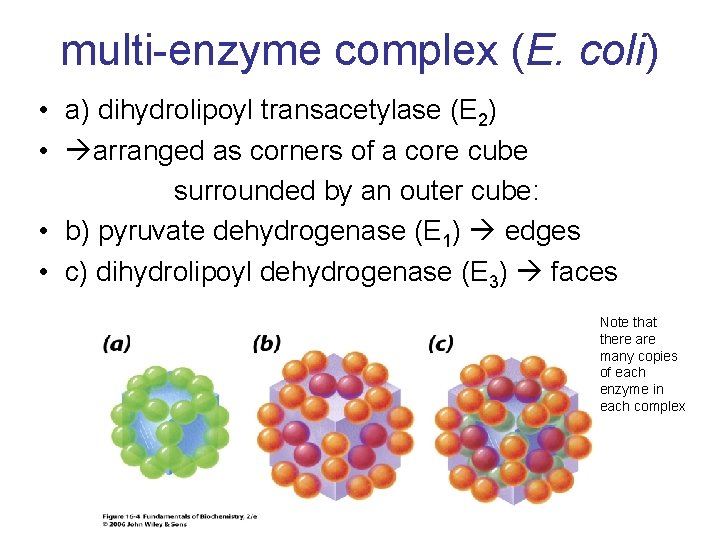
multi-enzyme complex (E. coli) • a) dihydrolipoyl transacetylase (E 2) • arranged as corners of a core cube surrounded by an outer cube: • b) pyruvate dehydrogenase (E 1) edges • c) dihydrolipoyl dehydrogenase (E 3) faces Note that there are many copies of each enzyme in each complex
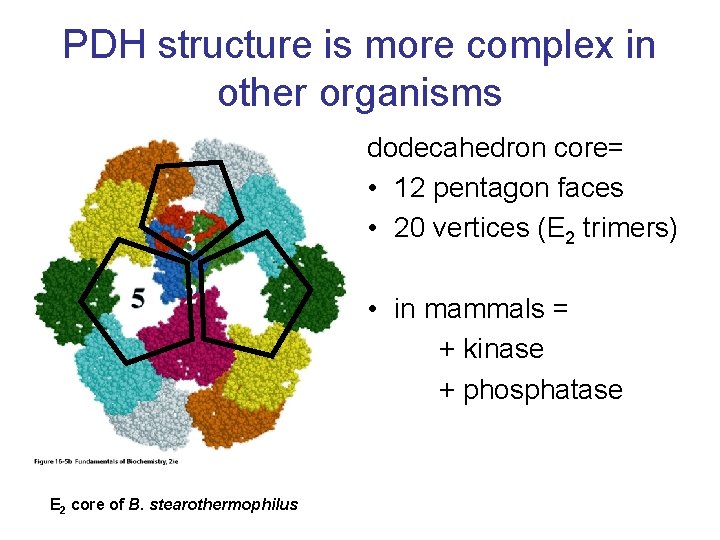
PDH structure is more complex in other organisms dodecahedron core= • 12 pentagon faces • 20 vertices (E 2 trimers) • in mammals = + kinase + phosphatase E 2 core of B. stearothermophilus
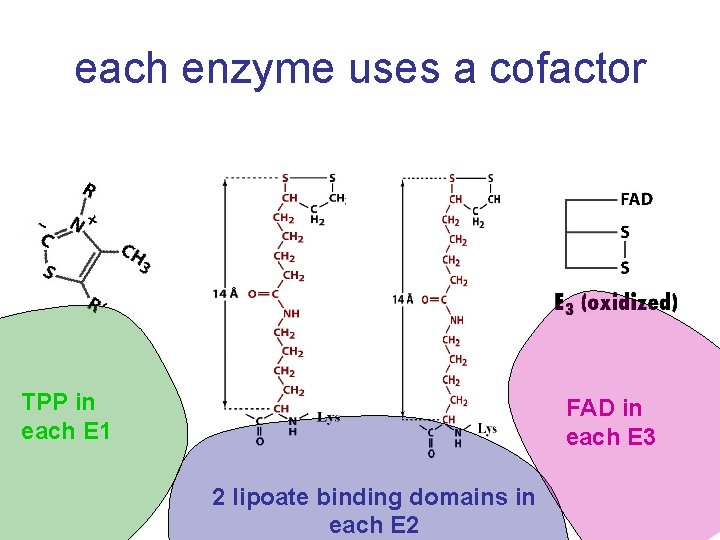
each enzyme uses a cofactor TPP in each E 1 FAD in each E 3 2 lipoate binding domains in each E 2
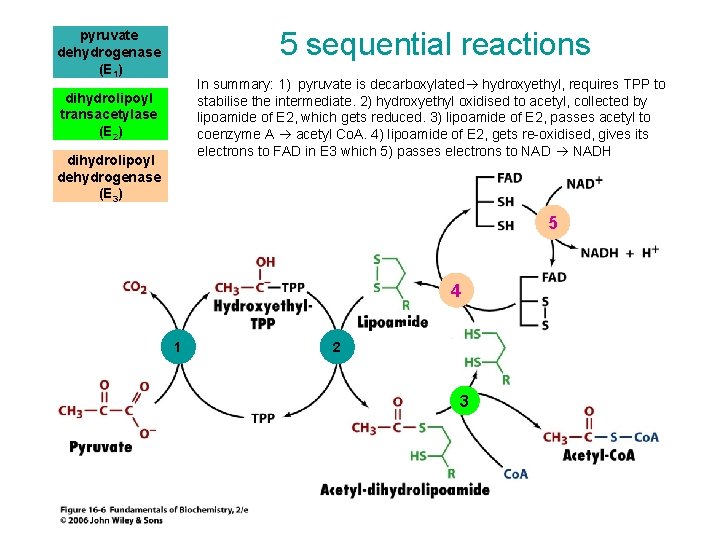
5 sequential reactions pyruvate dehydrogenase (E 1) In summary: 1) pyruvate is decarboxylated hydroxyethyl, requires TPP to stabilise the intermediate. 2) hydroxyethyl oxidised to acetyl, collected by lipoamide of E 2, which gets reduced. 3) lipoamide of E 2, passes acetyl to coenzyme A acetyl Co. A. 4) lipoamide of E 2, gets re-oxidised, gives its electrons to FAD in E 3 which 5) passes electrons to NADH dihydrolipoyl transacetylase (E 2) dihydrolipoyl dehydrogenase (E 3) 5 4 1 2 3
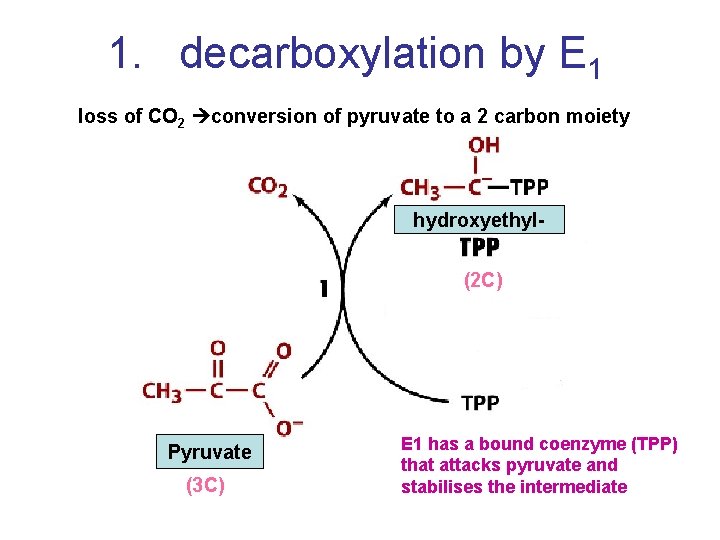
1. decarboxylation by E 1 loss of CO 2 conversion of pyruvate to a 2 carbon moiety hydroxyethyl(2 C) Pyruvate (3 C) E 1 has a bound coenzyme (TPP) that attacks pyruvate and stabilises the intermediate
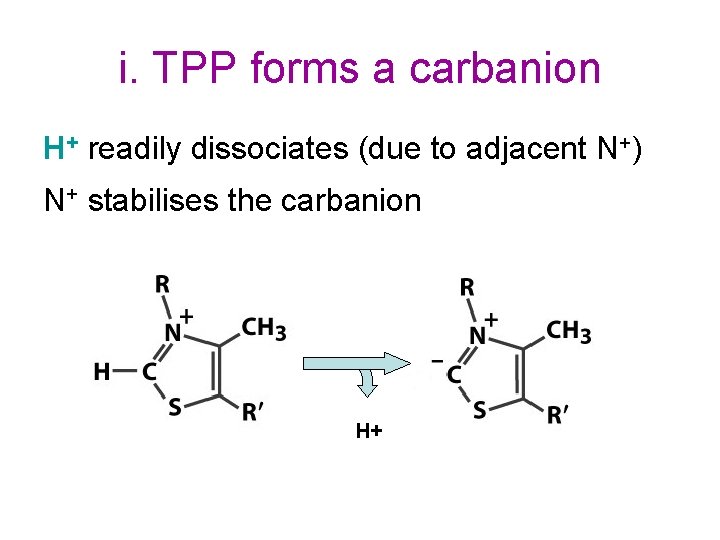
i. TPP forms a carbanion H+ readily dissociates (due to adjacent N+) N+ stabilises the carbanion H+
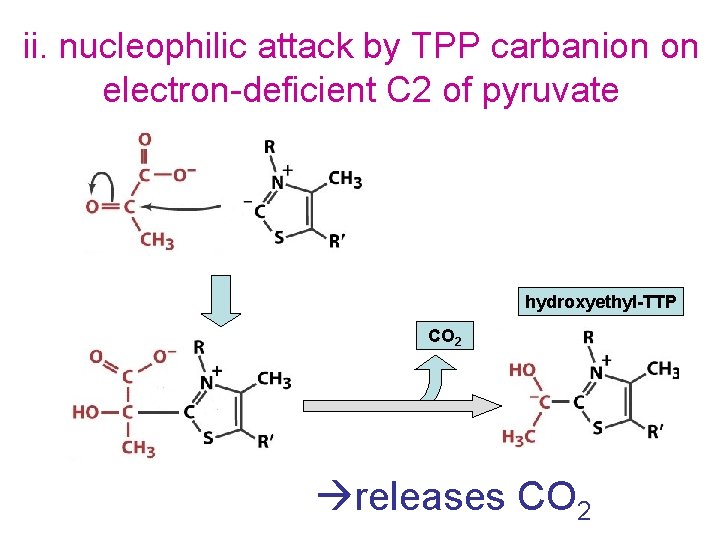
ii. nucleophilic attack by TPP carbanion on electron-deficient C 2 of pyruvate hydroxyethyl-TTP CO 2 releases CO 2
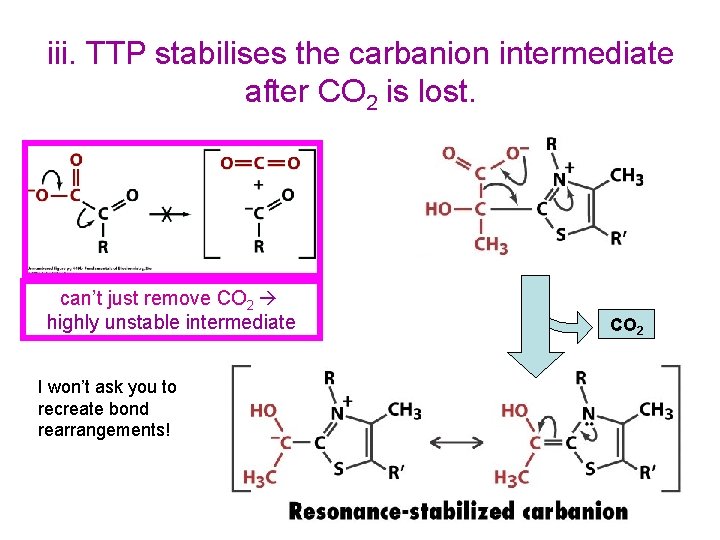
iii. TTP stabilises the carbanion intermediate after CO 2 is lost. can’t just remove CO 2 highly unstable intermediate I won’t ask you to recreate bond rearrangements! CO 2
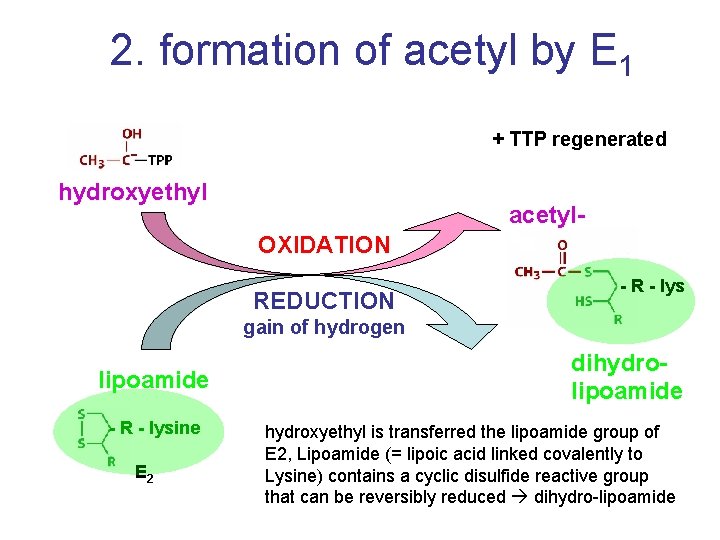
2. formation of acetyl by E 1 + TTP regenerated hydroxyethyl acetyl. OXIDATION REDUCTION - R - lys gain of hydrogen lipoamide - R - lysine E 2 dihydrolipoamide hydroxyethyl is transferred the lipoamide group of E 2, Lipoamide (= lipoic acid linked covalently to Lysine) contains a cyclic disulfide reactive group that can be reversibly reduced dihydro-lipoamide
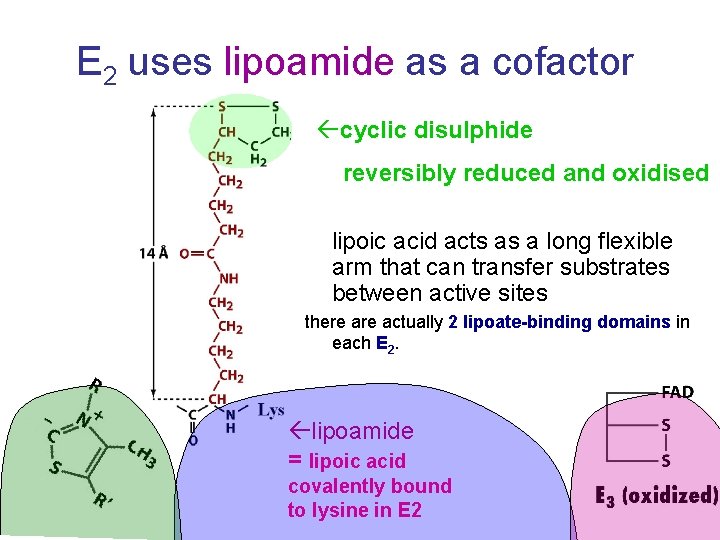
E 2 uses lipoamide as a cofactor cyclic disulphide reversibly reduced and oxidised lipoic acid acts as a long flexible arm that can transfer substrates between active sites there actually 2 lipoate-binding domains in each E 2. lipoamide = lipoic acid covalently bound to lysine in E 2
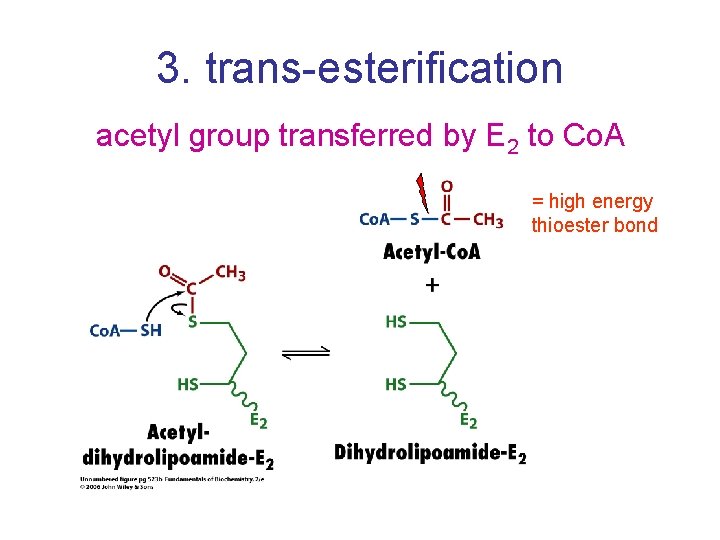
3. trans-esterification acetyl group transferred by E 2 to Co. A = high energy thioester bond

Thioesters: high energy bond • Form between carboxylic acid (COOH) and a thiol (SH) eg thiol in Coenzyme. A • eg Acetyl-Co. A is common to CHO, fat and protein metabolism • eg. In citric acid cycle, cleavage of thioester in succinyl-Co. A provides energy for synthesis of GTP

Lipoamide cofactor in E 2 So far…. • disulfide swings to outer shell to collect hydroxyethyl from TPP in E 1 • swings to E 2 to transfer acetyl to Co. A now we have acetyl-Co. A Remember: there are multiple copies of Kreb’s, FA synthesis each enzyme in complex next must regenerate lipoamide and produce NADH • So… lipoamide swings to E 3 to be reoxidised and transfer electrons to NADH via FAD
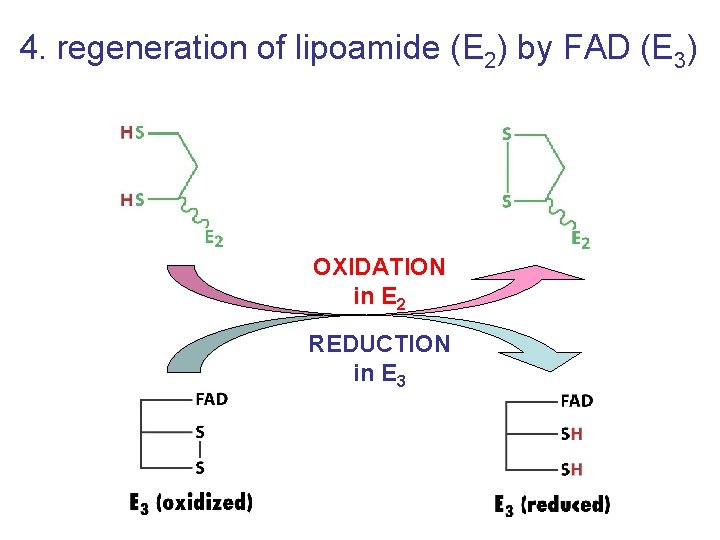
4. regeneration of lipoamide (E 2) by FAD (E 3) OXIDATION in E 2 REDUCTION in E 3
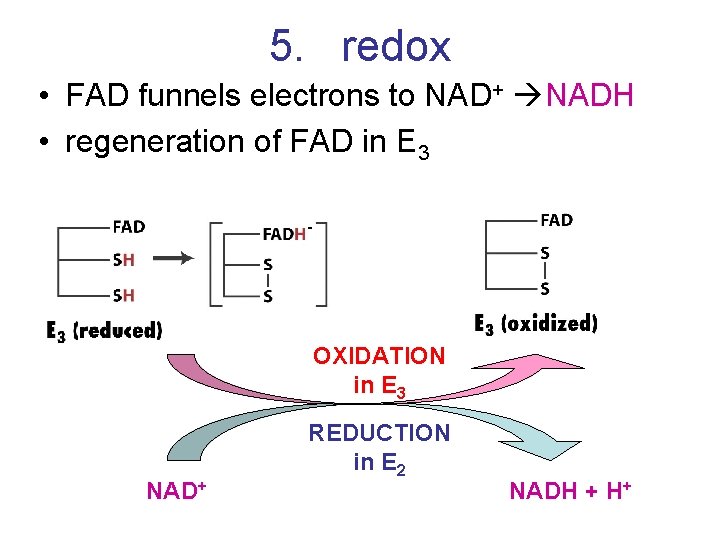
5. redox • FAD funnels electrons to NAD+ NADH • regeneration of FAD in E 3 OXIDATION in E 3 NAD+ REDUCTION in E 2 NADH + H+
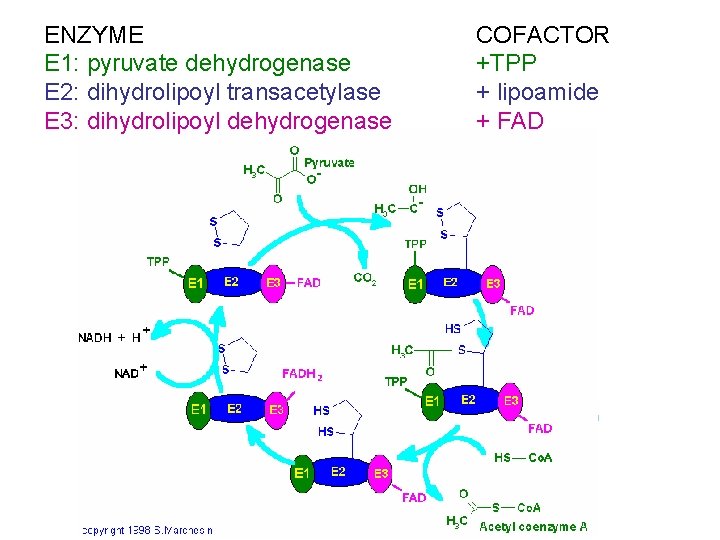
ENZYME E 1: pyruvate dehydrogenase E 2: dihydrolipoyl transacetylase E 3: dihydrolipoyl dehydrogenase COFACTOR +TPP + lipoamide + FAD
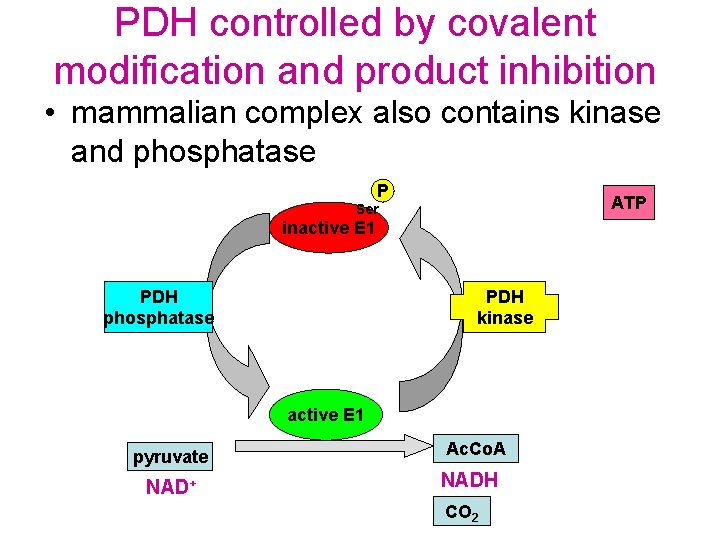
PDH controlled by covalent modification and product inhibition • mammalian complex also contains kinase and phosphatase P ATP Ser inactive E 1 PDH phosphatase PDH kinase active E 1 pyruvate NAD+ Ac. Co. A NADH CO 2
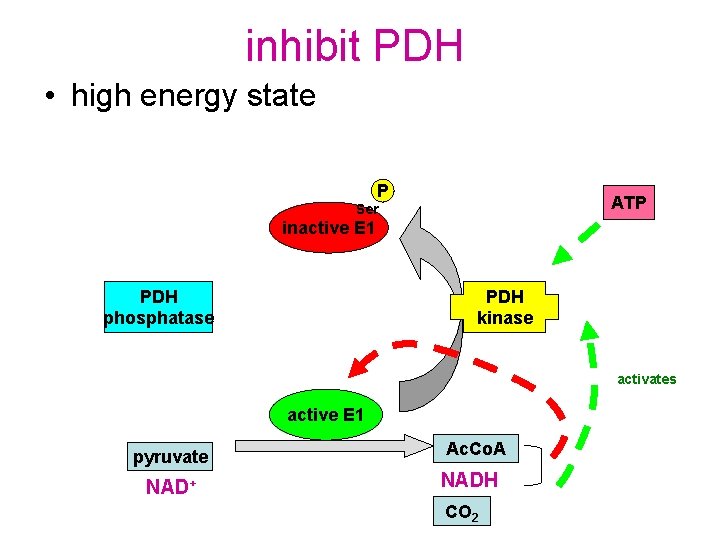
inhibit PDH • high energy state P ATP Ser inactive E 1 PDH phosphatase PDH kinase activates active E 1 pyruvate NAD+ Ac. Co. A NADH CO 2

inhibition by products in addition to activating PDH kinase, NADH and acetyl-Co. A: • compete with substrates for binding sites • drive E 2 and E 3 in reverse (these reactions are close to equilibrium) • E 2 not available to collect hydyrxyol from TPP • TPP cannot accept pyruvate
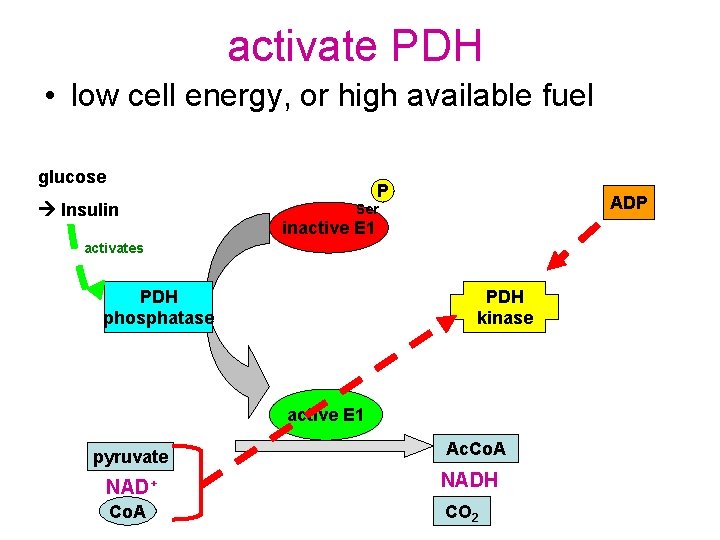
activate PDH • low cell energy, or high available fuel glucose Insulin P ADP Ser inactive E 1 activates PDH phosphatase PDH kinase active E 1 pyruvate Ac. Co. A NAD+ NADH Co. A CO 2
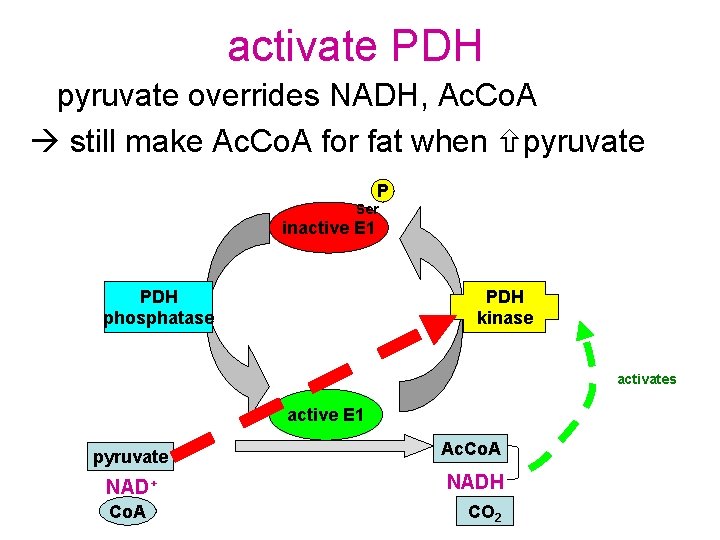
activate PDH pyruvate overrides NADH, Ac. Co. A still make Ac. Co. A for fat when pyruvate P Ser inactive E 1 PDH phosphatase PDH kinase activates active E 1 pyruvate Ac. Co. A NAD+ NADH Co. A CO 2
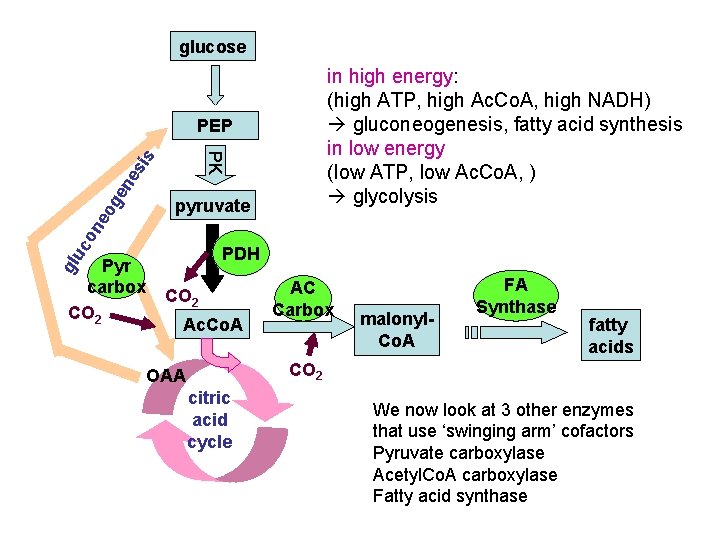
glucose in high energy: (high ATP, high Ac. Co. A, high NADH) gluconeogenesis, fatty acid synthesis in low energy (low ATP, low Ac. Co. A, ) glycolysis PEP on eo ge ne sis PK pyruvate uc PDH gl Pyr carbox CO 2 Ac. Co. A AC Carbox malonyl. Co. A FA Synthase fatty acids CO 2 OAA citric acid cycle We now look at 3 other enzymes that use ‘swinging arm’ cofactors Pyruvate carboxylase Acetyl. Co. A carboxylase Fatty acid synthase
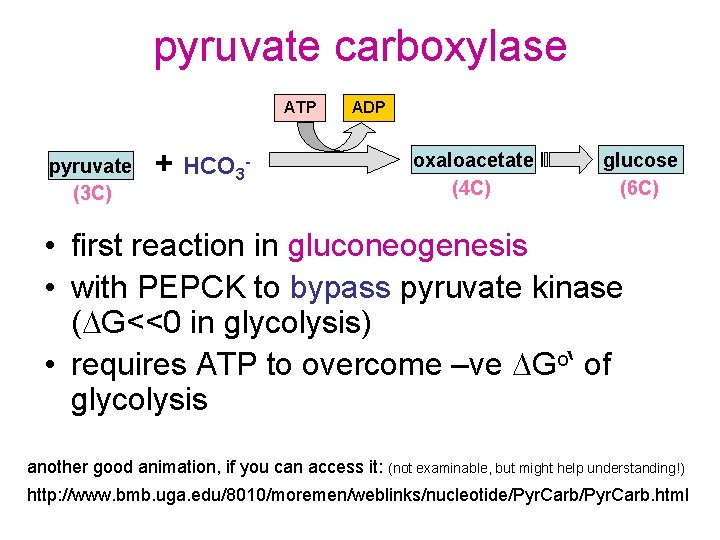
pyruvate carboxylase ATP pyruvate (3 C) + HCO 3 - ADP oxaloacetate (4 C) glucose (6 C) • first reaction in gluconeogenesis • with PEPCK to bypass pyruvate kinase (DG<<0 in glycolysis) • requires ATP to overcome –ve DGo‘ of glycolysis another good animation, if you can access it: (not examinable, but might help understanding!) http: //www. bmb. uga. edu/8010/moremen/weblinks/nucleotide/Pyr. Carb. html
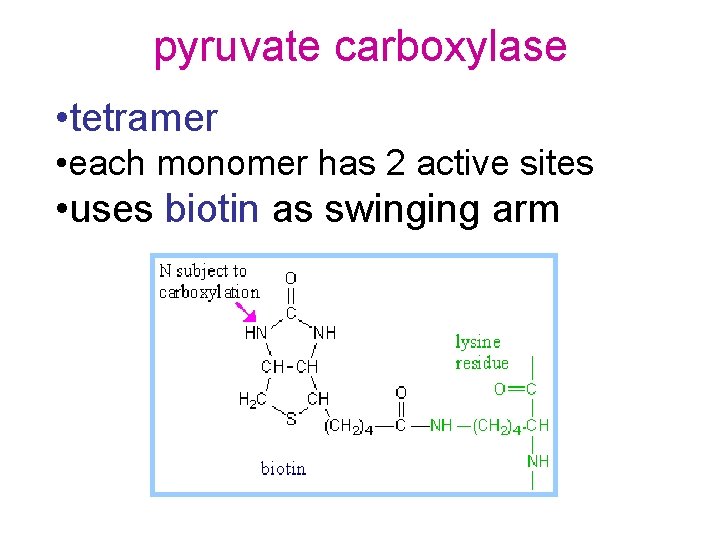
pyruvate carboxylase • tetramer • each monomer has 2 active sites • uses biotin as swinging arm
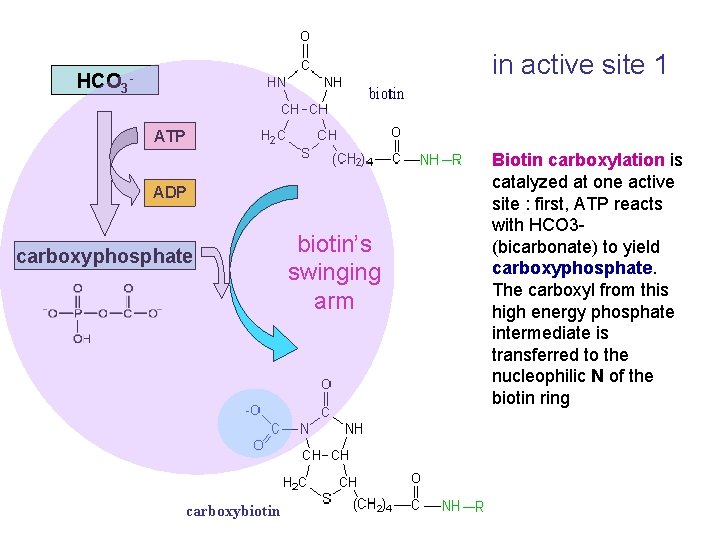
in active site 1 HCO 3 ATP biotin ADP carboxyphosphate carboxybiotin’s swinging arm Biotin carboxylation is catalyzed at one active site : first, ATP reacts with HCO 3(bicarbonate) to yield carboxyphosphate. The carboxyl from this high energy phosphate intermediate is transferred to the nucleophilic N of the biotin ring
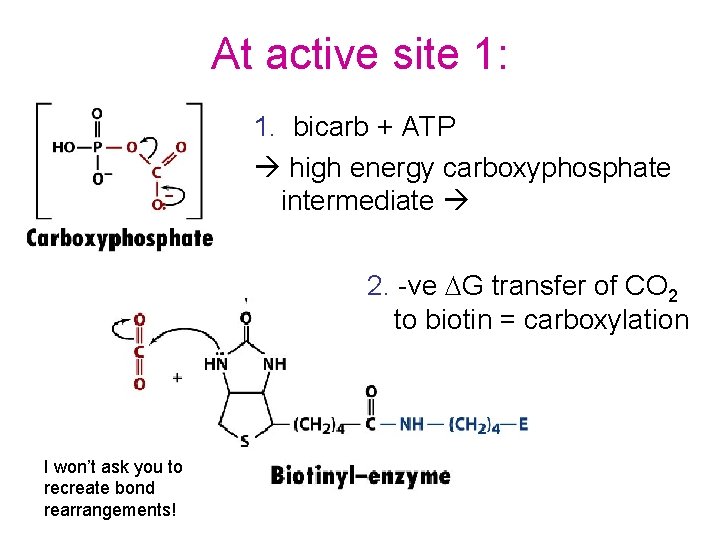
At active site 1: 1. bicarb + ATP high energy carboxyphosphate intermediate 2. -ve DG transfer of CO 2 to biotin = carboxylation I won’t ask you to recreate bond rearrangements!
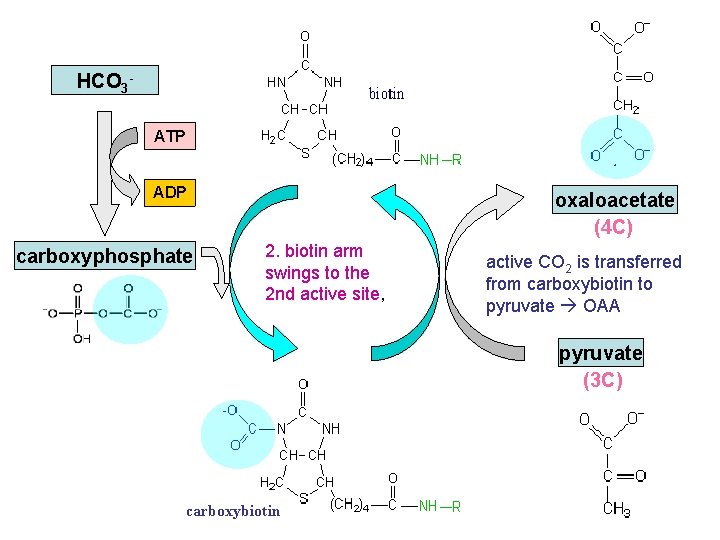
HCO 3 ATP biotin ADP carboxyphosphate oxaloacetate (4 C) 2. biotin arm swings to the 2 nd active site, active CO 2 is transferred from carboxybiotin to pyruvate OAA pyruvate (3 C) carboxybiotin
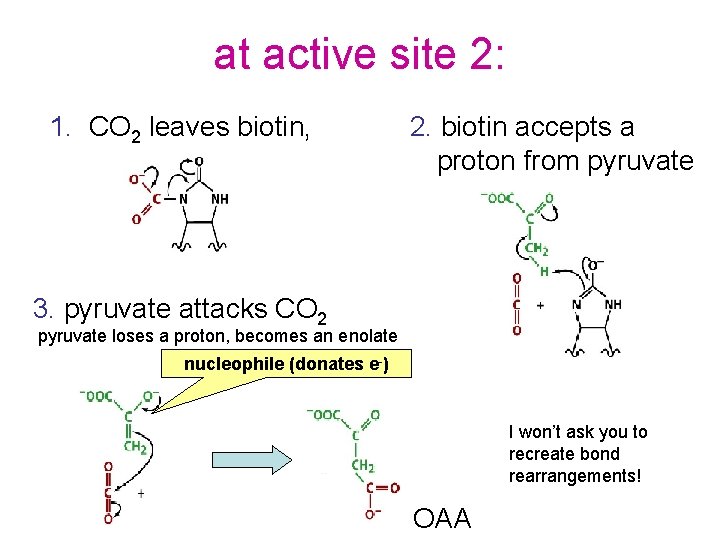
at active site 2: 1. CO 2 leaves biotin, 2. biotin accepts a proton from pyruvate 3. pyruvate attacks CO 2 pyruvate loses a proton, becomes an enolate nucleophile (donates e-) I won’t ask you to recreate bond rearrangements! OAA
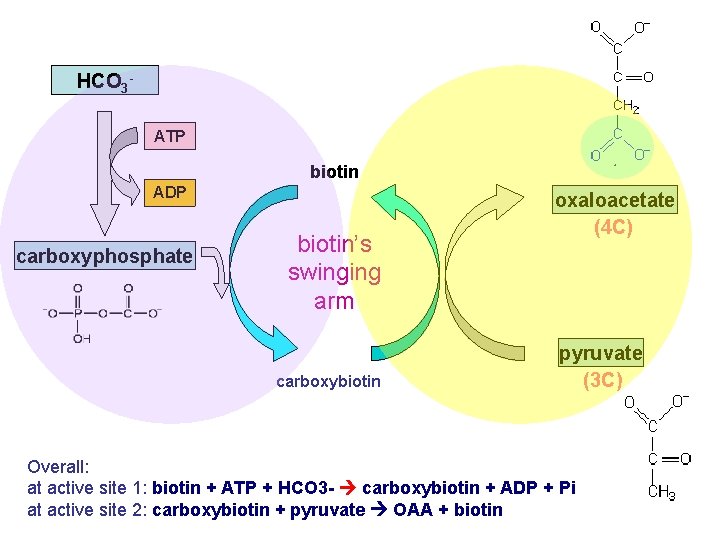
HCO 3 ATP biotin ADP carboxyphosphate biotin’s swinging arm carboxybiotin oxaloacetate (4 C) pyruvate (3 C) Overall: at active site 1: biotin + ATP + HCO 3 - carboxybiotin + ADP + Pi at active site 2: carboxybiotin + pyruvate OAA + biotin
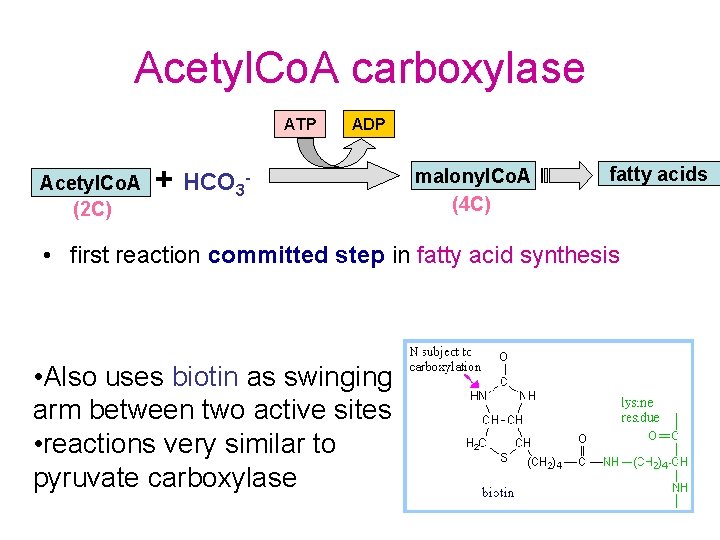
Acetyl. Co. A carboxylase ATP Acetyl. Co. A (2 C) ADP + HCO 3 - malonyl. Co. A (4 C) fatty acids • first reaction committed step in fatty acid synthesis • Also uses biotin as swinging arm between two active sites • reactions very similar to pyruvate carboxylase
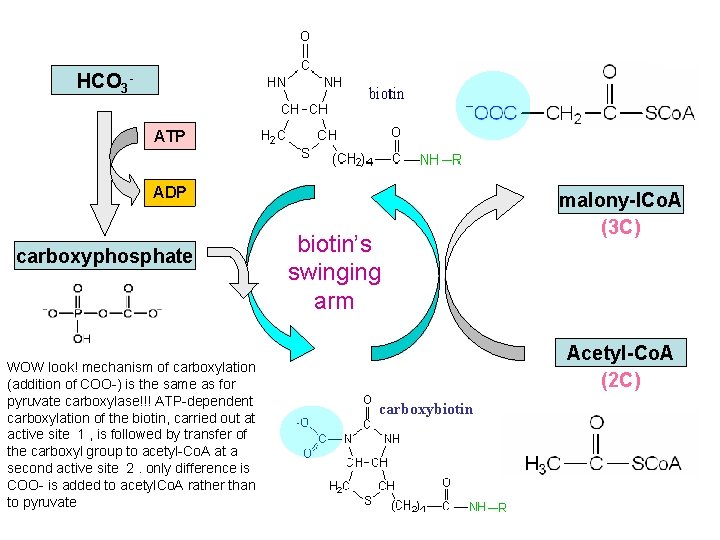
HCO 3 ATP biotin ADP carboxyphosphate WOW look! mechanism of carboxylation (addition of COO-) is the same as for pyruvate carboxylase!!! ATP-dependent carboxylation of the biotin, carried out at active site 1 , is followed by transfer of the carboxyl group to acetyl-Co. A at a second active site 2. only difference is COO- is added to acetyl. Co. A rather than to pyruvate biotin’s swinging arm malony-l. Co. A (3 C) Acetyl-Co. A (2 C) carboxybiotin
regulation of Ac. Co. A-Carboxylase The mammalian enzyme is regulated, by w phosphorylation by c. AMP dependent kinase w inhibition when energy ( c. AMP) w allosteric control by local metabolites. Conformational changes with regulation: w active = multimeric filamentous complexes. w inactive = dissociation to = monomeric form P

fatty acid synthase • dimer • 6 active sites are individual domains of a large protein – ? developed from gene fusion – has more catalytic activities than any enzyme! • has two prosthetic groups thioester bonds – thiol of cysteine (in condensing domain) – thiol of P-pantetheine (in acyl carrier domain) • acts as a long flexible arm transferring substrates between active sites
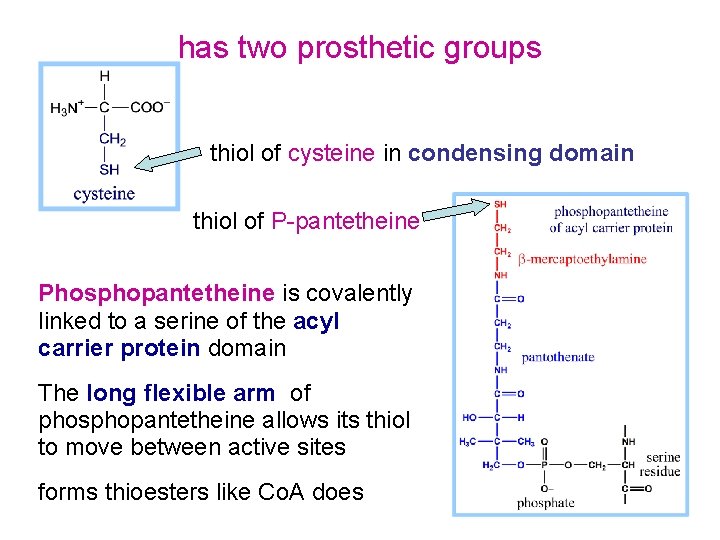
has two prosthetic groups thiol of cysteine in condensing domain thiol of P-pantetheine Phosphopantetheine is covalently linked to a serine of the acyl carrier protein domain The long flexible arm of phosphopantetheine allows its thiol to move between active sites forms thioesters like Co. A does
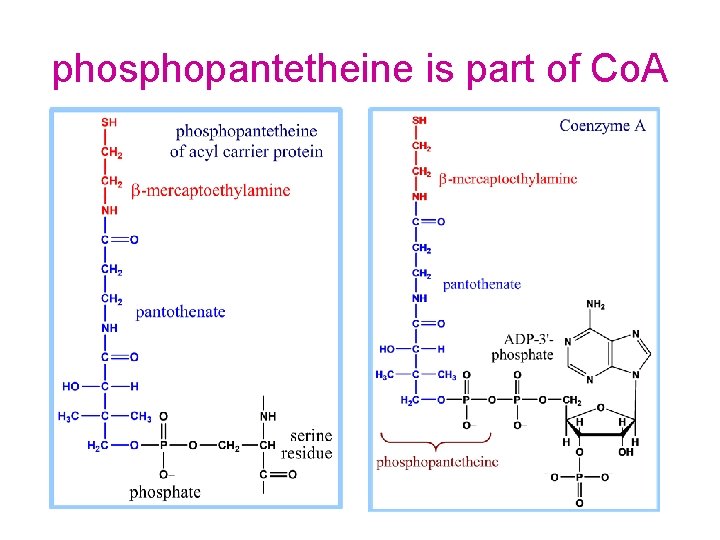
phosphopantetheine is part of Co. A
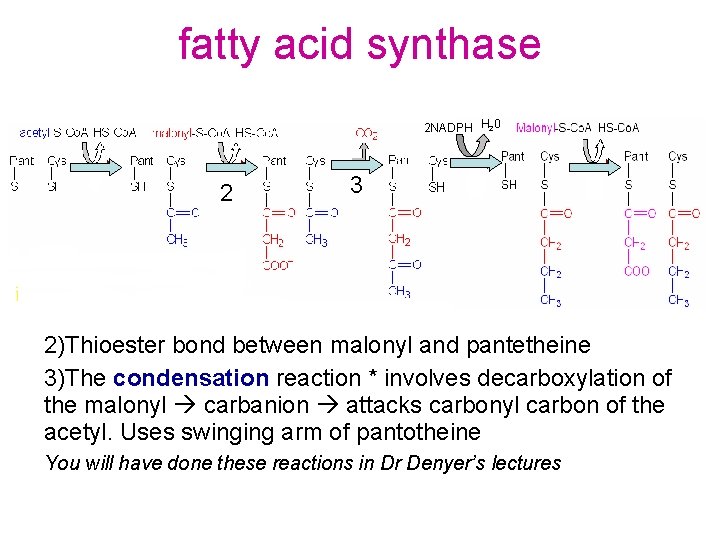
fatty acid synthase 2 NADPH H 20 2 3 2)Thioester bond between malonyl and pantetheine 3)The condensation reaction * involves decarboxylation of the malonyl carbanion attacks carbonyl carbon of the acetyl. Uses swinging arm of pantotheine You will have done these reactions in Dr Denyer’s lectures
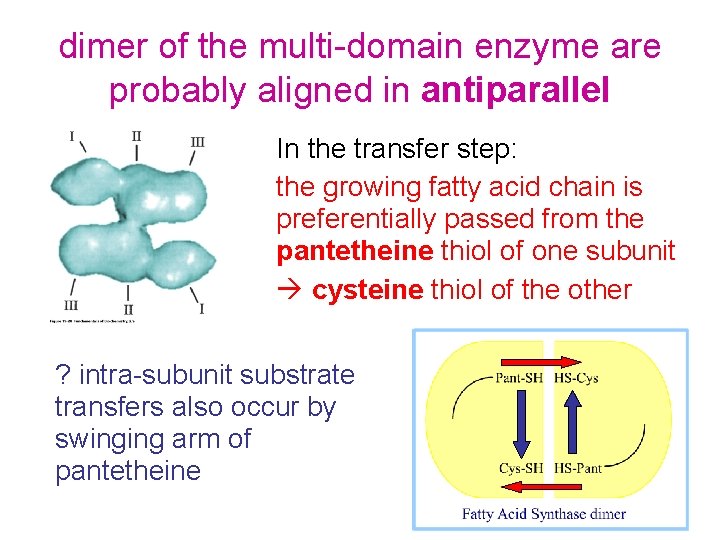
dimer of the multi-domain enzyme are probably aligned in antiparallel In the transfer step: the growing fatty acid chain is preferentially passed from the pantetheine thiol of one subunit cysteine thiol of the other ? intra-subunit substrate transfers also occur by swinging arm of pantetheine

essential dietary cofactors: cannot be made by mammals • thiamine = vitamin B 1 (in TPP) – deficiency = beri-beri – eg alcohol reduced uptakeof thiamine brain symptoms ( brain glucose metabolism) • • • riboflavin = vitamin B 2 (for FAD) niacin = vitamin B 3 (NAD) lipoic acid biotin pantothenic acid (vitamin B 5)

advantages of multi-active site enzymes and multi enzyme complexes • diffusion distance between substrate and active sites (usually the limiting factor in determining the reaction rate) reaction rate • chance of side reactions – substrates stay within complex • coordinated control of sequential reactions

Voet, Voet and Pratt (2 nd Ed) • • . DG and equilibrium pg 401 PDH pg 519 -524, regulation pg 533 TPP mechanism pg 450 thioester bonds pg 413 Pyruvate carboxylase pg 502, pg Acetyl. Co. A carboxylase pg 651 Fatty acid synthase pg 653 (much more detail than you need for this lecture!)
- Slides: 54