Constraintbased Modeling of Metabolic Networks Zhuo Wang Outline

Constraint-based Modeling of Metabolic Networks Zhuo Wang

Outline n n n Introduction to metabolism and metabolic networks Constraint-based modeling Mathematical formulation and methods ¨ Linear n programming Case study ¨ Human tissue-specific metabolic behavior ¨ Overproduction for microbial metabolic engineering
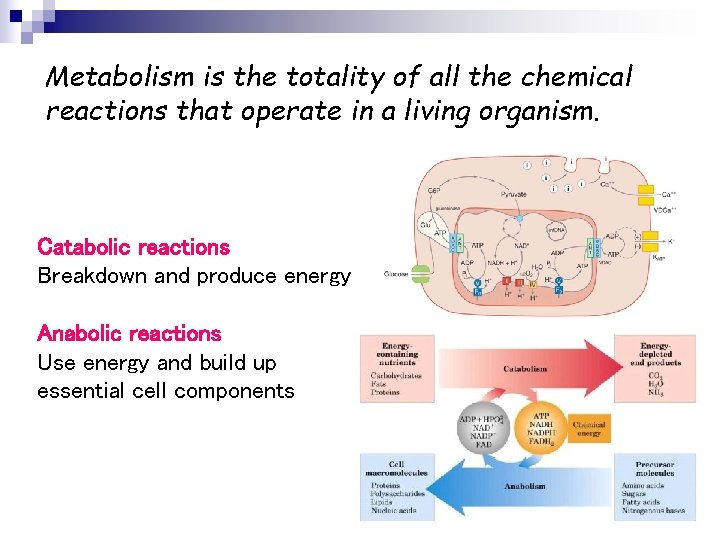
Metabolism is the totality of all the chemical reactions that operate in a living organism. Catabolic reactions Breakdown and produce energy Anabolic reactions Use energy and build up essential cell components

Why Study Metabolism? n It’s the essence of life. . n Tremendous importance in Medicine: In born errors of metabolism cause acute symptoms and even death on early age n Metabolic diseases (obesity肥胖症, diabetics糖尿病) are major sources of morbidity and mortality n Metabolic enzymes and their regulators gradually becoming viable drug targets n n Bioengineering: n n Efficient production of biological products The best understood cellular network
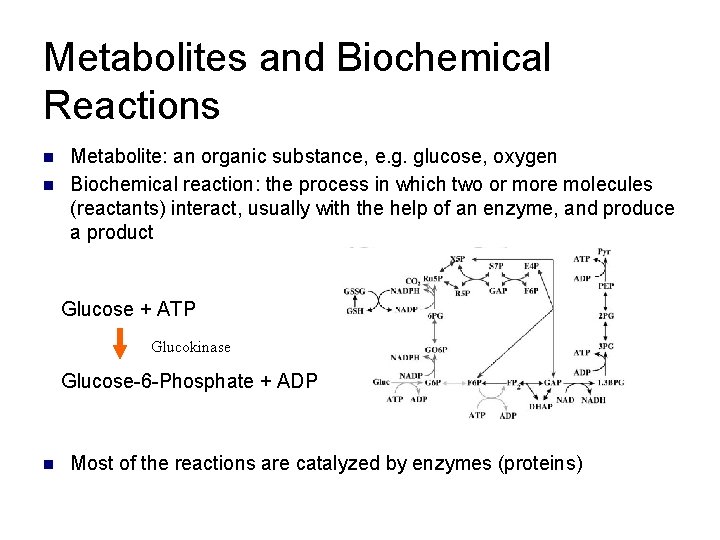
Metabolites and Biochemical Reactions n n Metabolite: an organic substance, e. g. glucose, oxygen Biochemical reaction: the process in which two or more molecules (reactants) interact, usually with the help of an enzyme, and produce a product Glucose + ATP Glucokinase Glucose-6 -Phosphate + ADP n Most of the reactions are catalyzed by enzymes (proteins)

Modeling the Network Function: Kinetic Models n n Dynamics of metabolic behavior over time ¨ Metabolite concentrations ¨ Enzyme activity rate – depends on enzyme concentrations and metabolite concentrations ¨ Solved using a set of differential equations Impossible to model large-scale networks ¨ Requires specific enzyme rates data ¨ Too complicated

Modeling the Network Function Kinetic models Accuracy • Dynamical systems • Requires kinetic constants (mostly unknown) Approx. kinetics Metabolic Constraint-based models • Optimization theory • Constrained space of possible, steady-state network behaviors • Probabilistic models, discrete models, etc’ Scale PPI Conventional functional models Topological analysis • Graph theory • Structural network properties: degree distribution, centrality, clusters, etc’
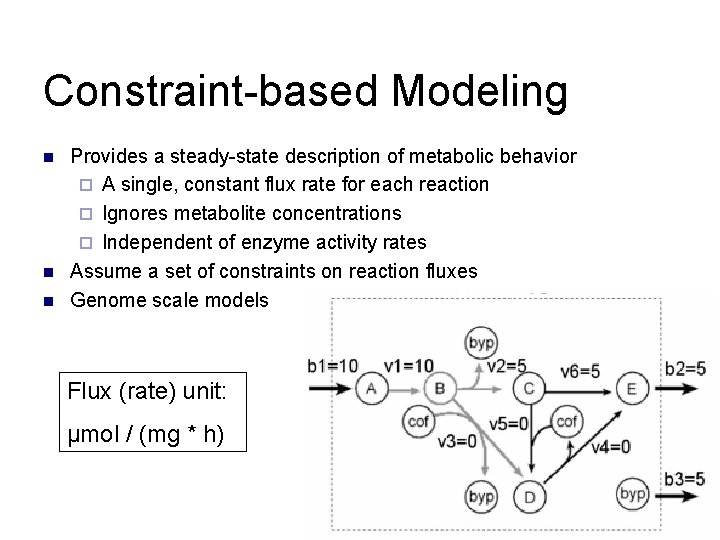
Constraint-based Modeling n n n Provides a steady-state description of metabolic behavior ¨ A single, constant flux rate for each reaction ¨ Ignores metabolite concentrations ¨ Independent of enzyme activity rates Assume a set of constraints on reaction fluxes Genome scale models Flux (rate) unit: μmol / (mg * h)

Constraint Based Modeling n Find a steady-state flux distribution through all biochemical reactions n Under the constraints: ¨ Mass balance: metabolite production and consumption rates are equal ¨ Thermodynamic: irreversibility of reactions ¨ Enzymatic capacity: bounds on enzyme rates ¨ Availability of nutrients

Additional Constraints n n n Transcriptional regulatory constraints (Covert, et. al. , 2002) ¨ Boolean representation of regulatory network Energy balance analysis (Beard, et. al. , 2002) ¨ Loops are not feasible according to thermodynamic principles Reaction directionality ¨ Depending on metabolite concentrations FBA solution space Meaningful solutions
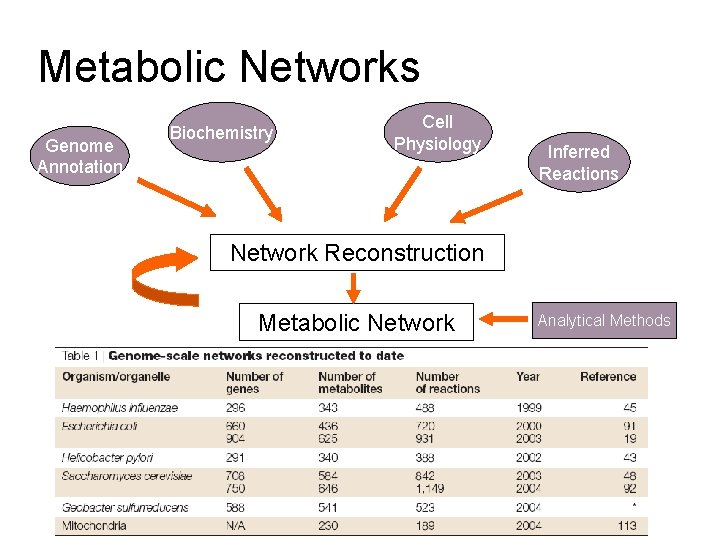
Metabolic Networks Genome Annotation Biochemistry Cell Physiology Inferred Reactions Network Reconstruction Metabolic Network Analytical Methods

Constraint-based modeling applications n Phenotype predictions: ¨ ¨ ¨ n Bioengineering: ¨ n Strain design – overproduce desired compounds Biomedical: ¨ n Growth rates across media Knockout lethality Nutrient uptake/secretion rates Intracellular fluxes Growth rate following adaptive evolution Predict drug targets for metabolic disorders Studying an array of questions regarding: Dispensability of metabolic genes ¨ Robustness and evolution of metabolic networks ¨

Phenotype Predictions: Knockout Lethality in E. coli n 86% of the predictions were consistent with the experimental observations
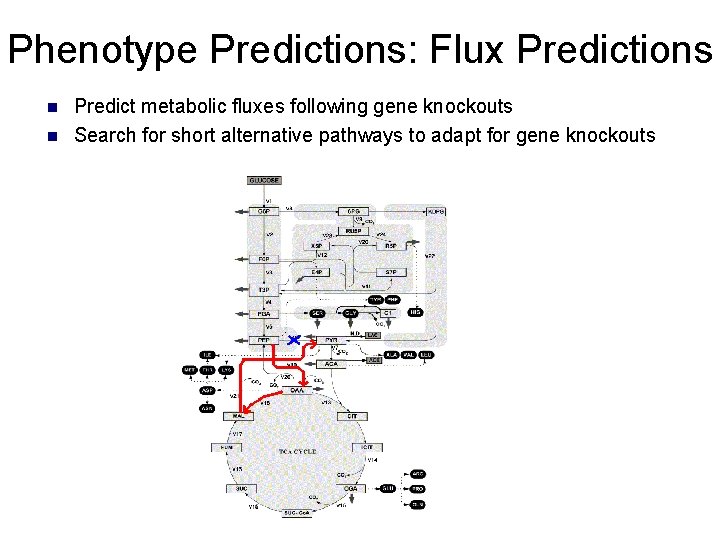
Phenotype Predictions: Flux Predictions n n Predict metabolic fluxes following gene knockouts Search for short alternative pathways to adapt for gene knockouts

Strain design: maximizing metabolite production rate n n Identify a set of gene whose knockout increases the production rate of some metabolite The knockout of reaction v 3 increases the production rate of metabolite F

Constraint-based Modeling: Mathematical Representation
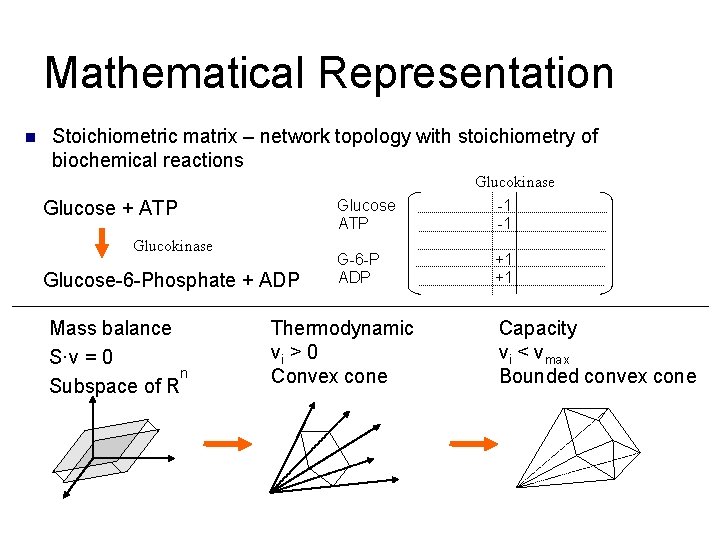
Mathematical Representation n Stoichiometric matrix – network topology with stoichiometry of biochemical reactions Glucokinase Glucose + ATP Glucokinase Glucose-6 -Phosphate + ADP Mass balance S·v = 0 n Subspace of R Glucose ATP -1 -1 G-6 -P ADP +1 +1 Thermodynamic vi > 0 Convex cone Capacity vi < vmax Bounded convex cone

Determination of Likely Physiological States n n n How to identify plausible physiological states? Optimization methods ¨ Maximal biomass production rate ¨ Minimal ATP production rate ¨ Minimal nutrient uptake rate Exploring the solution space ¨ Extreme pathways ¨ Elementary modes

Biomass Production Optimization n n Metabolic demands of precursors and cofactors required for 1 g of biomass of E. coli Classes of macromolecules: Amino Acids, Carbohydrates Ribonucleotides, Deoxyribonucleotides Lipids, Phospholipids Sterol, Fatty acids These precursors are removed from the metabolic network in the corresponding ratios n We define a growth reaction Z = 41. 2570 VATP - 3. 547 VNADH+18. 225 VNADPH + …. n
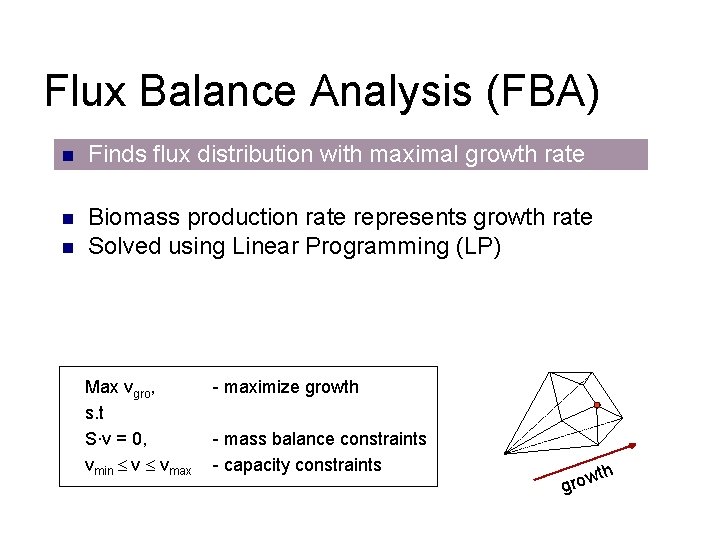
Flux Balance Analysis (FBA) n Finds flux distribution with maximal growth rate n Biomass production rate represents growth rate Solved using Linear Programming (LP) n Max vgro, s. t S∙v = 0, vmin v vmax - maximize growth - mass balance constraints - capacity constraints th w gro
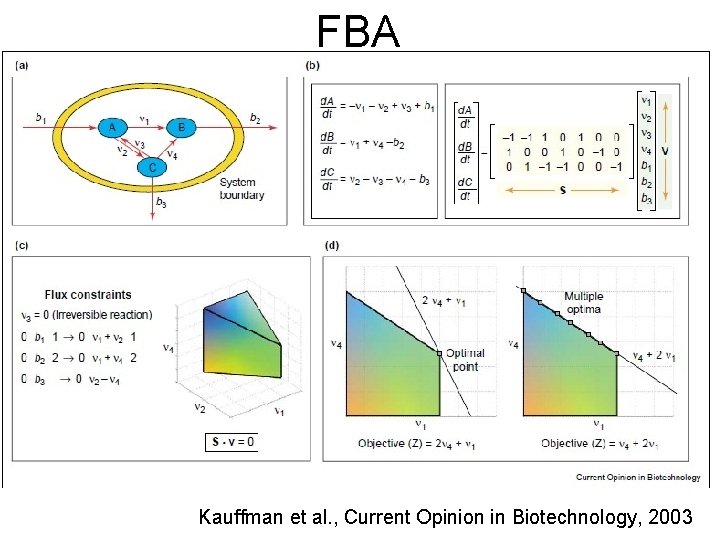
FBA Kauffman et al. , Current Opinion in Biotechnology, 2003
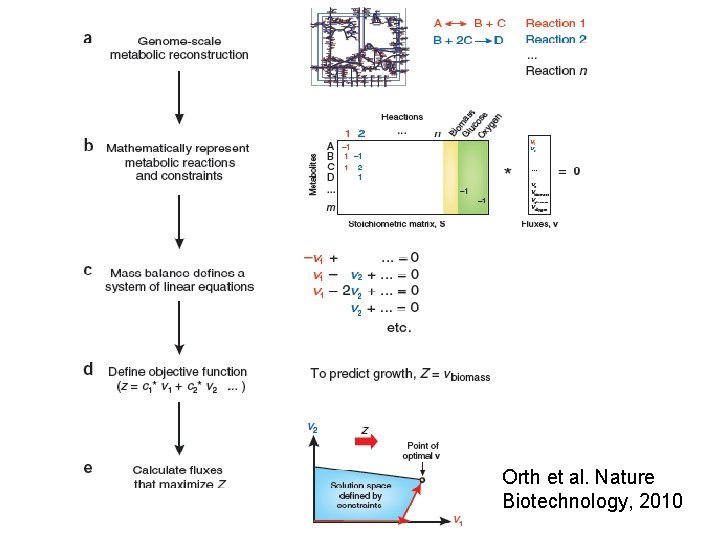
Orth et al. Nature Biotechnology, 2010

FBA Example (1)

FBA Example (2)

FBA Example (2)

Linear Programming Basics (1)
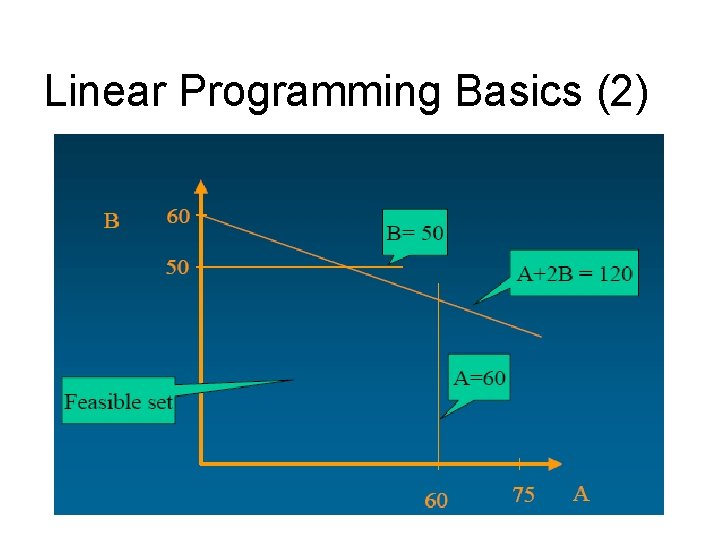
Linear Programming Basics (2)
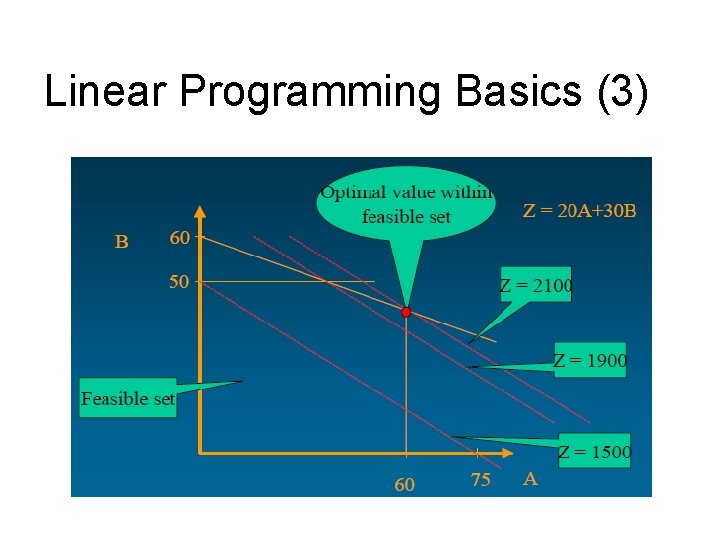
Linear Programming Basics (3)
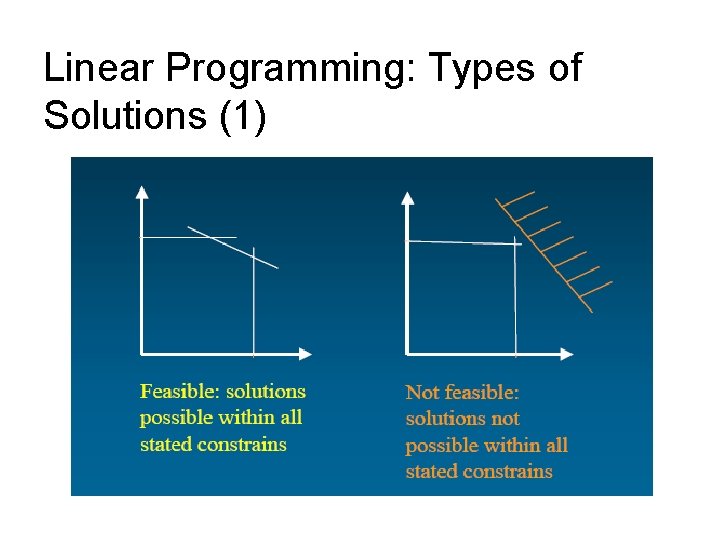
Linear Programming: Types of Solutions (1)

Linear Programming: Types of Solutions (2)

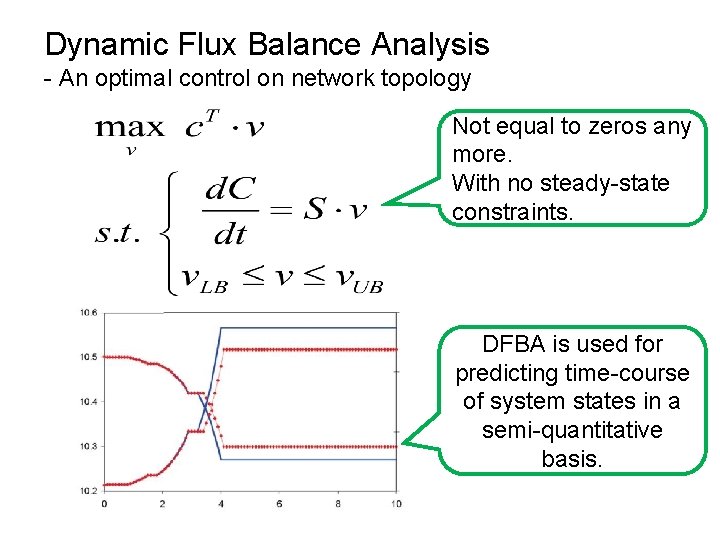
Dynamic Flux Balance Analysis - An optimal control on network topology Not equal to zeros any more. With no steady-state constraints. DFBA is used for predicting time-course of system states in a semi-quantitative basis.

Tools for metabolic flux analysis n n Flux. Analyzer: FBA COBRA Toolbox: FBA, DFBA, MOMA,sampling Flux. Explorer: FBA/MOMA,Manually construction network Bio-SPICE: Integrative platform with GUI http: //biospice. sourceforge. net/#Bio-SPICE_Core Fluxor Computational Analyzer – FBA Fluxor Spreadsheet - MOMA

COBRA n n n Becker, S. et al. , "Quantitative prediction of cellular metabolism with constraint-based models: The COBRA Toolbox", Nat. Protoc 2, 727 -738 (2007). Schellenberger J, Que R, Fleming RMT, Thiele I, Orth JD, Feist AM, Zielinski DC, Bordbar A, Lewis NE, Rahmanian S et al. , Quantitative prediction of cellular metabolism with constraint-based models: the COBRA Toolbox v 2. 0 Nat Protoc 6(9): 1290 -307 (2011). Ebrahim A, Lerman JA, Palsson BO, Hyduke DR. COBRApy: COnstraints-Based Reconstruction and Analysis for Python BMC Syst Bio 7: 74 (2013).

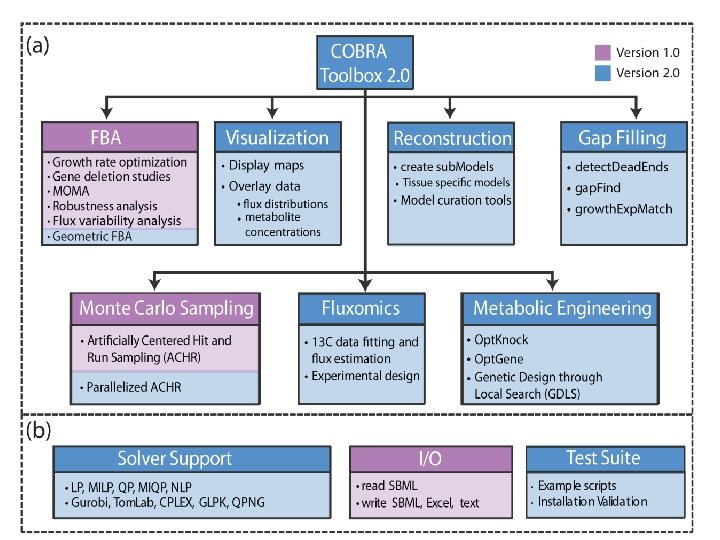
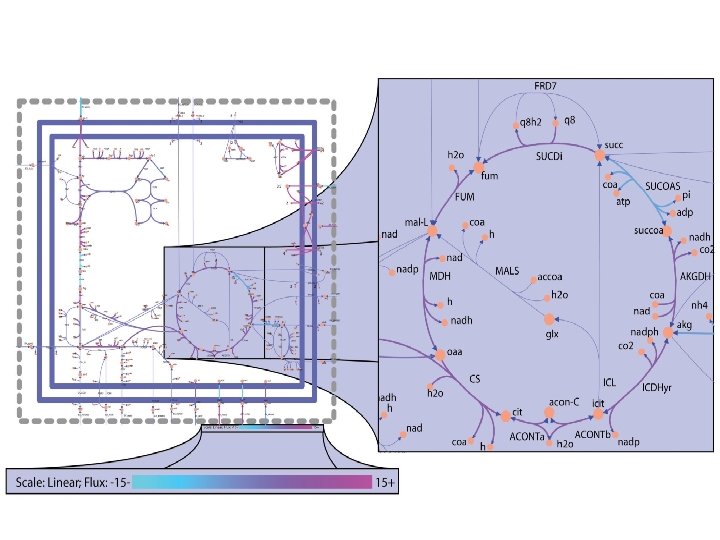

COBRA application n n COnstraint-Based Reconstruction and Analysis (COBRA) methods have been successfully employed in the field of microbial metabolic engineering and are being extended to modeling transcriptional and signaling networks and the field of public health. Specifically, COBRA methods have been used to guide metabolic pathway engineering, to model pathogens and host-pathogen interactions and to assess the impact of disease states on human metabolism

Where to get the models n n The biological network models that are analyzed with COBRA methods are constructed in a bottom-up fashion from bibliomic and experimental data and thus represent Biochemically, Genetically, and Genomically (Bi. GG) consistent knowledgebases Literature

Materials n Equipment • The COBRA Toolbox version 3. 0(the latest stable version) • A computer capable of running MATLAB R 2018 a(the latest stable version) • lib. SBML programming library 5. 16. 0(the latest stable version)(http: //www. sbml. org) • SBMLToolbox version 4. 1. 0 (the latest stable version) for MATLAB to allow reading and writing models in SBML format (http: //www. sbml. org) • A linear programming (LP) solver. Currently the COBRA Toolbox supports: o Gurobi (Gurobi Optimization. http: //www. gurobi. com) through Gurobi Mex (http: //www. convexoptimization. com/wikimization/index. php/Gurobi_mex) o CPLEX (ILOG Inc. ) through Tomlab (Tomlab Optimization Inc. http: //tomopt. com) o GLPK (http: //www. gnu. org/software/glpk) through glpkmex (http: //glpkmex. sourceforge. net) – Note, GLPK does not provide accurate solutions for Opt. Knock or GDLS calculations as implemented in the Toolbox.

Functions Initializing the Toolbox 1| Navigate to the directory where you installed the Toolbox: ≫ init. Cobra. Toolbox() 2| Save the paths added if desired: ≫ savepath() n Changing COBRA solvers 3| Set the solvers used by the COBRA Toolbox using the following function: ≫ change. Cobra. Solver(solver. Name, [solver. Type]) n

Run COBRA Toolbox test suite ≫ test. All() n Read and write SBML model ≫ model = read. Cb. Model([filename]); ≫ write. Cb. Model(model, format, [file. Name], [comp. Symbol. List], [comp. Name. List], [SBMLLevel], [SBMLVersion]); n Simulate optimal growth using flux-balance analysis (FBA) ≫ [solution] = optimize. Cb. Model(model, [osense. Str], [min. Norm], [allow. Loops]) n

Simulating deletion studies ≫[gr. Ratio, gr. Rate. KO, gr. Rate. WT, has. Effect, del. Rxns, flux. Solution] = single. Gene. Deletion(model, method, [gene. List]) n Flux Variability Analysis determine the minimum and maximum flux values that the reactions within the model can carry while obtaining a specific percentage of optimal growth rate ≫ [min. Flux max. Flux] = flux. Variability(model, opt. Percentage, [rxn. Name. List], [verb. Flag], [allow. Loops]) n

Metabolic engineering Opt. Knock—Opt. Knock runs the Opt. Knock algorithm to determine reaction sets to knock out for the overproduction of a specific product when the model is optimized for internal cellular objectives. ≫ [Opt. Knock. Sol, bi. Level. MILPproblem] = Opt. Knock(model, selected. Rxn. List, [options], [constr. Opt], [prev. Solutions], [verb. Flag]) n

n Extream Pathways and Elementary Flux Modes
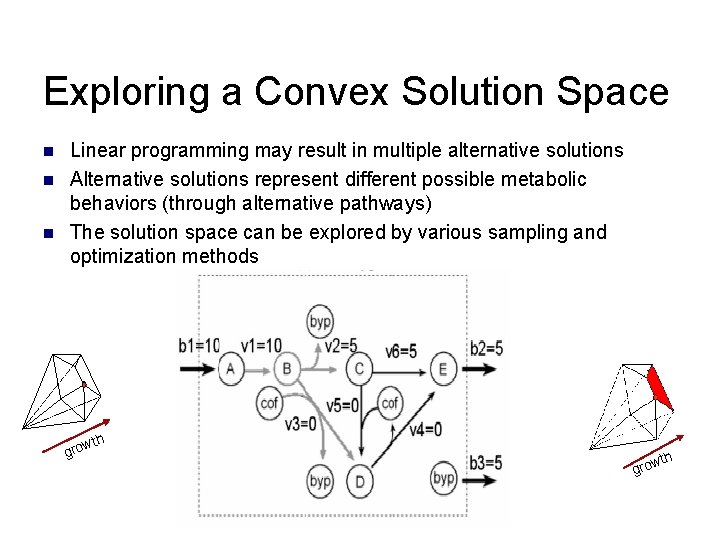
Exploring a Convex Solution Space n n n Linear programming may result in multiple alternative solutions Alternative solutions represent different possible metabolic behaviors (through alternative pathways) The solution space can be explored by various sampling and optimization methods wth gro h wt gro

Topological Methods n Not biased by a statement of an objective n Network based pathways: ¨ Extreme Pathways (Schilling, et. al. , 1999) ¨ Elementary Flux Modes (Schuster, el. al. , 1999) Decomposing flux distribution into extreme pathways Extreme pathways defining phenotypic phase planes Uniform random sampling n n n
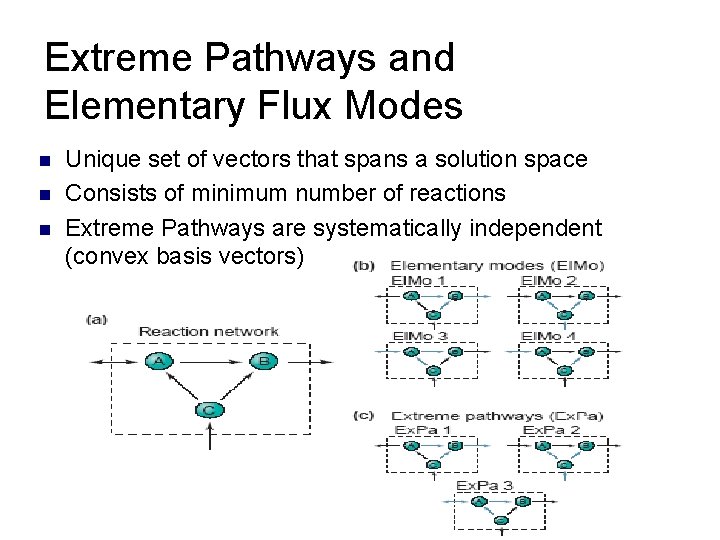
Extreme Pathways and Elementary Flux Modes n n n Unique set of vectors that spans a solution space Consists of minimum number of reactions Extreme Pathways are systematically independent (convex basis vectors)
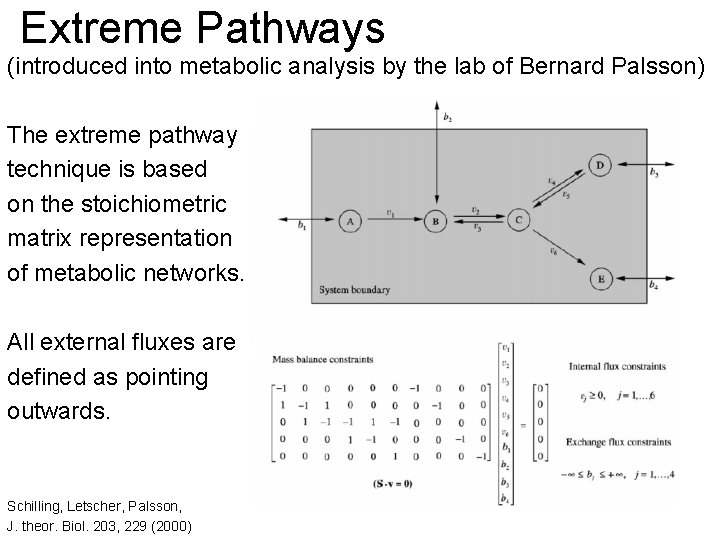
Extreme Pathways (introduced into metabolic analysis by the lab of Bernard Palsson) The extreme pathway technique is based on the stoichiometric matrix representation of metabolic networks. All external fluxes are defined as pointing outwards. Schilling, Letscher, Palsson, J. theor. Biol. 203, 229 (2000)
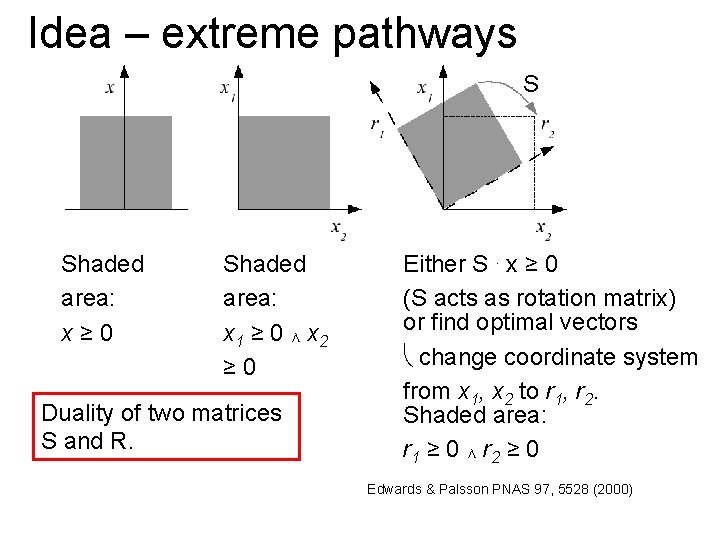
Idea – extreme pathways S Shaded area: x≥ 0 Shaded area: x 1 ≥ 0 ∧ x 2 ≥ 0 Duality of two matrices S and R. Either S. x ≥ 0 (S acts as rotation matrix) or find optimal vectors change coordinate system from x 1, x 2 to r 1, r 2. Shaded area: r 1 ≥ 0 ∧ r 2 ≥ 0 Edwards & Palsson PNAS 97, 5528 (2000)
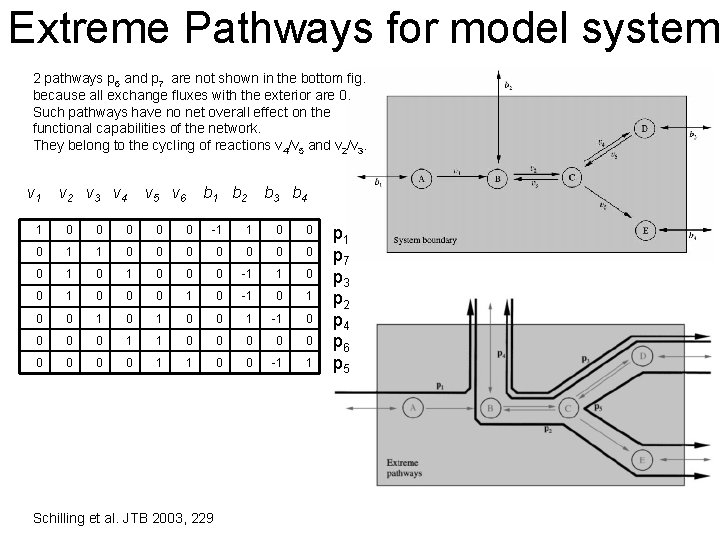
Extreme Pathways for model system 2 pathways p 6 and p 7 are not shown in the bottom fig. because all exchange fluxes with the exterior are 0. Such pathways have no net overall effect on the functional capabilities of the network. They belong to the cycling of reactions v 4/v 5 and v 2/v 3. v 1 v 2 v 3 v 4 v 5 v 6 b 1 b 2 b 3 b 4 1 0 0 0 -1 1 0 0 0 1 0 0 0 -1 1 0 0 0 1 0 -1 0 1 0 0 1 -1 0 0 1 1 0 0 0 0 0 1 1 0 0 -1 1 Schilling et al. JTB 2003, 229 p 1 p 7 p 3 p 2 p 4 p 6 p 5
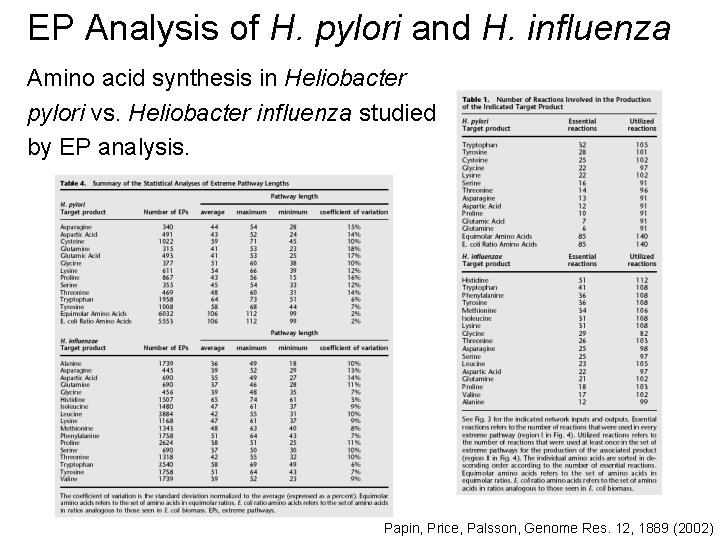
EP Analysis of H. pylori and H. influenza Amino acid synthesis in Heliobacter pylori vs. Heliobacter influenza studied by EP analysis. Papin, Price, Palsson, Genome Res. 12, 1889 (2002)

Summary – Extreme Pathways Extreme Pathway Analysis is a standard technique for analysis of metabolic networks. Number of EPs can become extremely large – hard to interpret. EP is an excellent basis for studying systematic effects of reaction cut sets. It will be very important to consider the interplay of metabolic and regulatory networks.

Assignment 1. 2. What is the principle of Flux Balance Analysis and describe the procedure to do FBA? Please read the following literatures, and list which software and tools are required to be installed to use COBRA for flux analysis. Becker et. al. (2007) Quantitative prediction of cellular metabolism with constraint-based models: the COBRA Toolbox. Nature Protocols 2: 727 -738. Schellenberger J, et al. (2011) Quantitative prediction of cellular metabolism with constraint-based models: the COBRA Toolbox v 2. 0 Nature Protocols 6(9): 1290 -307.
- Slides: 54