Connecting Function Process and Component Outline Chris Use














![xrefs: not necessarily equivalent glycolysis is_a some_has_part GO: new equivalent glycolysis [human] some_has_part glucose-6 xrefs: not necessarily equivalent glycolysis is_a some_has_part GO: new equivalent glycolysis [human] some_has_part glucose-6](https://slidetodoc.com/presentation_image_h2/a382c122be2a1544603165d58a150c1f/image-32.jpg)
![xrefs: not necessarily equivalent glycolysis xref some_has_part glycolysis [human] some_has_part glucose-6 phosphate isomerase activity xrefs: not necessarily equivalent glycolysis xref some_has_part glycolysis [human] some_has_part glucose-6 phosphate isomerase activity](https://slidetodoc.com/presentation_image_h2/a382c122be2a1544603165d58a150c1f/image-33.jpg)




![Conclusions [so far] • Implement Low-Hanging-High-Yield-Fruit ASAP – MF part_of BP – regulates • Conclusions [so far] • Implement Low-Hanging-High-Yield-Fruit ASAP – MF part_of BP – regulates •](https://slidetodoc.com/presentation_image_h2/a382c122be2a1544603165d58a150c1f/image-43.jpg)


































- Slides: 90

Connecting Function, Process and Component

Outline • Chris: – Use cases: why we need to do this – Specific details • The Low hanging Fruit • Pathways and the has_part relation – Mining links from Reactome and Metacyc • Jen/Harold: – Biological examples

What this talk doesn’t contain • no philosophy talk – no pontificating on the nature of functions

Why bother? • To improve the ontology • To fill in annotation gaps • As an aid to annotation – Suggest new annotations – Avoid redundant annotation effort – Annotation cross-products • Better integration with pathway databases • To present annotations to users in more useful ways – e. g. more informative Ami. GO displays

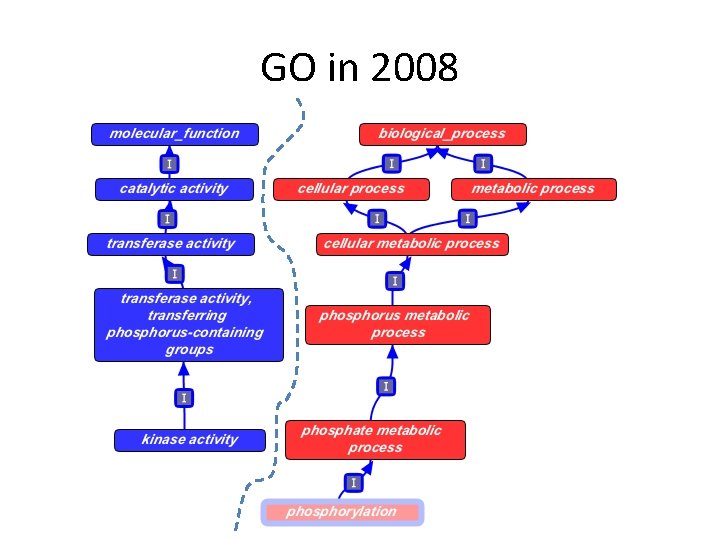
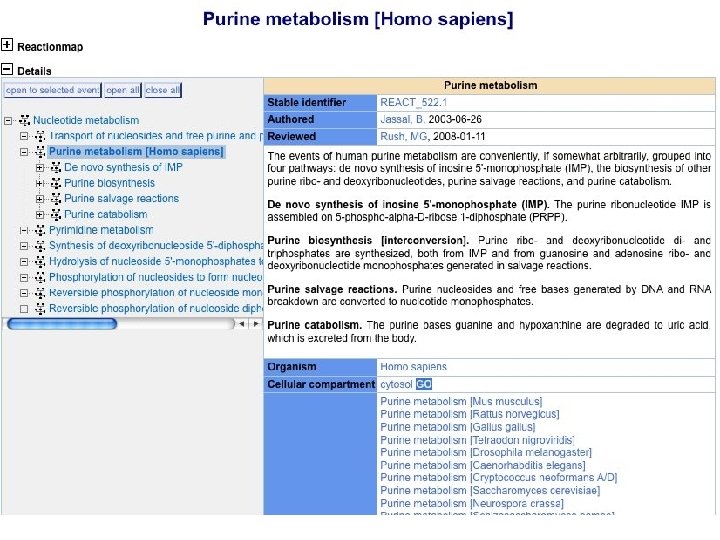
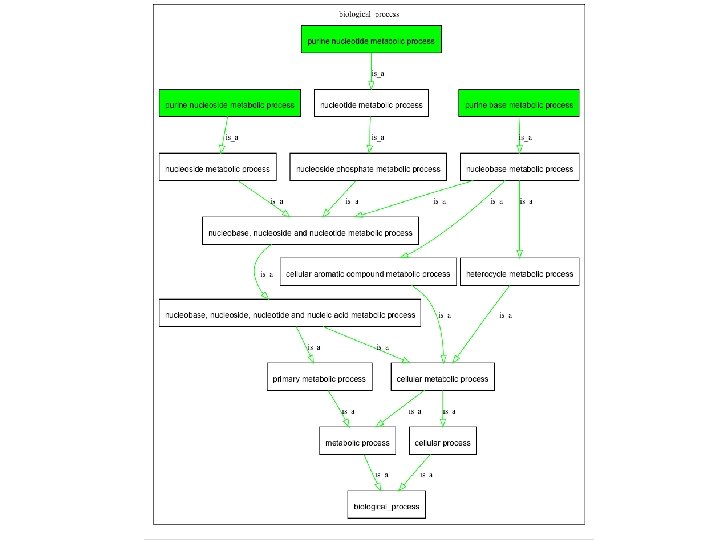
GO in 2008

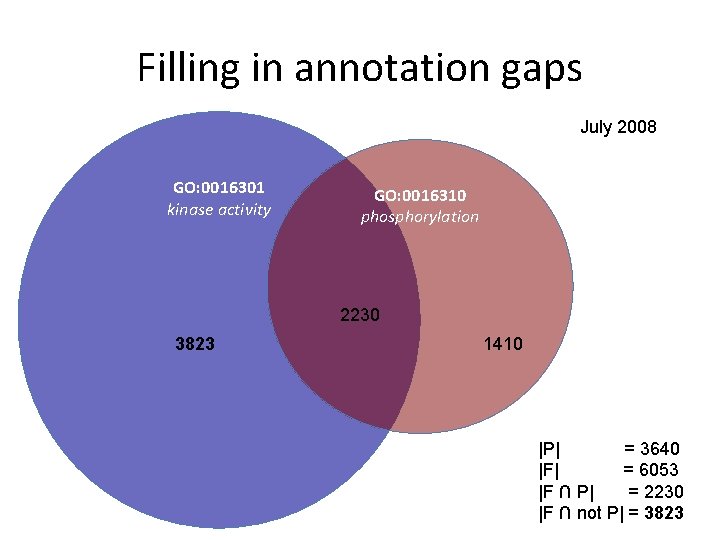
Filling in annotation gaps July 2008 GO: 0016301 kinase activity GO: 0016310 phosphorylation 2230 3823 1410 |P| = 3640 |F| = 6053 |F ∩ P| = 2230 |F ∩ not P| = 3823

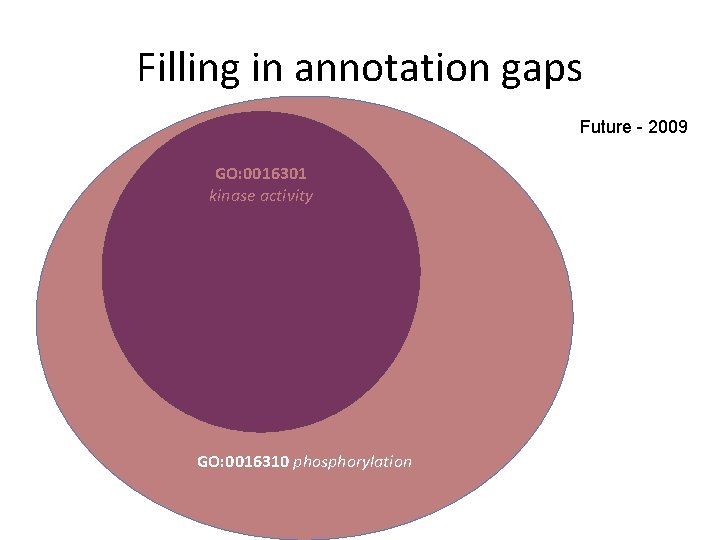
Filling in annotation gaps Future - 2009 GO: 0016301 kinase activity GO: 0016310 phosphorylation

Aid to annotation • GO: 0006096 ! glycolysis • • • GO: 0003872 ! 6 -phosphofructokinase activity GO: 0004332 ! fructose-bisphosphate aldolase activity GO: 0004347 ! glucose-6 -phosphate isomerase activity GO: 0004365 ! glyceraldehyde-3 -phosphate dehydrogenase (phosphorylating) activity GO: 0004618 ! phosphoglycerate kinase activity GO: 0004619 ! phosphoglycerate mutase activity GO: 0004634 ! phosphopyruvate hydratase activity GO: 0004743 ! pyruvate kinase activity GO: 0004807 ! triose-phosphate isomerase activity

Improved presentation to users

Specifics • Low Hanging Fruit – Function to process links • Mostly part_of links • Some regulates links • Pathways – Process to function • has_part – Mining from pathways databases & curation

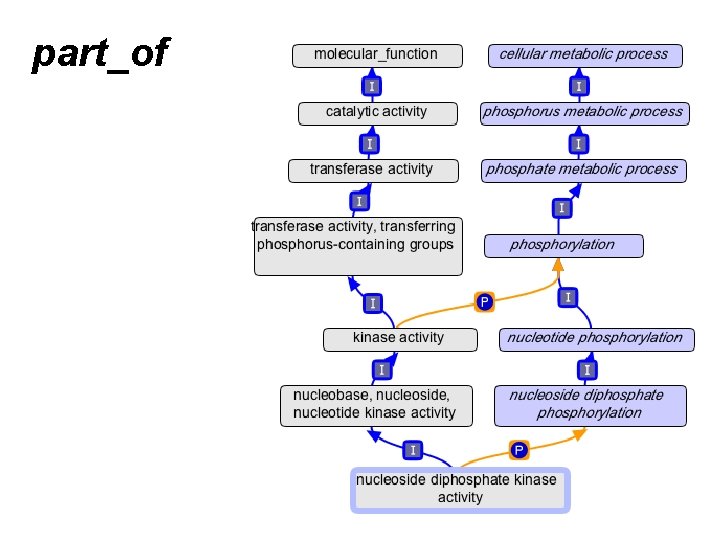
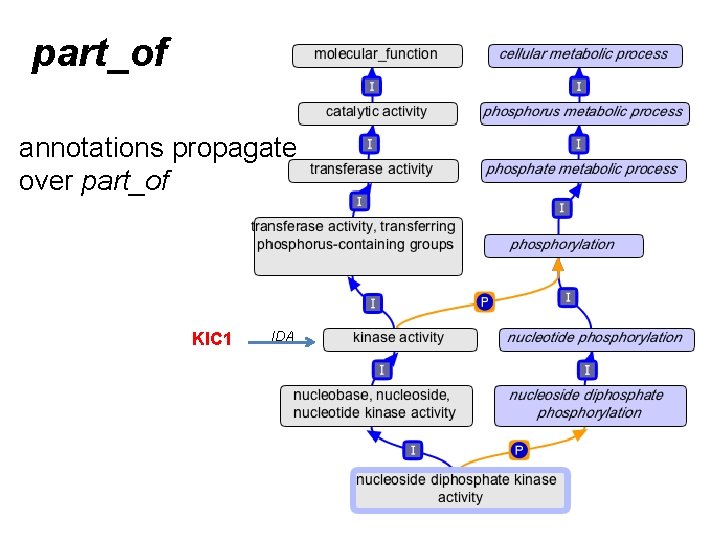
part_of

part_of annotations propagate over part_of KIC 1 IDA

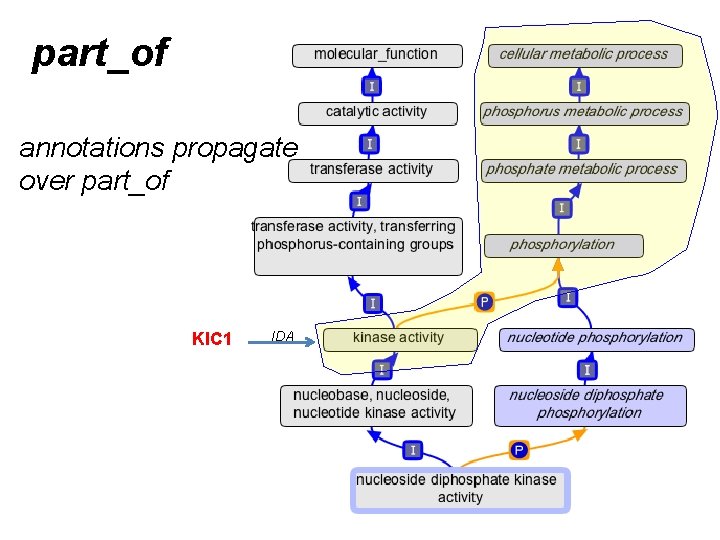
part_of annotations propagate over part_of KIC 1 IDA

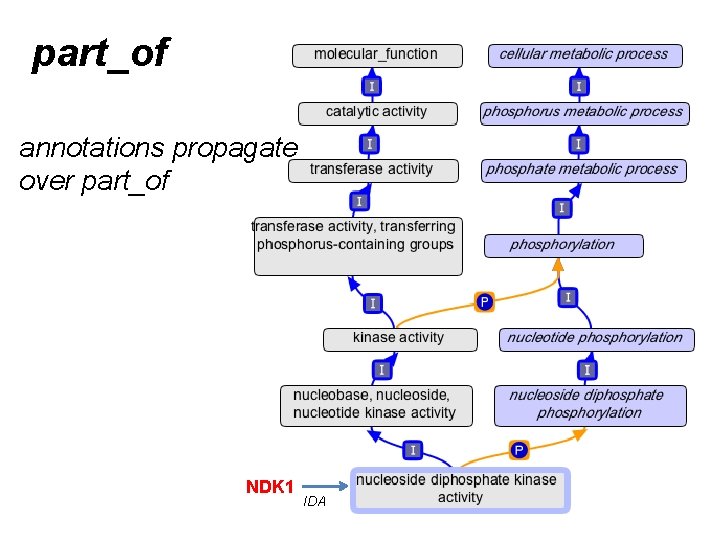
part_of annotations propagate over part_of NDK 1 IDA

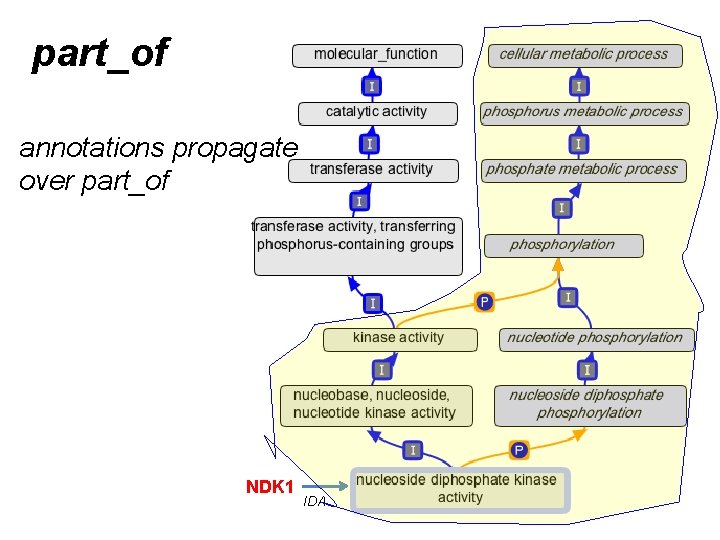
part_of annotations propagate over part_of NDK 1 IDA

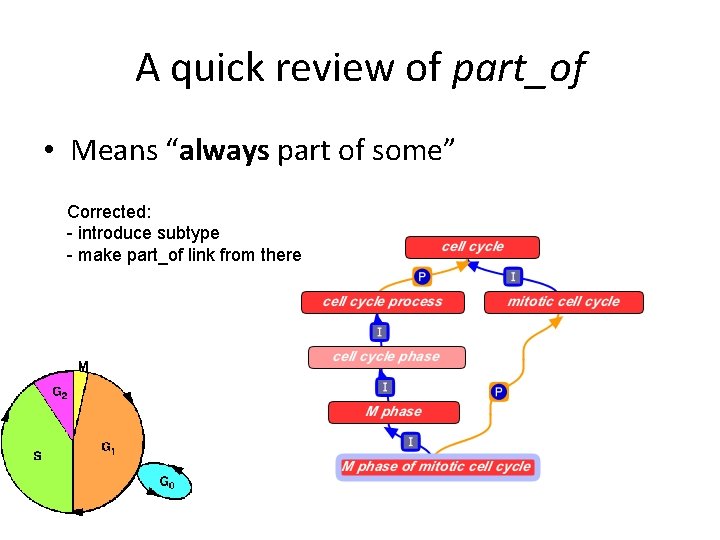
A quick review of part_of • Means “always part of some” – Example: • nucleus part_of cell • EVERY nucleus is part_of SOME cell

A quick review of part_of • Means “always part of some” – Counter example: • M phase part_of mitotic cell cycle WRONG • EVERY M phase is part_of SOME mitotic cell cycle

A quick review of part_of • Means “always part of some” Corrected: - introduce subtype - make part_of link from there

Guide to using part_of for MF • We make a link between MF and BP when – EVERY instance of the activity is executed in the context of that BP • Example: – kinase activity part_of phosphorylation • Counter-example: – 6 -phosphofructokinase activity part_of glycolysis • gene product annotations always propagate over part_of – from the MF, to the BP • NOT the reverse – and also over is_a, as usual

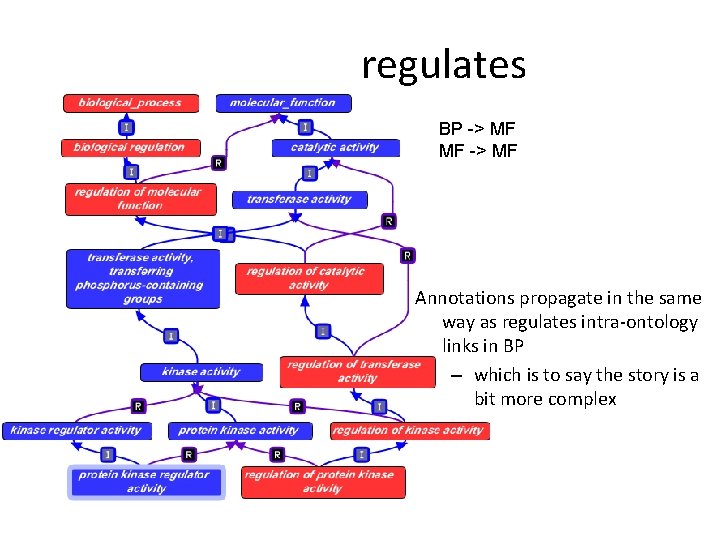
regulates BP -> MF MF -> MF Annotations propagate in the same way as regulates intra-ontology links in BP – which is to say the story is a bit more complex

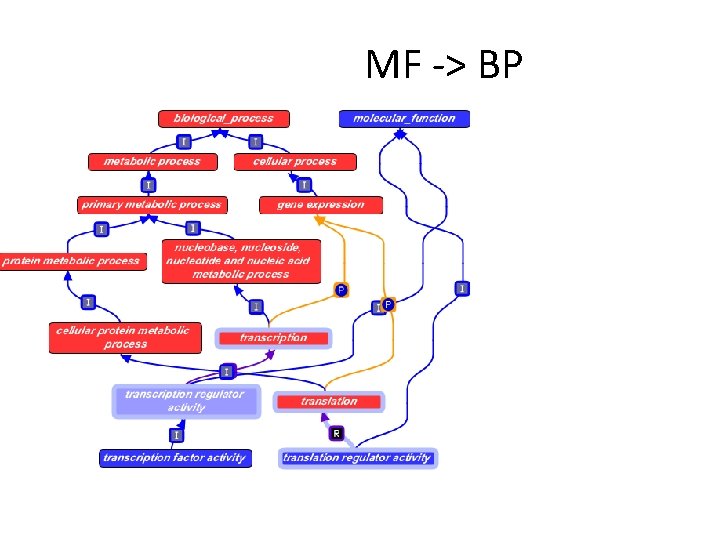
MF -> BP

Current progress on low hanging fruit • part_of – – – 134 F->P links safe to add http: //www. geneontology. org/scratch/fp-links/ Many derived from CHEBI cross-products Mostly transporters, kinases, synthetases. . Lots more if we include metabolism • Regulates – 110 links • These will be implemented in the live GO on 2009/mm/dd

Connecting process to function : pathways • Can we use part_of with pathways? – rarely – Results in true path violations – Example: • glycolysis – 6 -phosphofructokinase activity • Solution – has_part

Pathways and their parts • GO: 0006096 ! glycolysis • • • part_of • • GO: 0003872 ! 6 -phosphofructokinase activity GO: 0004332 ! fructose-bisphosphate aldolase activity GO: 0004347 ! glucose-6 -phosphate isomerase activity GO: 0004365 ! glyceraldehyde-3 -phosphate dehydrogenase (phosphorylating) activity GO: 0004618 ! phosphoglycerate kinase activity GO: 0004619 ! phosphoglycerate mutase activity GO: 0004634 ! phosphopyruvate hydratase activity GO: 0004743 ! pyruvate kinase activity GO: 0004807 ! triose-phosphate isomerase activity

Pathways and their parts • GO: 0006096 ! glycolysis • • • has_part • • GO: 0003872 ! 6 -phosphofructokinase activity GO: 0004332 ! fructose-bisphosphate aldolase activity GO: 0004347 ! glucose-6 -phosphate isomerase activity GO: 0004365 ! glyceraldehyde-3 -phosphate dehydrogenase (phosphorylating) activity GO: 0004618 ! phosphoglycerate kinase activity GO: 0004619 ! phosphoglycerate mutase activity GO: 0004634 ! phosphopyruvate hydratase activity GO: 0004743 ! pyruvate kinase activity GO: 0004807 ! triose-phosphate isomerase activity

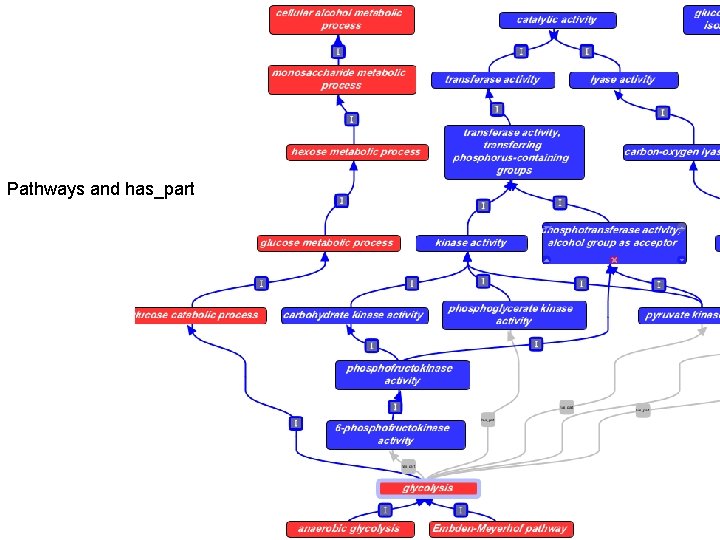
Pathways and has_part

Annotations do NOT propagate • …but the links can be used to suggest annotations

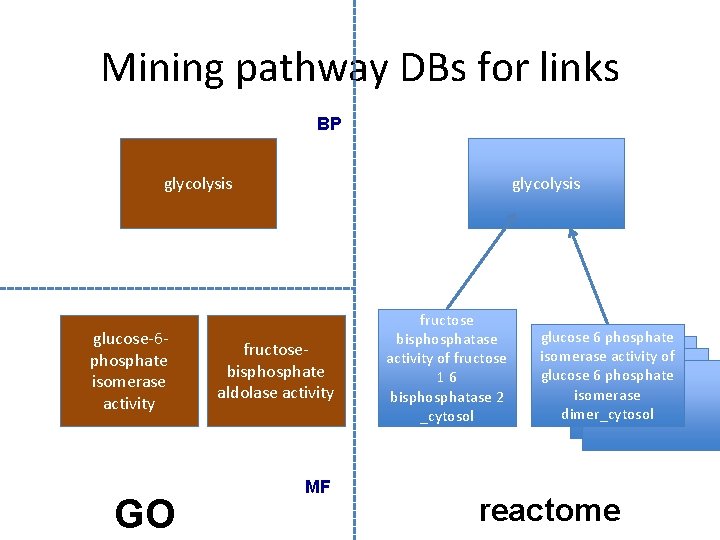
Mining pathway DBs for links BP glycolysis glucose-6 phosphate isomerase activity GO glycolysis fructosebisphosphate aldolase activity MF fructose bisphosphatase activity of fructose 16 bisphosphatase 2 _cytosol glucose 6 phosphate isomerase activity of glucose 6 phosphate isomerase dimer_cytosol reactome

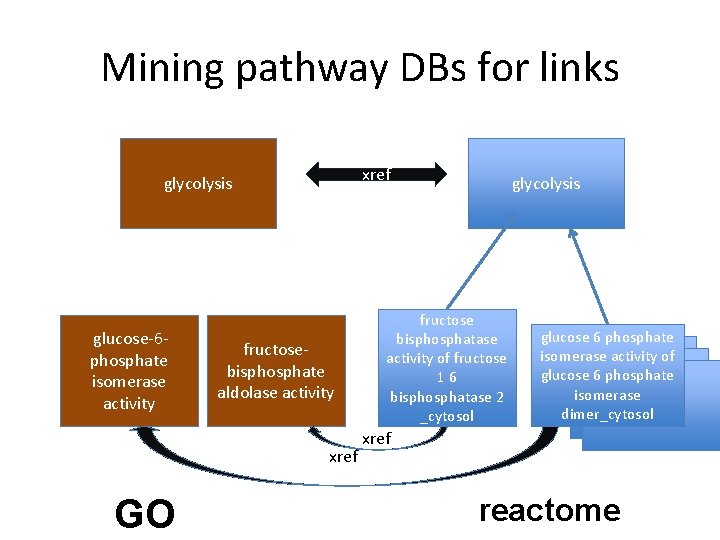
Mining pathway DBs for links xref glycolysis glucose-6 phosphate isomerase activity fructosebisphosphate aldolase activity xref GO glycolysis fructose bisphosphatase activity of fructose 16 bisphosphatase 2 _cytosol glucose 6 phosphate isomerase activity of glucose 6 phosphate isomerase dimer_cytosol xref reactome

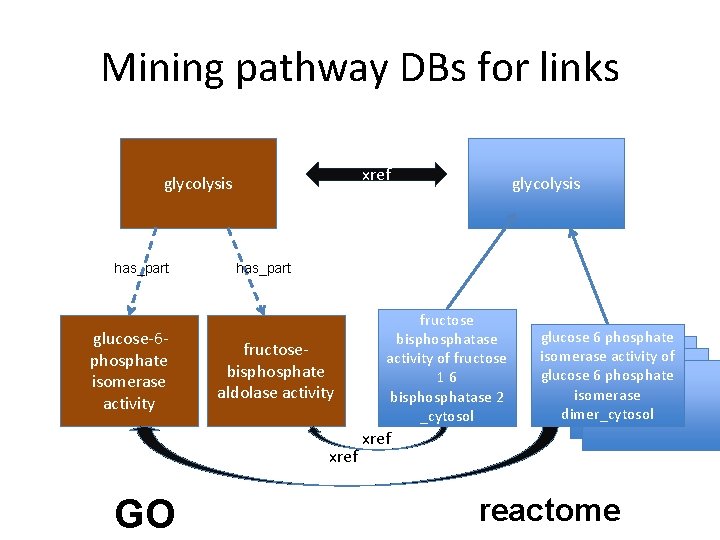
Mining pathway DBs for links xref glycolysis has_part glucose-6 phosphate isomerase activity has_part fructosebisphosphate aldolase activity xref GO glycolysis fructose bisphosphatase activity of fructose 16 bisphosphatase 2 _cytosol glucose 6 phosphate isomerase activity of glucose 6 phosphate isomerase dimer_cytosol xref reactome

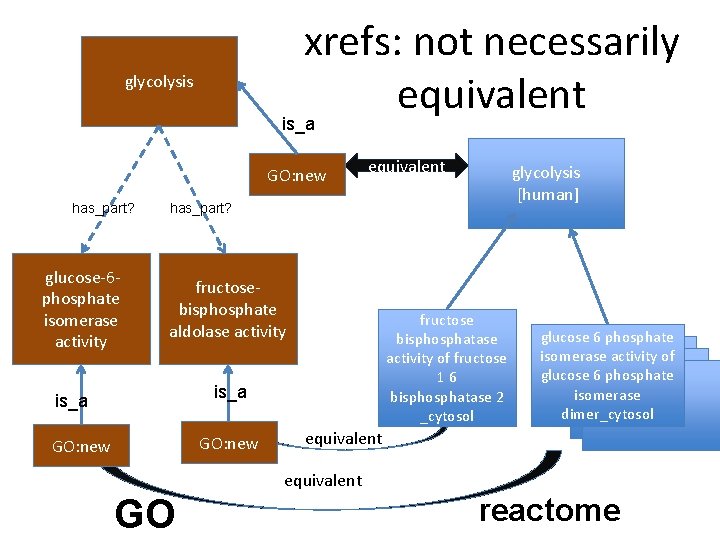
xrefs: not necessarily equivalent glycolysis is_a GO: new has_part? glucose-6 phosphate isomerase activity equivalent glycolysis [human] has_part? fructosebisphosphate aldolase activity fructose bisphosphatase activity of fructose 16 bisphosphatase 2 _cytosol is_a GO: new glucose 6 phosphate isomerase activity of glucose 6 phosphate isomerase dimer_cytosol equivalent GO reactome
![xrefs not necessarily equivalent glycolysis isa somehaspart GO new equivalent glycolysis human somehaspart glucose6 xrefs: not necessarily equivalent glycolysis is_a some_has_part GO: new equivalent glycolysis [human] some_has_part glucose-6](https://slidetodoc.com/presentation_image_h2/a382c122be2a1544603165d58a150c1f/image-32.jpg)
xrefs: not necessarily equivalent glycolysis is_a some_has_part GO: new equivalent glycolysis [human] some_has_part glucose-6 phosphate isomerase activity fructosebisphosphate aldolase activity fructose bisphosphatase activity of fructose 16 bisphosphatase 2 _cytosol is_a GO: new glucose 6 phosphate isomerase activity of glucose 6 phosphate isomerase dimer_cytosol equivalent GO reactome
![xrefs not necessarily equivalent glycolysis xref somehaspart glycolysis human somehaspart glucose6 phosphate isomerase activity xrefs: not necessarily equivalent glycolysis xref some_has_part glycolysis [human] some_has_part glucose-6 phosphate isomerase activity](https://slidetodoc.com/presentation_image_h2/a382c122be2a1544603165d58a150c1f/image-33.jpg)
xrefs: not necessarily equivalent glycolysis xref some_has_part glycolysis [human] some_has_part glucose-6 phosphate isomerase activity fructosebisphosphate aldolase activity fructose bisphosphatase activity of fructose 16 bisphosphatase 2 _cytosol glucose 6 phosphate isomerase activity of glucose 6 phosphate isomerase dimer_cytosol xref GO reactome

Progress on mining links from pathway databases • Methods – We used biopax OWL dumps from Meta. Cyc and Reactome • For Metacyc, used the xrefs from GO to Meta. Cyc • For Reactome, used the xrefs from Reactome to GO • Results – 2896 links between BP and MF • Reactome: 1249 • Meta. Cyc: 1697 – Good correlation with curated links • Issues – technical: inconsistencies in biopax representation – curational: xrefs incomplete or could be improved



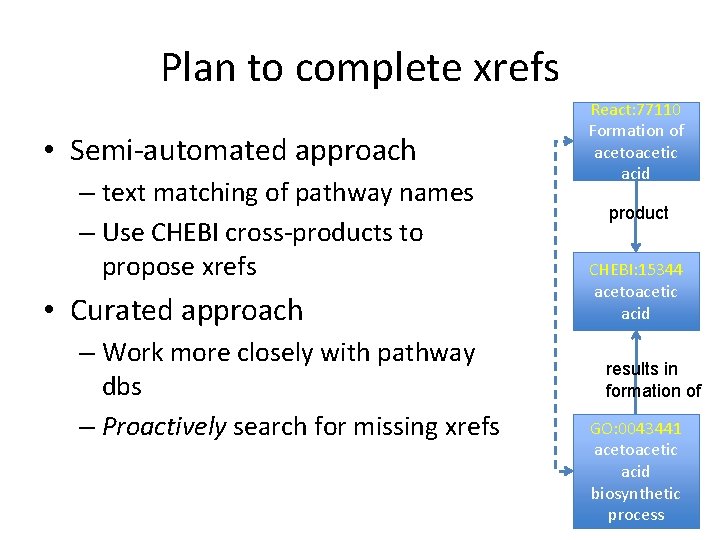
Plan to complete xrefs • Semi-automated approach – text matching of pathway names – Use CHEBI cross-products to propose xrefs • Curated approach – Work more closely with pathway dbs – Proactively search for missing xrefs React: 77110 Formation of acetoacetic acid product CHEBI: 15344 acetoacetic acid results in formation of GO: 0043441 acetoacetic acid biosynthetic process

More advanced co-annotation • Annotation cross-products

Component-function • We can make links from Cellular component too – Example: • histone deacetylase complex

Implementation plan • 2009/02/nn

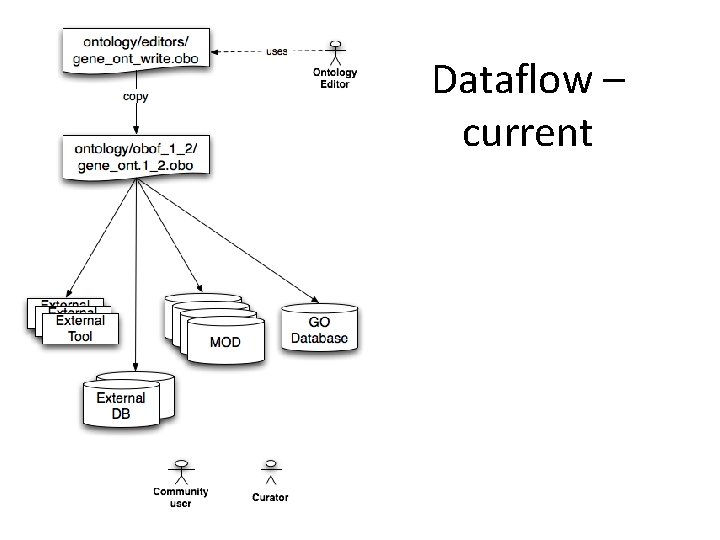
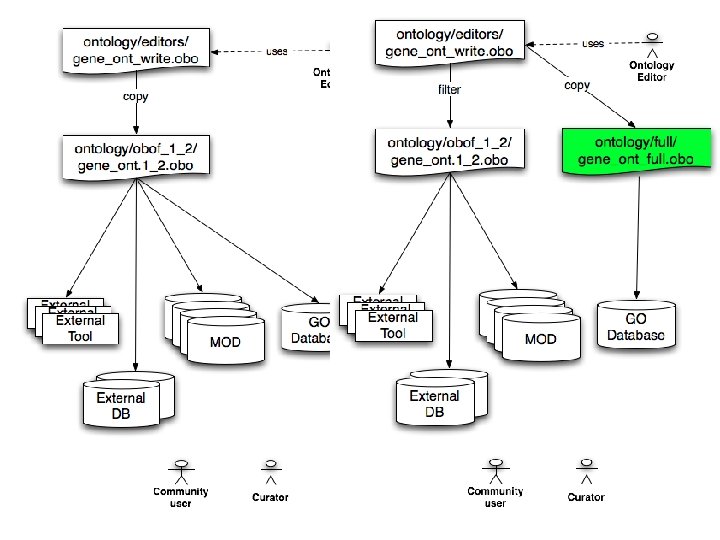
Dataflow – current

Dataflow – current
![Conclusions so far Implement LowHangingHighYieldFruit ASAP MF partof BP regulates Conclusions [so far] • Implement Low-Hanging-High-Yield-Fruit ASAP – MF part_of BP – regulates •](https://slidetodoc.com/presentation_image_h2/a382c122be2a1544603165d58a150c1f/image-43.jpg)
Conclusions [so far] • Implement Low-Hanging-High-Yield-Fruit ASAP – MF part_of BP – regulates • Pathways – How much curation effort? • curate xrefs only and mine all links? • Curate has_part links too? – Work with pathway dbs to unify exchange formats and make data interoperable

Action items • Resuscitate xref dataflow with reactome

• stop here

• I told you to stop

• Don’t say I didn’t warn you…

Relations in GO for 2009

Intro • We have many relations ready to GO live in the scratch directory – within GO ontologies – across GO ontologies – between GO and external ontologies – Both cross product (N+S conditions) and regular links • Requires a fundamental change in how we and our users think about GO and annotations – Tools that make use of these will better serve users

Relations in GO • In the beginning there was is_a and part_of – Benefits: simplicity • We could effectively ignore relations • Most tools and users effectively do this – Speculation: recent introduction of regulates had no effect on majority of users – Drawbacks: lack of expressivity • We need more relations – – Regulation Spatial relations has_part for Process-Function annotations

Example of a relation rule in GO • Rule: – A is_a B, B is_a C A is_a C • Example: • We can generalize this by having a rule for transitive relations – transitive r, A r B, B r C A r C • We can also write this as a composition rule: – is_a – Open question: • does this notation help or hinder? ?

Transitivity • We currently have two transitive relations in GO: – is_a – part_of • Example: – mitotic prophase part_of mitosis – In GO, part_of is an all-some relation • regulates is not defined to be transitive in GO • (but the majority of tools still treat it as if it were!) • Example:

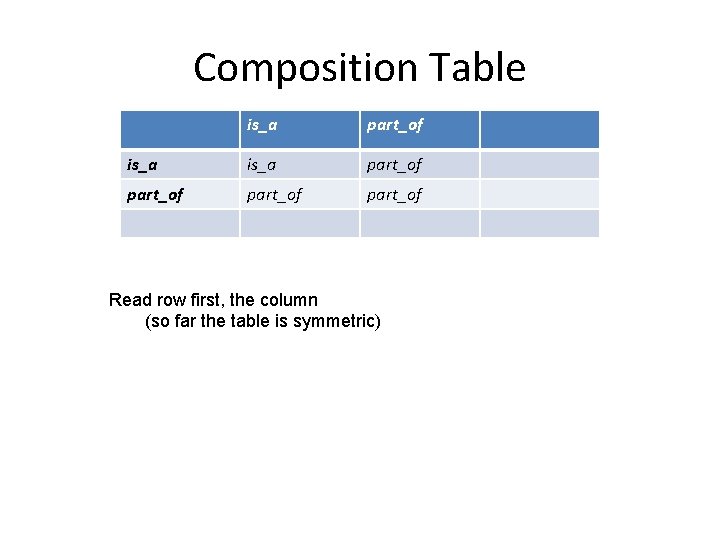
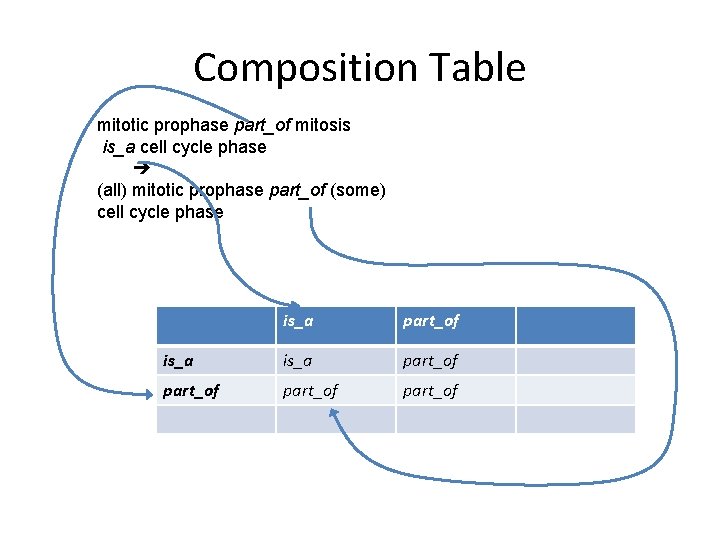
Composition with is_a • Any relation that follows the all-some pattern composes with is_a to itself • Example: – (all) nucleus part_of (some) cell • Composition: – is_a. R R – R. is_a R • Example: – (all) mitotic prophase part_of (some) mitosis – mitosis is_a cell cycle phase • – (all) mitotic prophase part_of (some) cell cycle phase

Composition Table is_a part_of part_of Read row first, the column (so far the table is symmetric)

Composition Table mitotic prophase part_of mitosis is_a cell cycle phase (all) mitotic prophase part_of (some) cell cycle phase is_a part_of part_of

Chained compositions A part_f B is_a C is_a D part_of E A part_of B is_a D part_of E A part_of E order of reduction does not matter is_a part_of part_of

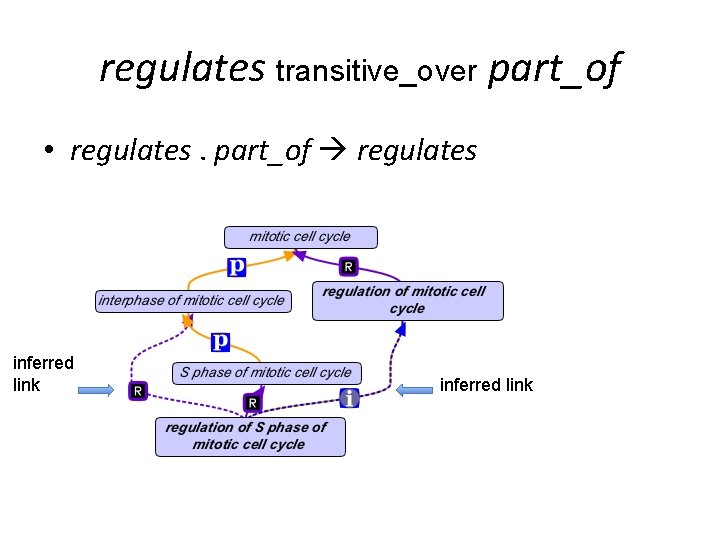
regulates transitive_over part_of • regulates. part_of regulates inferred link

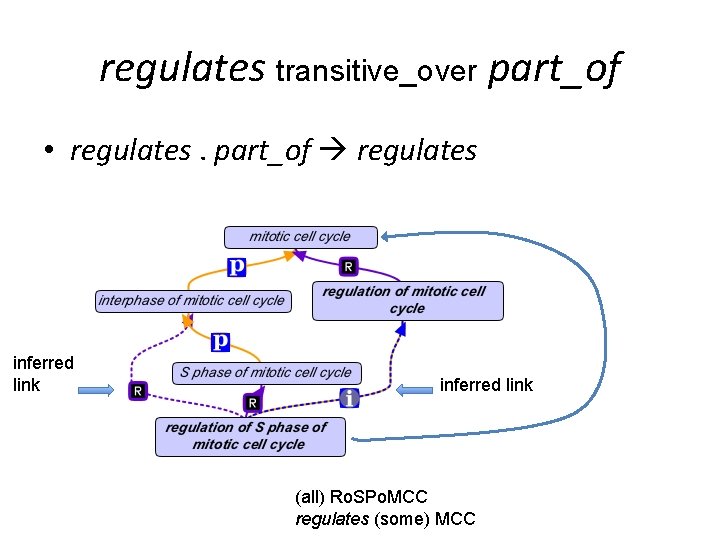
regulates transitive_over part_of • regulates. part_of regulates inferred link (all) Ro. SPo. MCC regulates (some) MCC

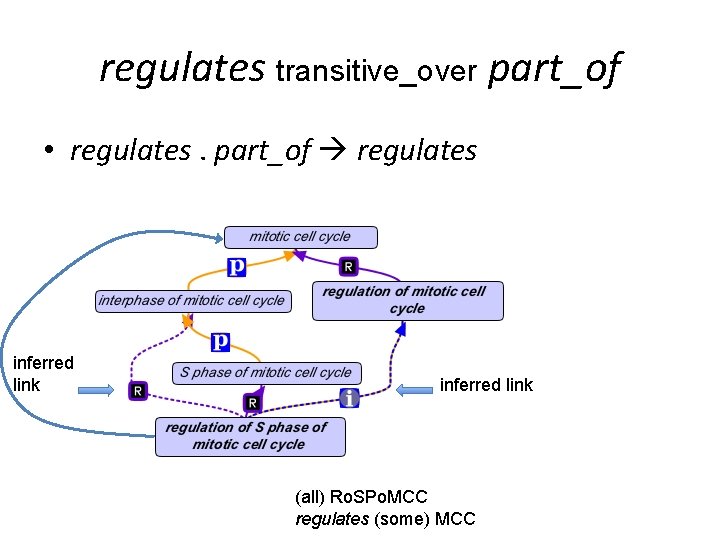
regulates transitive_over part_of • regulates. part_of regulates inferred link (all) Ro. SPo. MCC regulates (some) MCC

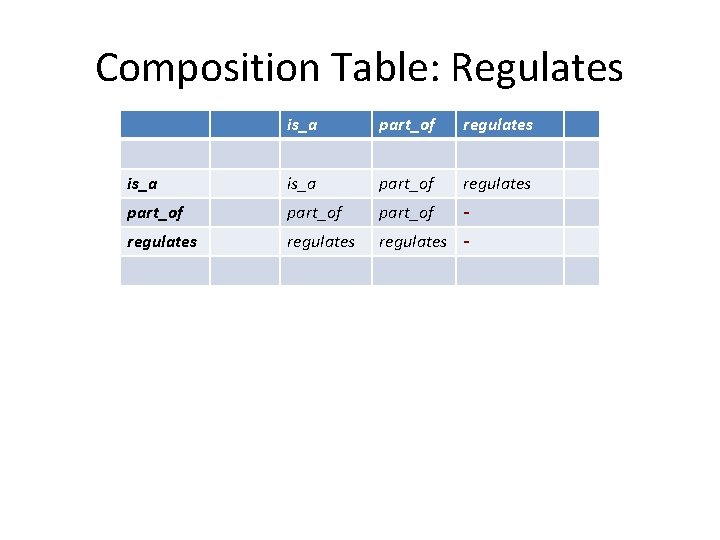
Composition Table: Regulates is_a part_of regulates part_of - regulates -

Composition Table: Regulates is_a part_of regulates part_of - regulates - regulates. part_of regulates

Composition Table: Regulates is_a part_of regulates part_of N/A regulates - part_of. regulates N/A

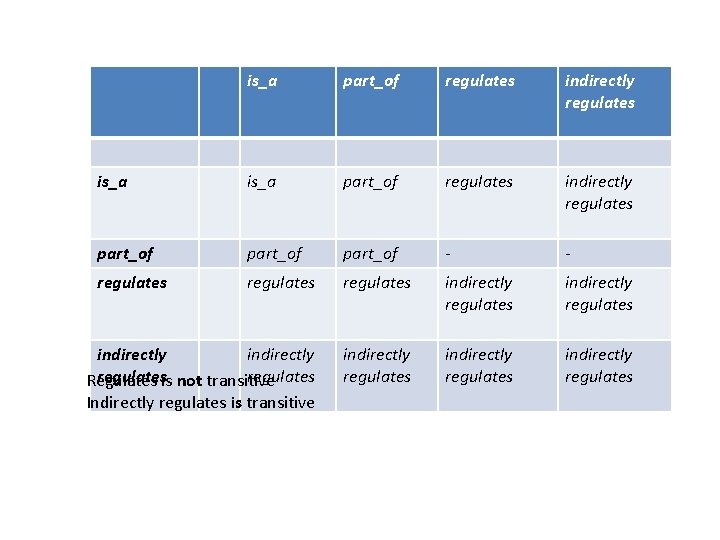
is_a part_of regulates part_of - regulates indirectly regulates We have the option of defining additional relations These may be entirely implicit (i. e. we would never assert indirectly regulates in GO)

is_a part_of regulates indirectly regulates part_of - - regulates indirectly regulatesis not transitive regulates Regulates Indirectly regulates is transitive indirectly regulates

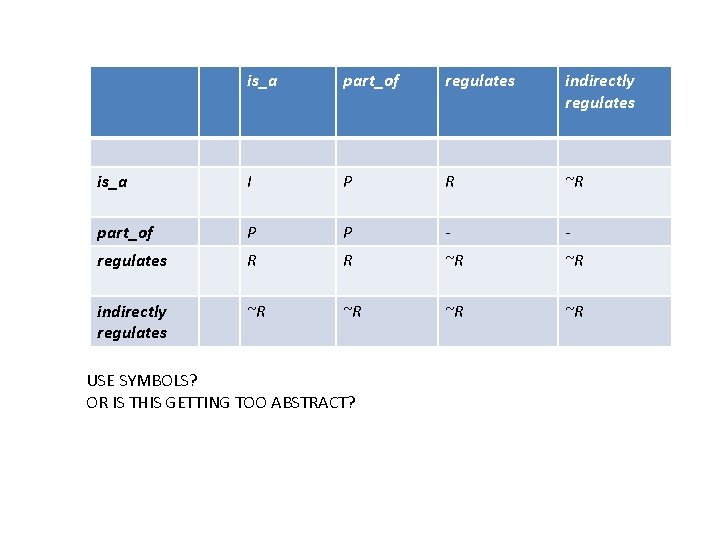
is_a part_of regulates indirectly regulates is_a I P R ~R part_of P P - - regulates R R ~R ~R indirectly regulates ~R ~R USE SYMBOLS? OR IS THIS GETTING TOO ABSTRACT?

Sub-relations • regulates – negatively_regulates – positively_regulates

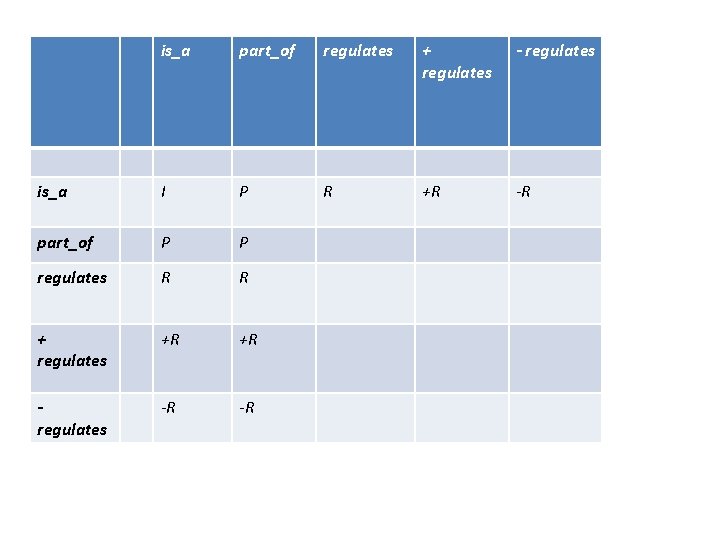
is_a part_of regulates + regulates - regulates is_a I P R +R -R part_of P P regulates R R + regulates +R +R regulates -R -R

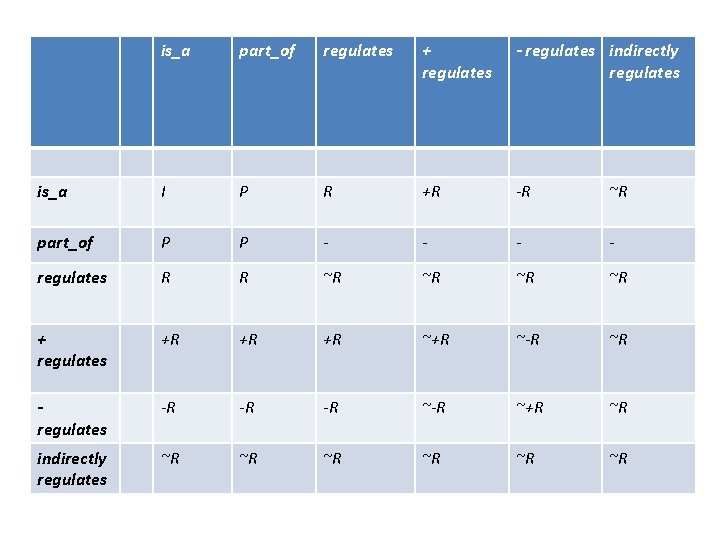
is_a part_of regulates + regulates - regulates indirectly regulates is_a I P R +R -R ~R part_of P P - - regulates R R ~R ~R + regulates +R +R +R ~-R ~R regulates -R -R -R ~+R ~R indirectly regulates ~R ~R ~R

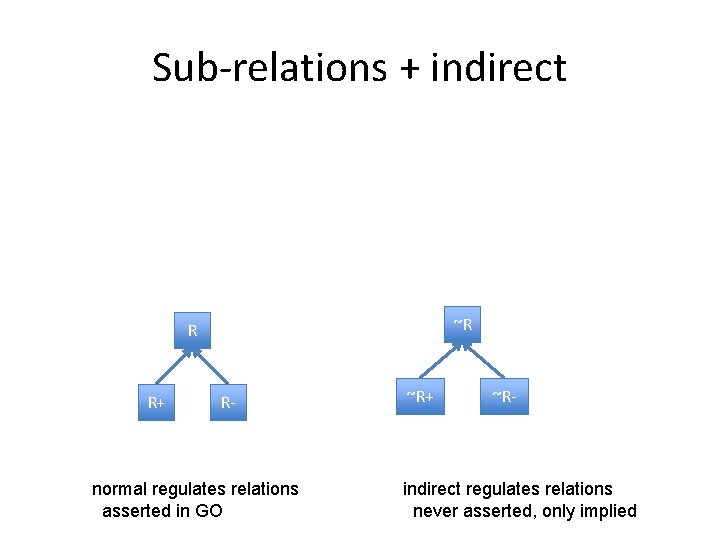
Sub-relations + indirect ~R R R+ R- normal regulates relations asserted in GO ~R+ ~R- indirect regulates relations never asserted, only implied

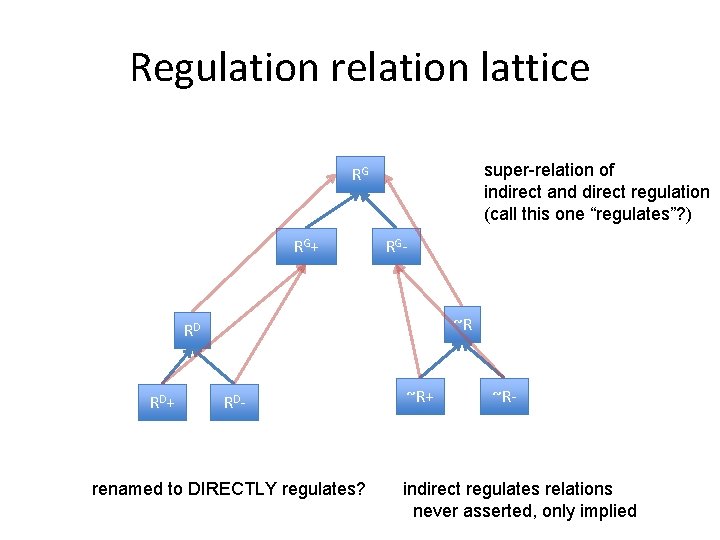
Regulation relation lattice super-relation of indirect and direct regulation (call this one “regulates”? ) RG RG+ RG- ~R RD RD+ RD- renamed to DIRECTLY regulates? ~R+ ~R- indirect regulates relations never asserted, only implied

has_part • NOT the inverse of part_of at the ontology level • Example: – nucleus part_of cell: YES • every nucleus is part_of some cell – by definition; e. g. extruded nuclei are ex-nuclei – cell has_part nucleus: NO • not every cell has_part nucleus – mammalian erythrocytes, bacteria • Example: – <pf example here> – <summarise pf progress>

Annotations and relations • not just an ontology issue – this is of relevance to annotations too… • The current simple methodology of propagating annotations up the graph only works for a small subset of relations – To understand how annotations and new relations interact we must think in terms of gene product relations

Gene product relations • What is the relation between a gene product and – A molecular function? – A biological process? – A cellular component? • Why care? • What’s wrong with “annotated_to”? – We need to define these relations: • to do justice to the biology • to be able to deal with new relations within the GO itself

Why we should care • How should annotation queries, analysis tools (slimmers, enrichment tools) etc treat the (pseudo-)new regulates relation? • How should we recommend the processfunction links be vizualized? • How should these links be treated in queries?

Proposed relations for gene products • For MF and BP: – has_potential – has_function_during • For CC: – localized_to/acts_in Names TBD MFs are ontologically like BPs (bfo processes)…. – This is more specific than has_location • A gene product may travel through different locations – Formally: • GP localized_to CC : GP executes some function in CC

How to read a GAF • <gene product> <rel> <GO term> • gene product may not be explicitly in GAF – that’s OK – gene as proxy • The relation does NOT apply to the gene however • genes are only localized_to chromosomes, and only participate in gene expression. It’s the products that do the work • <rel> is implicit, depending on F, C or P • Examples:

Annotation relation composition • is_a – always propagate over is_a • localized_to. is_a localized_to • has_function_in. is_a has_function_in • part_of • localized_to. part_of localized_to • has_function_in. part_of has_function_in • This is effectively what we do with gene product annotations now • post-hoc logical justification for why it’s OK to propagate

Annotation relation composition: regulates • regulates – localized_to. regulates NEVER POSSIBLE • localized_to never has a process as target • regulates always has process as subject – has_function_in. regulates regulator_of • This introduces an addition implicit relation that can be used to sum gene product results – Fake Ami. GO screenshot here


Annotation relation composition: inter -ontology links • We have 183 CC->MF/BP links in scratch • regulates – localized_to. has_function_in ? ? may_contribute_to? ? • Example: • RPS 25 A localized_to ribosome • ribosome has_function_in protein biosynthesis – • RPS 25 A ? ? has_function_in? ? protein biosynthesis • No need for curator to make explicit IC claims • • Q: we never want “may” in relation names? Can we make a stronger claim? How does a curator know when to make an IC claim here? Potential confusion with contributes_to qualifier

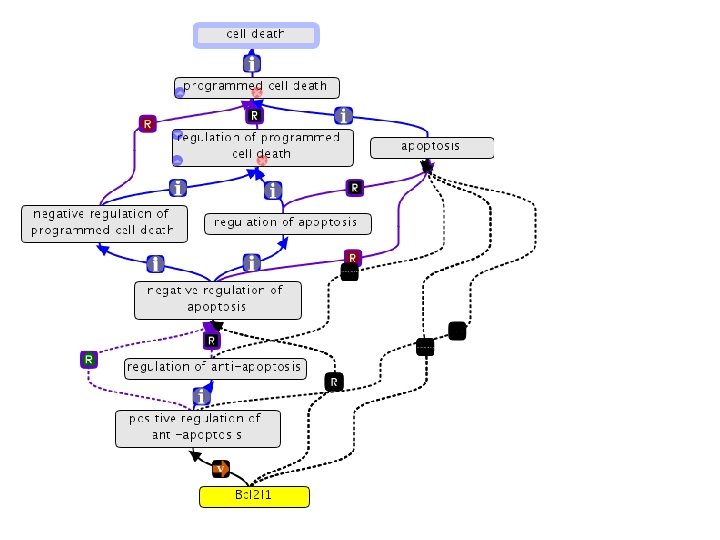
Annotation relations and has_part • Need some graphical illustrations • See – http: //wiki. geneontology. org/index. php/Has_part – for now

Qualifiers • Annotation qualifiers (contributes_to) have the effect of modifying the relation – NOT is not a qualifier – it is a logical operator • We can add new relations to the qualifier column – gene. Product. A acted_on_during protein secretion by the type II secretion system

Secondary taxon IDs

Cell component relations • We have 674 xp defs within CC in scratch – adjacent_to – surrounds/surrounded_by – spans – overlaps • Use case: reactome • Can we say anything about gene products here? – we can perform spatial gene product queries

Spatial reasoning – spans. adjacent_to overlaps (? ? TBD!!) – SUN-KASH complex spans nuclear inner membrance – nuclear inner membrane adjacent_to nuclear lumen – – SUN-KASH complex overlaps nuclear lumen

Links from BP to external ontologies • Process-continuant links – A has_function_in cysteine biosynthesis • A ? ? has_participant? ? cysteine • this is true but can we make stronger claims – A has_function_in heart development • A has_participant heart • c. f. heart process, TAZ gene • How can we use this? – Browse GO annotations via other ontologies – Enrichment using anatomy terms… – Ami. GO screenshots

what next?

Won’t this confuse users? • We will provide a pre-made inferred relation table for all of GO – we could do this for gps too but it would be over a billion entries. . • We can always distribute a dumb. GO – just is_a and part_of, not even regulates • Need more guidance on how this can be used

Discussion

What’s next? • Move relations into GO editors file – post OE 2 – CC-self • spatial relations – BP->MF • has_part • regulates – BP->BP • has_part (? ? ) – External onts • Dual releases? dumb. GO and full. GO? • Fix GOC tools (Ami. GO, slimmer, enrichment, graphviz, ref. G) to deal appropriately – OE 2 should already be fine • Educate non-GOC folks