Connectivity Interest Group Meeting 1 Background for Interest

“Connectivity” Interest Group Meeting 1: Background for Interest Group Structural vs Functional vs Effective connectivity Basics of Dynamic Causal Modelling Rik Henson (Thanks to Andre Marrieros for some slides)

28 April: Alex Clarke (CSL) Synchronisation in the ventral stream supports complex semantic processing: evidence from MEG 12 May: James Rowe (CBU/CS) Advanced DCM: Bayesian model selection, random/fixed effects, model families 26 May: Michael Ewbank (CBU) Example application of above to f. MRI data on adaptation to visual images of bodies in EBA/FBA 9 June: Bernhard Staresina (CBU) Measures of power-power, power-phase, phase-phase, etc using intracranial EEG 23 June: Dan Wakeman (CBU) DCM for source-reconstructed extracranial MEG/EEG data 7 July: Elisabeth Von Dem Hagen (CBU) ICA analysis of resting state f. MRI data 21 July: John Griffiths (CSL) Connecting structural and dynamic connectivity: conduction delays, fibre geometry, and tissue microstructure • Stop for Summer (August)? • Additional topics/speakers?
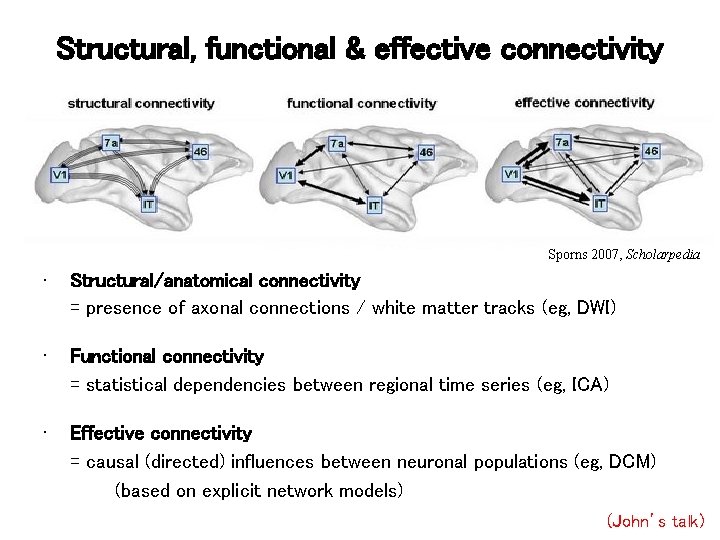
Structural, functional & effective connectivity Sporns 2007, Scholarpedia • Structural/anatomical connectivity = presence of axonal connections / white matter tracks (eg, DWI) • Functional connectivity = statistical dependencies between regional time series (eg, ICA) • Effective connectivity = causal (directed) influences between neuronal populations (eg, DCM) (based on explicit network models) (John’s talk)
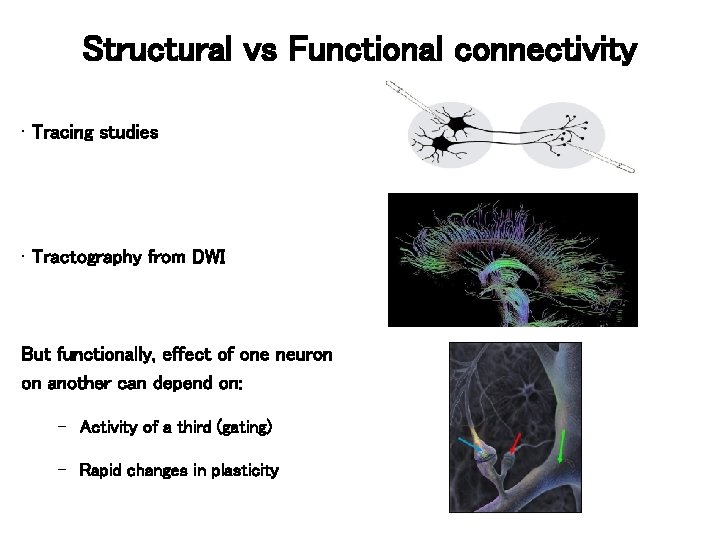
Structural vs Functional connectivity • Tracing studies • Tractography from DWI But functionally, effect of one neuron on another can depend on: – Activity of a third (gating) – Rapid changes in plasticity
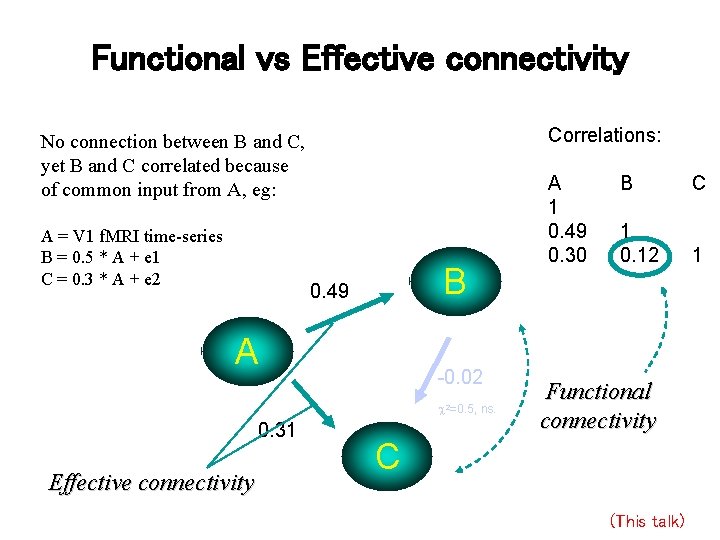
Functional vs Effective connectivity Correlations: No connection between B and C, yet B and C correlated because of common input from A, eg: A = V 1 f. MRI time-series B = 0. 5 * A + e 1 C = 0. 3 * A + e 2 B 0. 49 A -0. 02 2=0. 5, 0. 31 Effective connectivity A 1 0. 49 0. 30 ns. B C 1 0. 12 1 Functional connectivity C (This talk)

Functional connectivity • Useful when no model, no experimental perturbation (eg resting state) • Popular examples: seed-voxel correlations, PCA, ICA, etc • With frequency decomposition (eg EEG/MEG): coherence, phase-locking, (Bernhard’s talk) nonlinear synchrony, etc • Graph-theory summaries of functional networks • Correlations in f. MRI timeseries could be spurious haemodynamics (e. g, effects of heart-rate/breathing on vascular network) • Condition-dependent changes in functional connectivity (e. g, changes in seed voxel regression slope; “psycho-physiological interactions“ (PPIs)) (Elizabeth’s talk) (Ed Bullmore? )

Effective-connectivity: Definitions of Causality? 1. Temporal precedence (e. g, Granger Causality, DCM) 2. Network model inference (e. g, SEM, MAR, DCM) 3. Indirect experimental manipulations (e. g, PPI, DCM) 4. Direct experimental interventions (e. g, lesion, drugs) 5. …
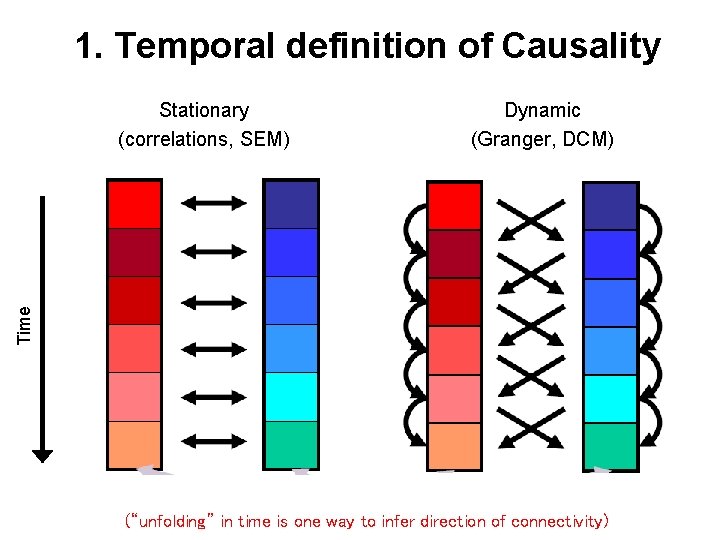
1. Temporal definition of Causality Dynamic (Granger, DCM) Time Stationary (correlations, SEM) (“unfolding” in time is one way to infer direction of connectivity)

1. Note on temporal causality and f. MRI • Problem with time-based measures of connectivity arises with f. MRI: BOLD timeseries is not direct reflection of Neural timeseries – (e. g, peak BOLD response in motor cortex can precede that in visual cortex in a visually-cued motor task, owing to different neural-BOLD mappings) • This compromises methods like Granger Causality and Multivariate Auto. Regressive models (MAR) (and PPIs) that operate directly on f. MRI data (Friston, 2010; Smith et al, 2011) • Note that this does not preclude these methods (eg MAR) for MEG/EEG timeseries, assuming these are more direct measures of neural activity

Effective-connectivity: Definitions of Causality? 1. Temporal precedence (e. g, Granger Causality, DCM) 2. Network model inference (e. g, SEM, MAR, DCM) 3. Indirect experimental manipulations (e. g, PPI, DCM) 4. Direct experimental interventions (e. g, lesion, drugs) 5. …

2. Explicit Network Models of Causality • (Bivariate) correlations such as Granger Causality (though see Partial Directed Coherence? ) do not use an explicit network (graph) model • Structural Equation Modelling (SEM) can test different network models, by simply comparing predicted with observed covariance matrices, but. . . – has no dynamical model (stationary covariances) – has no neural-BOLD model – cannot test some graphs, eg loops (no temporal definition of direction) – restricted to classical inference comparing nested models

Effective-connectivity: Definitions of Causality? 1. Temporal precedence (e. g, Granger Causality, DCM) 2. Network model inference (e. g, SEM, MAR, PDC, DCM) 3. Indirect experimental manipulations (e. g, PPI, DCM) 4. Direct experimental interventions (e. g, lesion, drugs) 5. …

3. Experimental Manipulation and Causality • Even dynamic (time-lagged), network (directed graph) models (eg PDC) may give little insight into theoretically-relevant causality – e. g, driving of Region B by A, but not C by A, could simply reflect stronger structural connectivity from A to B than from A to C • Want to be able to model dynamic changes in connectivity as a function of experimentally-controlled perturbations over time (eg, PPI, DCM) – e. g, driving of Region B by A depends on attention to a specific stimulus attribute (by controlling periods of attention to that attribute vs another)

=> Development of DCM 1. Dynamic: based on first-order differential equations - at level of neural activity, with separate haemodynamic model for f. MRI 2. Causal: based on explicit directed graph models 3. Modelling: designed to test experimental manipulations - “bilinear” approximation to interactive dynamics 4. (Estimated in a Bayesian context, allowing formal comparison of any number/type of models…)
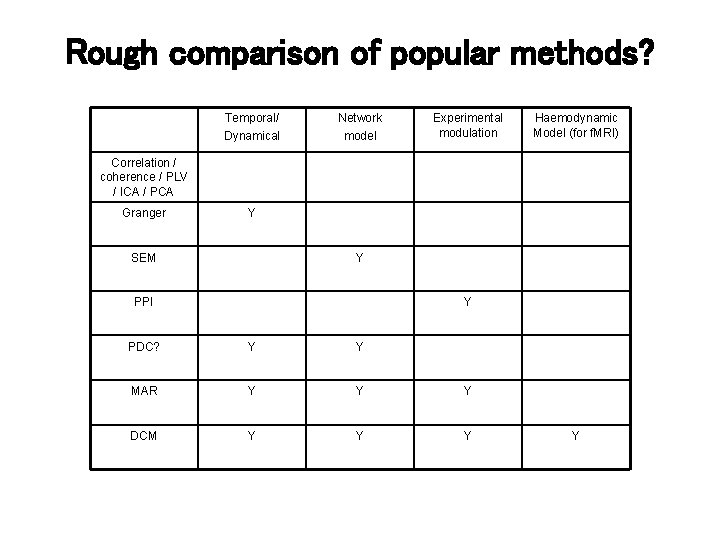
Rough comparison of popular methods? Temporal/ Dynamical Network model Experimental modulation Haemodynamic Model (for f. MRI) Correlation / coherence / PLV / ICA / PCA Granger Y SEM Y PPI Y PDC? Y Y MAR Y Y Y DCM Y Y

DCM for f. MRI. . .
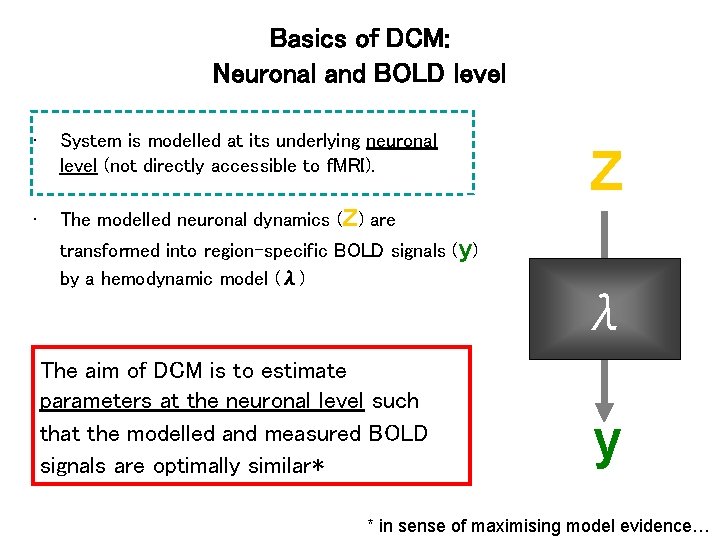
Basics of DCM: Neuronal and BOLD level • System is modelled at its underlying neuronal level (not directly accessible to f. MRI). • The modelled neuronal dynamics (Z) are Z transformed into region-specific BOLD signals (y) by a hemodynamic model (λ) λ The aim of DCM is to estimate parameters at the neuronal level such that the modelled and measured BOLD signals are optimally similar* y * in sense of maximising model evidence…
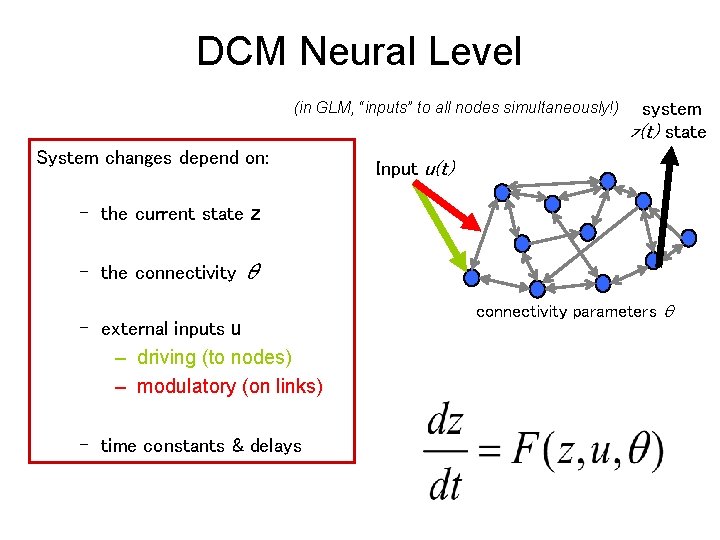
DCM Neural Level (in GLM, “inputs” to all nodes simultaneously!) System changes depend on: system z(t) state Input u(t) – the current state z – the connectivity θ – external inputs u – driving (to nodes) – modulatory (on links) – time constants & delays connectivity parameters
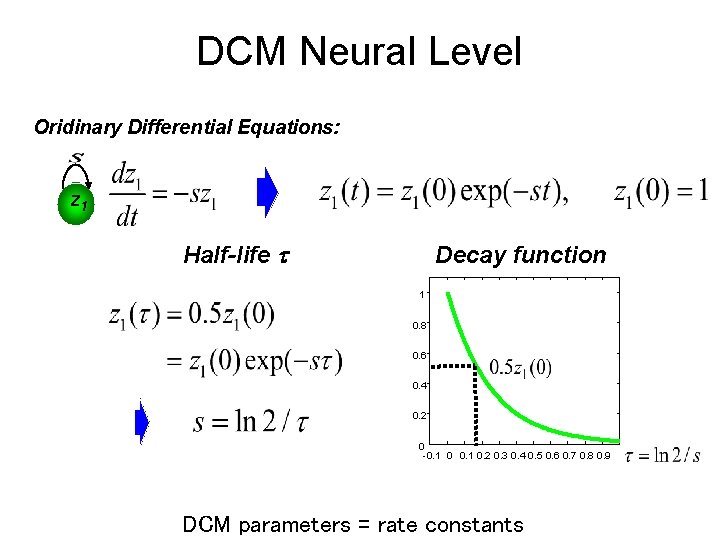
DCM Neural Level Oridinary Differential Equations: z 1 Half-life Decay function 1 0. 8 0. 6 0. 4 0. 2 0 -0. 1 0. 2 0. 3 0. 4 0. 5 0. 6 0. 7 0. 8 0. 9 DCM parameters = rate constants
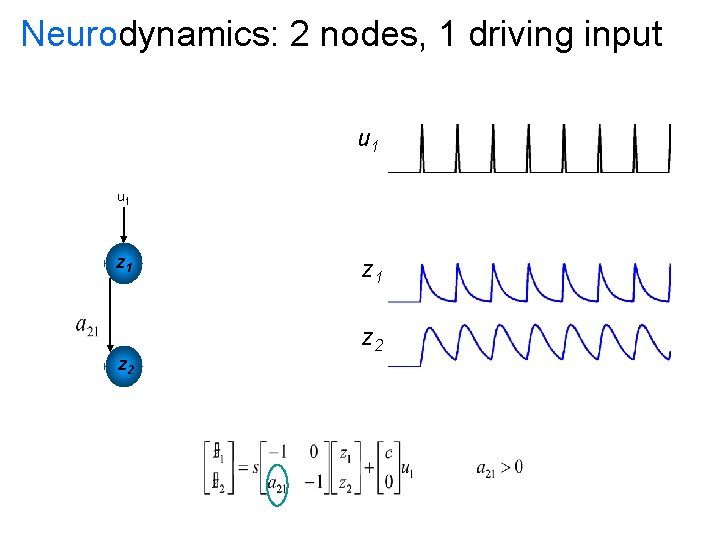
Neurodynamics: 2 nodes, 1 driving input u 1 u 2 z 1 z 2
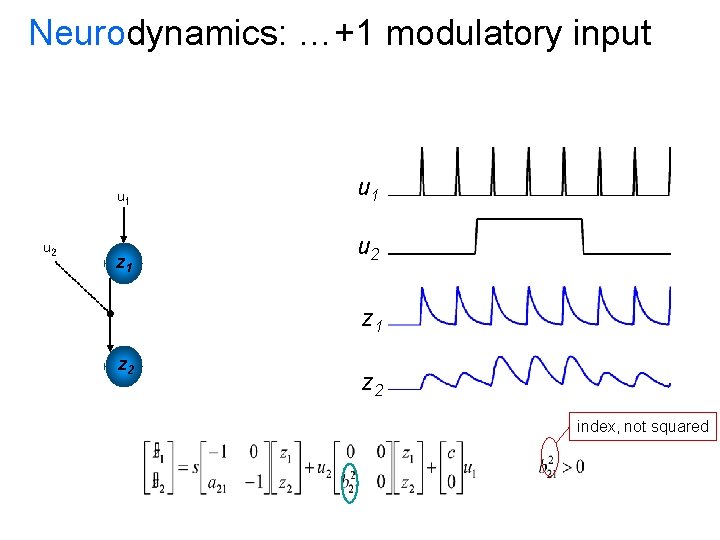
Neurodynamics: …+1 modulatory input u 1 u 2 z 1 z 2 index, not squared
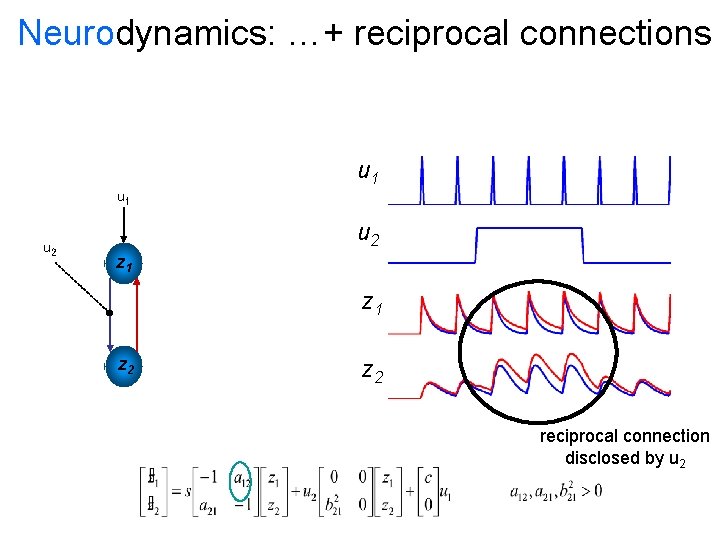
Neurodynamics: …+ reciprocal connections u 1 u 2 z 1 z 2 reciprocal connection disclosed by u 2
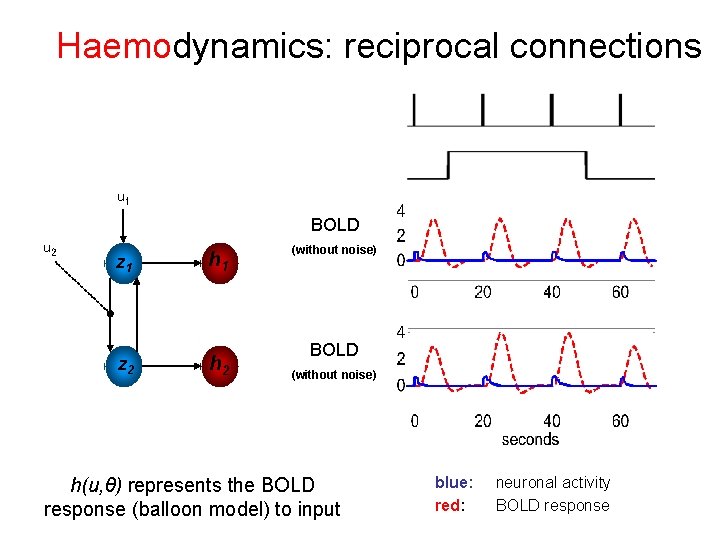
Haemodynamics: reciprocal connections u 1 BOLD u 2 z 1 z 2 h 1 h 2 (without noise) BOLD (without noise) h(u, θ) represents the BOLD response (balloon model) to input blue: red: neuronal activity BOLD response
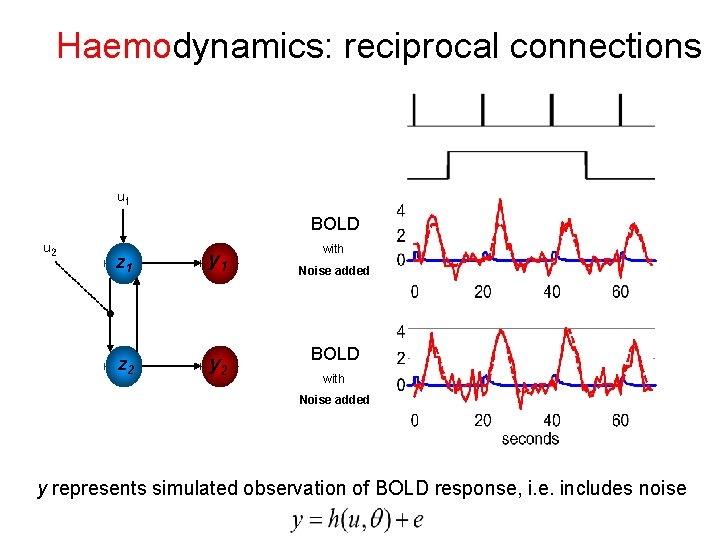
Haemodynamics: reciprocal connections u 1 BOLD u 2 z 1 y 1 z 2 y 2 with Noise added BOLD with Noise added y represents simulated observation of BOLD response, i. e. includes noise
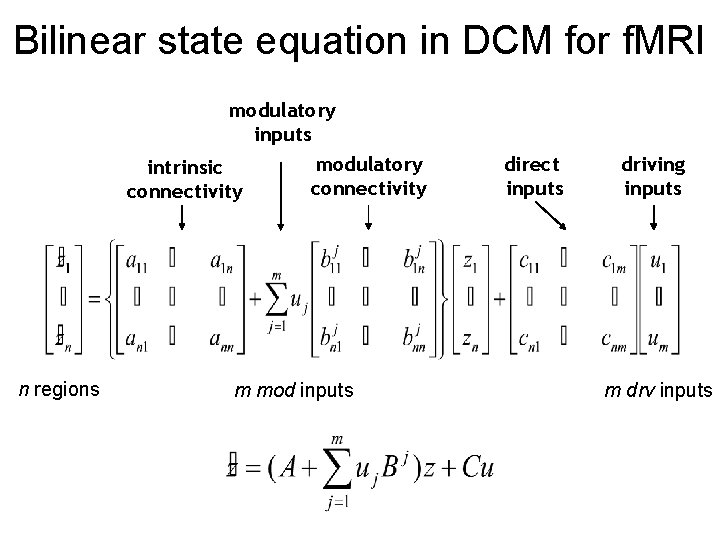
Bilinear state equation in DCM for f. MRI modulatory inputs modulatory intrinsic connectivity n regions m mod inputs direct inputs driving inputs m drv inputs
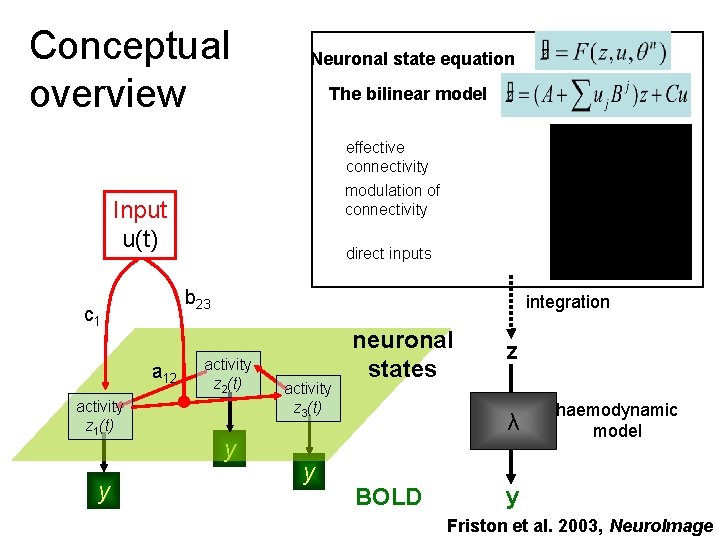
Conceptual overview Neuronal state equation The bilinear model effective connectivity modulation of connectivity Input u(t) b 23 c 1 a 12 activity z 1(t) y direct inputs integration activity z 2(t) y activity z 3(t) y neuronal states z λ BOLD haemodynamic model y Friston et al. 2003, Neuro. Image
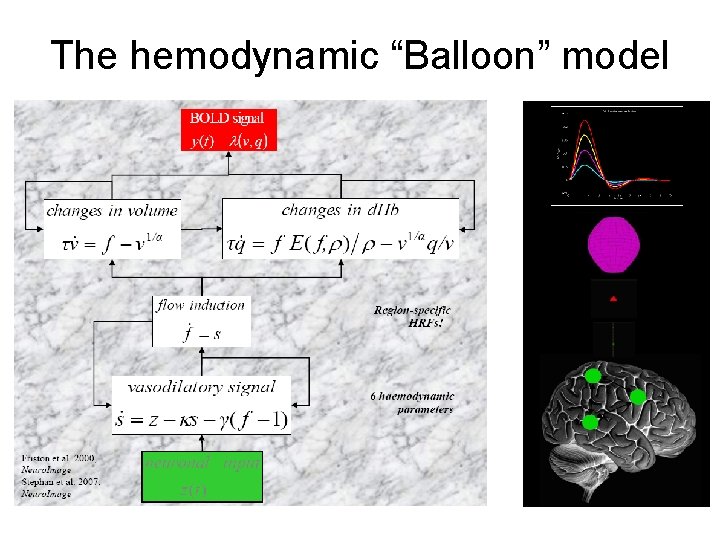
The hemodynamic “Balloon” model
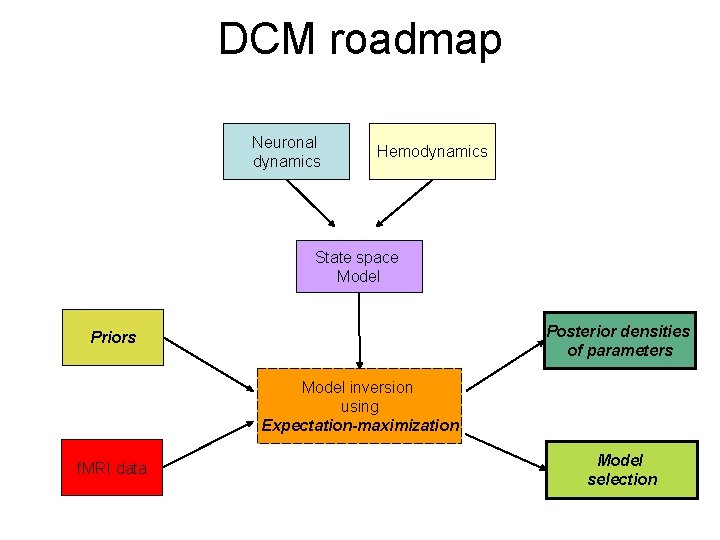
DCM roadmap Neuronal dynamics Hemodynamics State space Model Posterior densities of parameters Priors Model inversion using Expectation-maximization f. MRI data Model selection
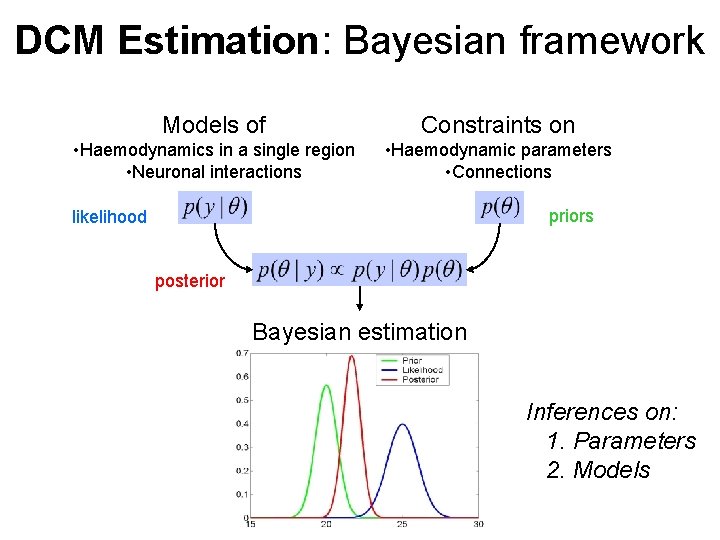
DCM Estimation: Bayesian framework Models of Constraints on • Haemodynamics in a single region • Neuronal interactions • Haemodynamic parameters • Connections priors likelihood posterior Bayesian estimation Inferences on: 1. Parameters 2. Models
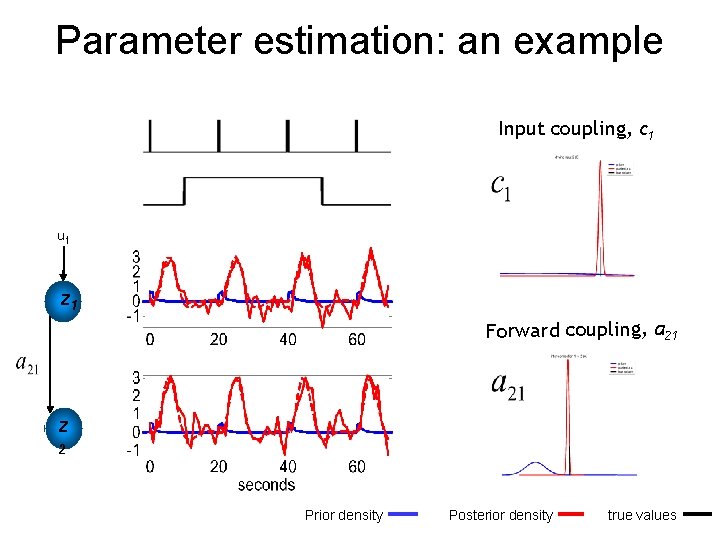
Parameter estimation: an example Input coupling, c 1 u 1 z 1 Forward coupling, a 21 z 2 Prior density Posterior density true values
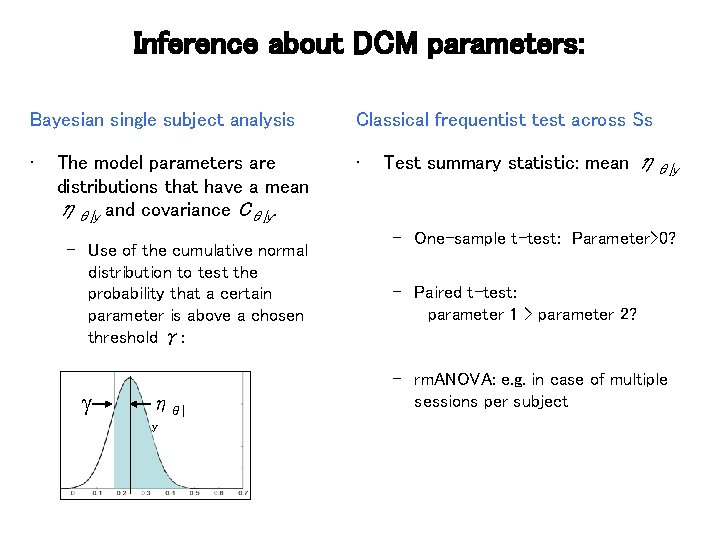
Inference about DCM parameters: Bayesian single subject analysis Classical frequentist test across Ss • • The model parameters are distributions that have a mean ηθ|y and covariance Cθ|y. – Use of the cumulative normal distribution to test the probability that a certain parameter is above a chosen threshold γ: ηθ| y Test summary statistic: mean ηθ|y – One-sample t-test: Parameter>0? – Paired t-test: parameter 1 > parameter 2? – rm. ANOVA: e. g. in case of multiple sessions per subject
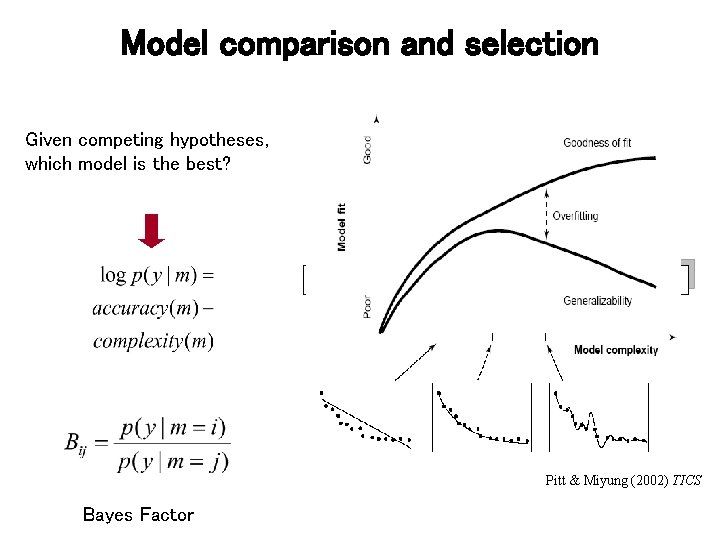
Model comparison and selection Given competing hypotheses, which model is the best? Pitt & Miyung (2002) TICS Bayes Factor
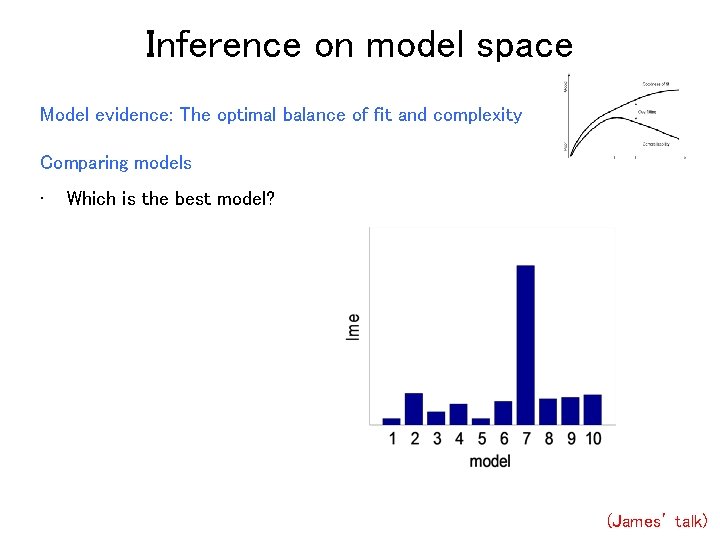
Inference on model space Model evidence: The optimal balance of fit and complexity Comparing models • Which is the best model? (James’ talk)
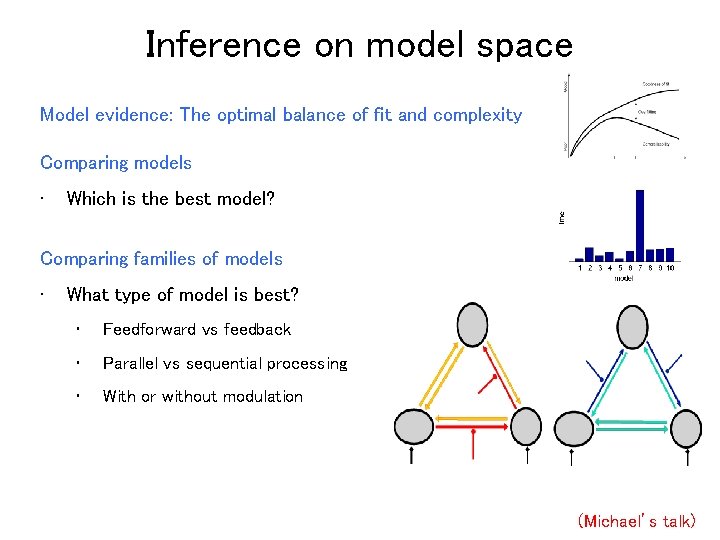
Inference on model space Model evidence: The optimal balance of fit and complexity Comparing models • Which is the best model? Comparing families of models • What type of model is best? • Feedforward vs feedback • Parallel vs sequential processing • With or without modulation (Michael’s talk)
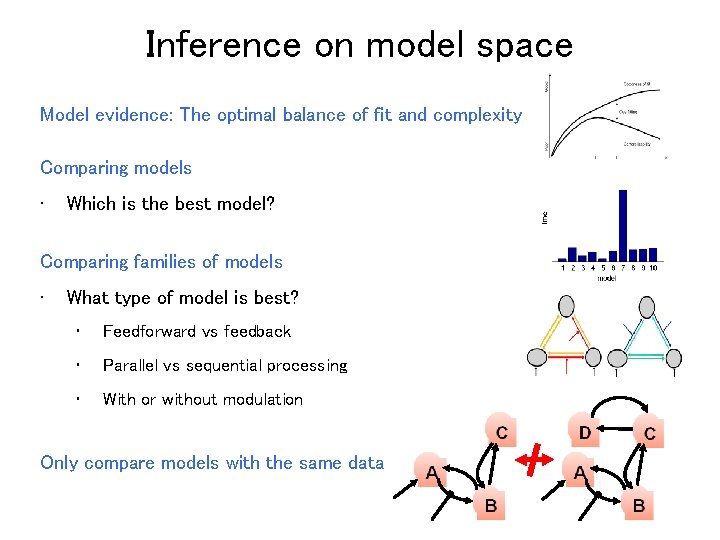
Inference on model space Model evidence: The optimal balance of fit and complexity Comparing models • Which is the best model? Comparing families of models • What type of model is best? • Feedforward vs feedback • Parallel vs sequential processing • With or without modulation Only compare models with the same data
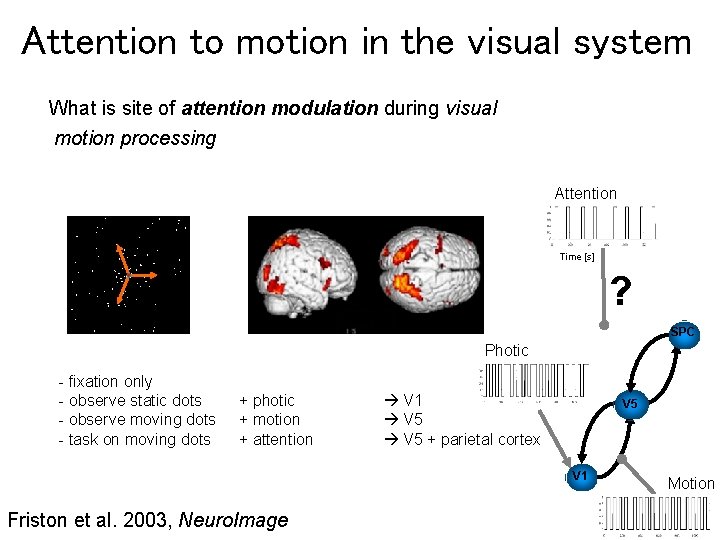
Attention to motion in the visual system What is site of attention modulation during visual motion processing Attention Time [s] ? SPC Photic - fixation only - observe static dots - observe moving dots - task on moving dots + photic + motion + attention V 1 V 5 + parietal cortex V 5 V 1 Friston et al. 2003, Neuro. Image Motion
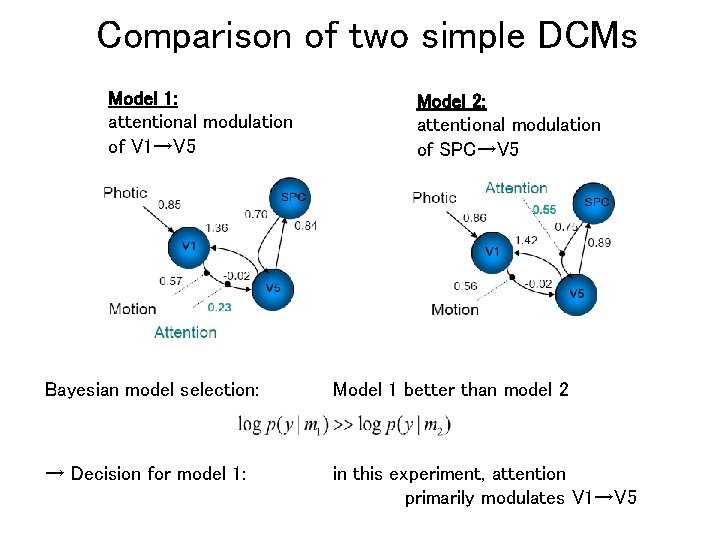
Comparison of two simple DCMs Model 1: attentional modulation of V 1→V 5 Model 2: attentional modulation of SPC→V 5 Bayesian model selection: Model 1 better than model 2 → Decision for model 1: in this experiment, attention primarily modulates V 1→V 5

So, DCM…. • enables one to infer hidden neuronal processes • allows one to test mechanistic hypotheses about observed effects – uses a deterministic differential equation to model neuro-dynamics (represented by matrices A, B and C) • is informed by anatomical and physiological principles • uses a Bayesian framework to estimate model parameters • is a generic approach to modelling experimentally perturbed dynamic systems. – provides an observation model for neuroimaging data, e. g. f. MRI, M/EEG – DCM is not model or modality specific (Models will change and the method extended to other modalities e. g. ERPs, LFPs)

Variants of DCM • DCM for f. MRI – “non-linear” DCM: modulatory input (B) equal to activity in another region – “two-state” DCM: inhibitory and excitatory neuronal subpopulations – “stochastic” DCM: random element to activity (e. g, for resting state) • DCM for E/MEG – “evoked” responses (complex neuronal model based on physiology) – “induced” responses (within/across frequency power coupling; no physiological model (more like DCM for f. MRI)) – “steady-state” responses – with (e. g, EEG/MEG) or without (e. g, LFP, i. EEG) a forward (head) model

Some useful references • The first DCM paper: Dynamic Causal Modelling (2003). Friston et al. Neuro. Image 19: 1273 -1302. • Physiological validation of DCM for f. MRI: Identifying neural drivers with functional MRI: an electrophysiological validation (2008). David et al. PLo. S Biol. 6 2683– 2697 • Hemodynamic model: Comparing hemodynamic models with DCM (2007). Stephan et al. Neuro. Image 38: 387 -401 • Nonlinear DCMs: Nonlinear Dynamic Causal Models for FMRI (2008). Stephan et al. Neuro. Image 42: 649 -662 • Two-state model: Dynamic causal modelling for f. MRI: A two-state model (2008). Marreiros et al. Neuro. Image 39: 269 -278 • Group Bayesian model comparison: Bayesian model selection for group studies (2009). Stephan et al. Neuro. Image 46: 1004 -10174 • 10 Simple Rules for DCM (2010). Stephan et al. Neuro. Image 52.

Thank you for your attention!!!
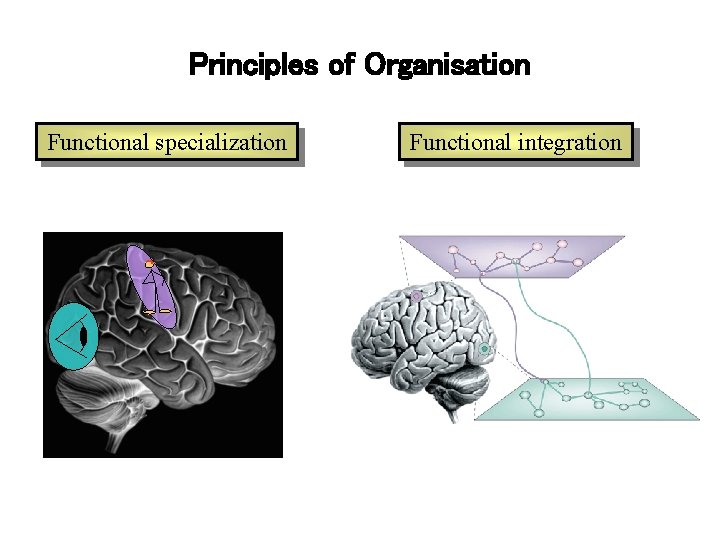
Principles of Organisation Functional specialization Functional integration
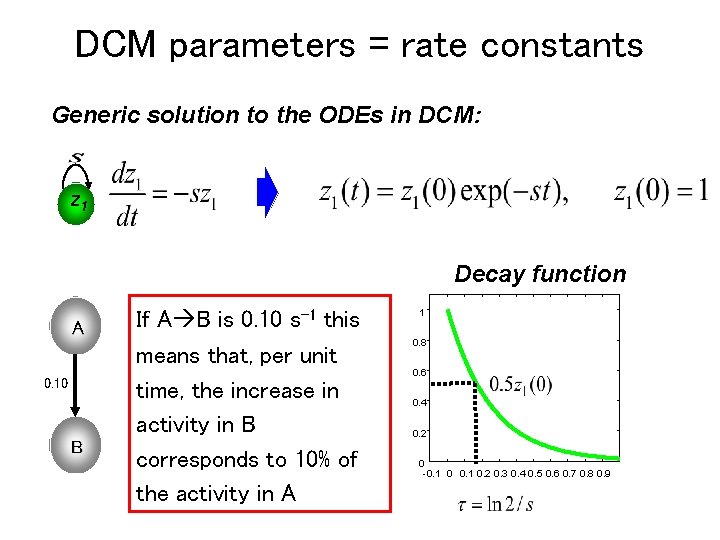
DCM parameters = rate constants Generic solution to the ODEs in DCM: z 1 Decay function A 0. 10 B If A B is 0. 10 s-1 this means that, per unit time, the increase in activity in B corresponds to 10% of the activity in A 1 0. 8 0. 6 0. 4 0. 2 0 -0. 1 0. 2 0. 3 0. 4 0. 5 0. 6 0. 7 0. 8 0. 9
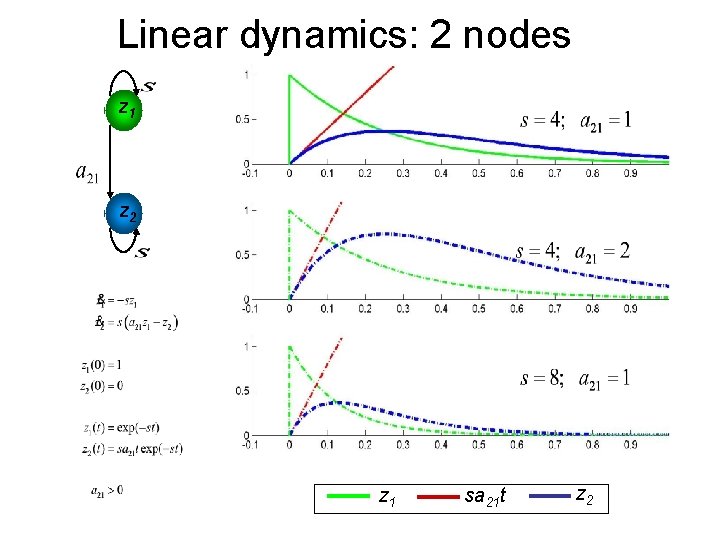
Linear dynamics: 2 nodes z 1 z 2 z 1 sa 21 t z 2
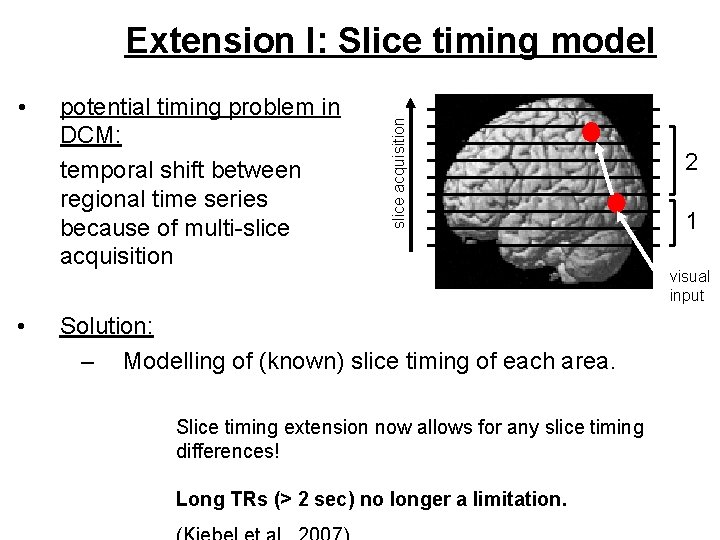
• • potential timing problem in DCM: temporal shift between regional time series because of multi-slice acquisition Extension I: Slice timing model Solution: – Modelling of (known) slice timing of each area. Slice timing extension now allows for any slice timing differences! Long TRs (> 2 sec) no longer a limitation. 2 1 visual input
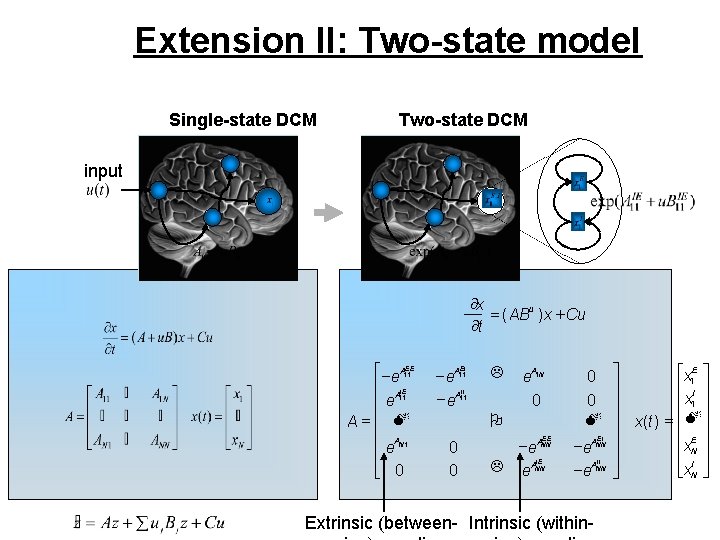
Extension II: Two-state model Single-state DCM Two-state DCM input ¶x = ( ABu ) x + Cu ¶t é -e. A 11 EE ê AIE ê e 11 A=ê M ê A ê e N 1 ê ë 0 EI -e. A 11 II -e. A 11 L A e 1 N 0 O 0 0 EE -e. ANN L AIE e NN 0 0 M EI -e. ANN II -e. ANN Extrinsic (between- Intrinsic (within- ù éx 1 E ù ú ê I ú ú êx 1 ú ú x( t ) = ê Mú ú ê Eú ú êx. N ú ú êx. I ú ëNû û
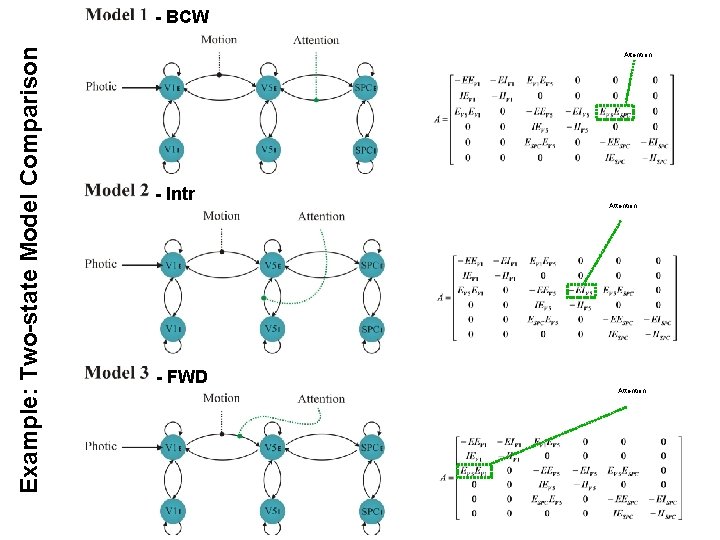
Example: Two-state Model Comparison - BCW b Attention - Intr - FWD Attention
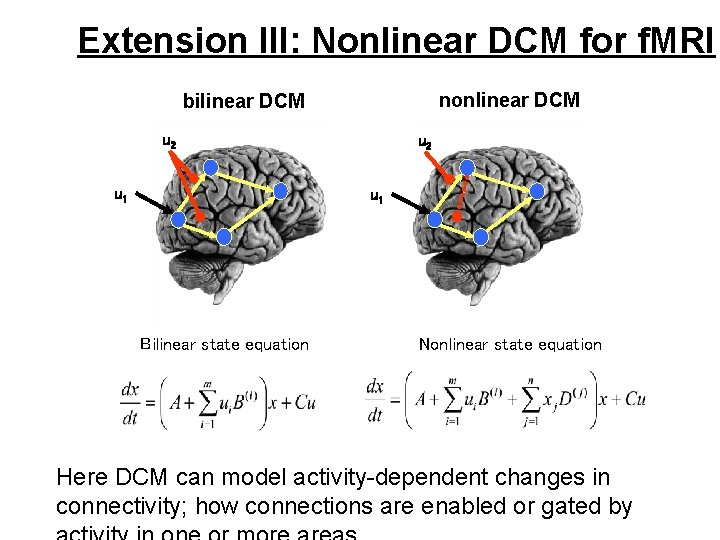
Extension III: Nonlinear DCM for f. MRI nonlinear DCM bilinear DCM u 2 u 1 Bilinear state equation Nonlinear state equation Here DCM can model activity-dependent changes in connectivity; how connections are enabled or gated by

Extension III: Nonlinear DCM for f. MRI Can V 5 activity during attention to motion be explained by allowing activity in SPC to modulate the V 1 -to-V 5 connection? attention . 0. 19 (100%) SPC 0. 03 (100%) 1. 65 visual (100%) stimulation 0. 01 (97. 4%) V 1 V 5 0. 04 (100%) motion The posterior density of indicates that this gating existed with 97% confidence. (The D matrix encodes which of the n neural units gate which connections in the system)
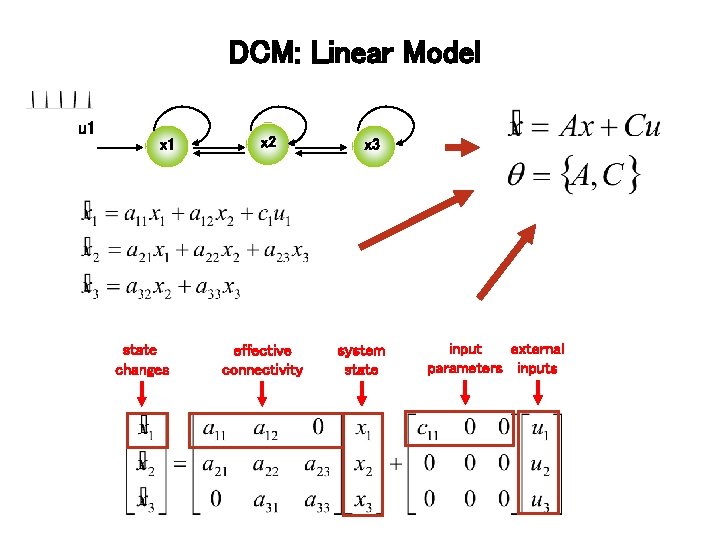
DCM: Linear Model u 1 x 1 state changes x 2 effective connectivity x 3 system state input external parameters inputs
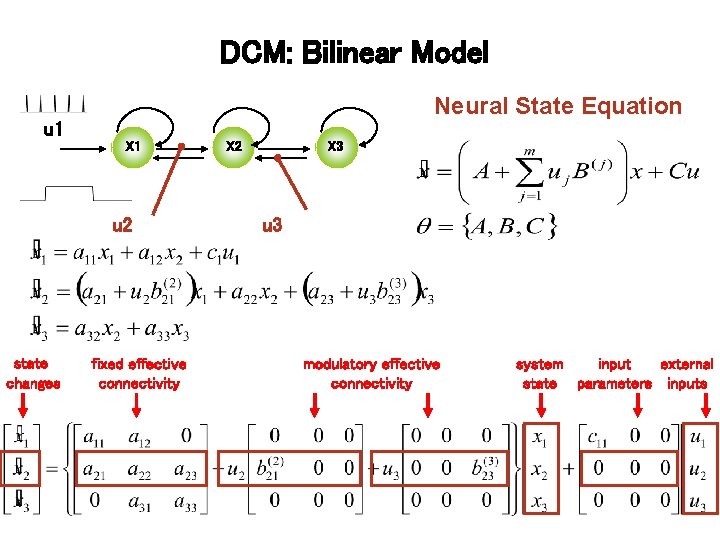
DCM: Bilinear Model Neural State Equation u 1 X 1 u 2 state changes fixed effective connectivity X 2 X 3 u 3 modulatory effective connectivity system input external state parameters inputs

Planning a DCM-compatible study • Suitable experimental design: – any design that is suitable for a GLM – preferably multi-factorial (e. g. 2 x 2) • e. g. one factor that varies the driving (sensory) input • and one factor that varies the contextual input • Hypothesis and model: – Define specific a priori hypothesis – Which parameters are relevant to test this hypothesis? – If you want to verify that intended model is suitable to test this hypothesis, then use simulations – Define criteria for inference – What are the alternative models to test?

Overview: parameter estimation • Specify model (neuronal and haemodynamic level) • Make it an observation model by adding measurement error e and confounds X (e. g. drift). stimulus function u neuronal state equation parameters hidden states state equation • Bayesian parameter estimation using expectation-maximization. • Result: (Normal) posterior parameter distributions, given by mean ηθ|y and Covariance Cθ|y. ηθ|y modeled BOLD response observation model
- Slides: 53