Conj 532 Cytokine JakSTAT Pathways Thelper cell subsets
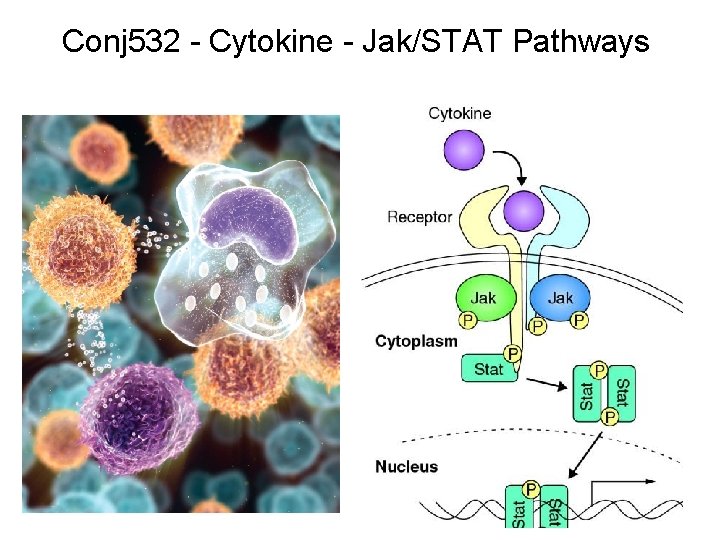
Conj 532 - Cytokine - Jak/STAT Pathways
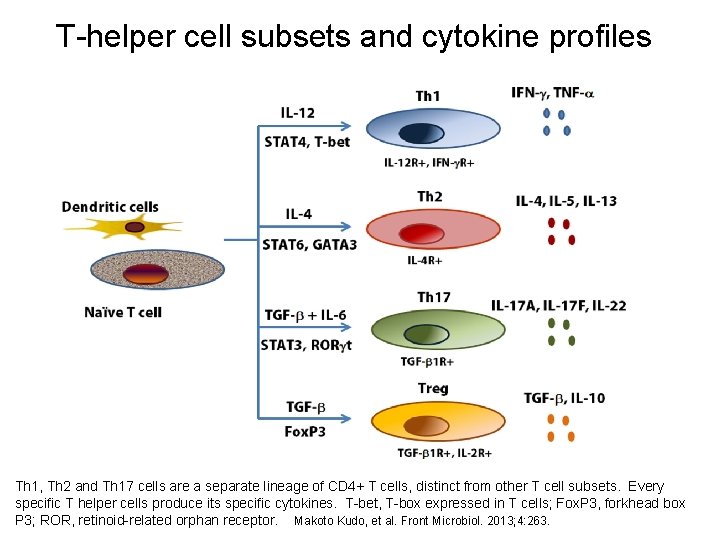
T-helper cell subsets and cytokine profiles Th 1, Th 2 and Th 17 cells are a separate lineage of CD 4+ T cells, distinct from other T cell subsets. Every specific T helper cells produce its specific cytokines. T-bet, T-box expressed in T cells; Fox. P 3, forkhead box P 3; ROR, retinoid-related orphan receptor. Makoto Kudo, et al. Front Microbiol. 2013; 4: 263.
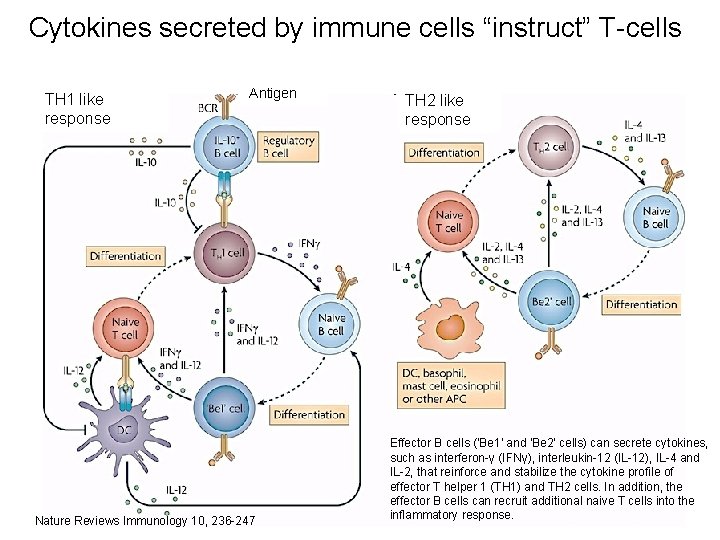
Cytokines secreted by immune cells “instruct” T-cells TH 1 like response Antigen Nature Reviews Immunology 10, 236 -247 TH 2 like response Effector B cells ('Be 1' and 'Be 2' cells) can secrete cytokines, such as interferon-γ (IFNγ), interleukin-12 (IL-12), IL-4 and IL-2, that reinforce and stabilize the cytokine profile of effector T helper 1 (TH 1) and TH 2 cells. In addition, the effector B cells can recruit additional naive T cells into the inflammatory response.
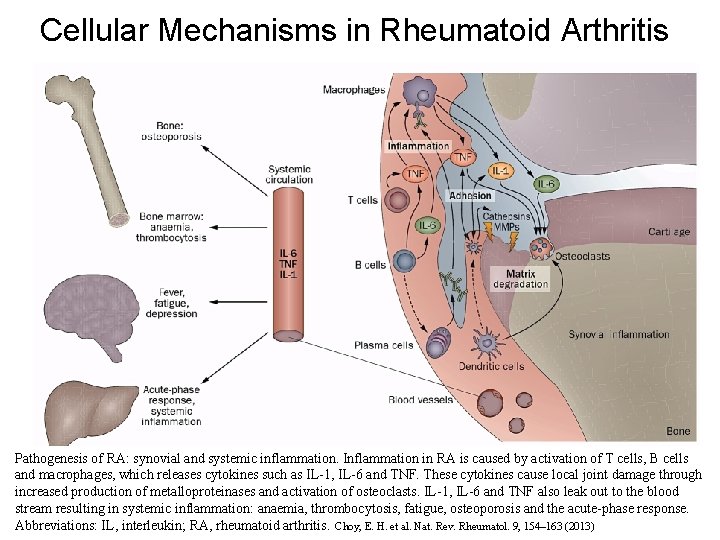
Cellular Mechanisms in Rheumatoid Arthritis Pathogenesis of RA: synovial and systemic inflammation. Inflammation in RA is caused by activation of T cells, B cells and macrophages, which releases cytokines such as IL-1, IL-6 and TNF. These cytokines cause local joint damage through increased production of metalloproteinases and activation of osteoclasts. IL-1, IL-6 and TNF also leak out to the blood stream resulting in systemic inflammation: anaemia, thrombocytosis, fatigue, osteoporosis and the acute-phase response. Abbreviations: IL, interleukin; RA, rheumatoid arthritis. Choy, E. H. et al. Nat. Rev. Rheumatol. 9, 154– 163 (2013)
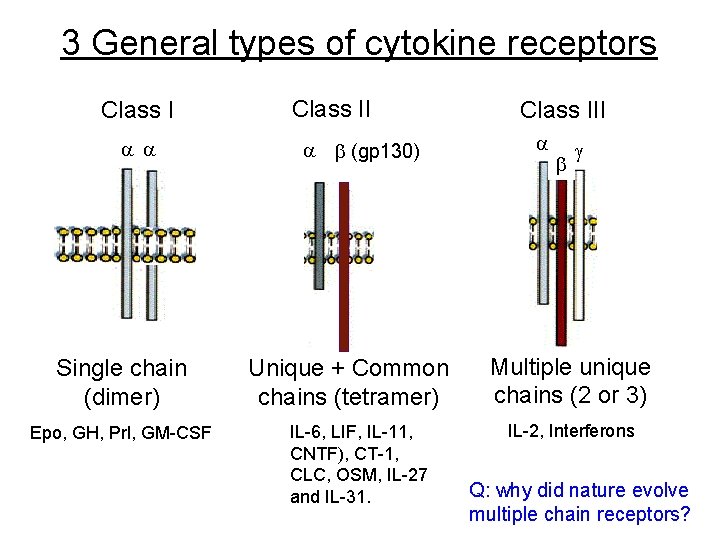
3 General types of cytokine receptors Class I a a Single chain (dimer) Epo, GH, Prl, GM-CSF Class II a b (gp 130) Unique + Common chains (tetramer) IL-6, LIF, IL-11, CNTF), CT-1, CLC, OSM, IL-27 and IL-31. Class III a b g Multiple unique chains (2 or 3) IL-2, Interferons Q: why did nature evolve multiple chain receptors?
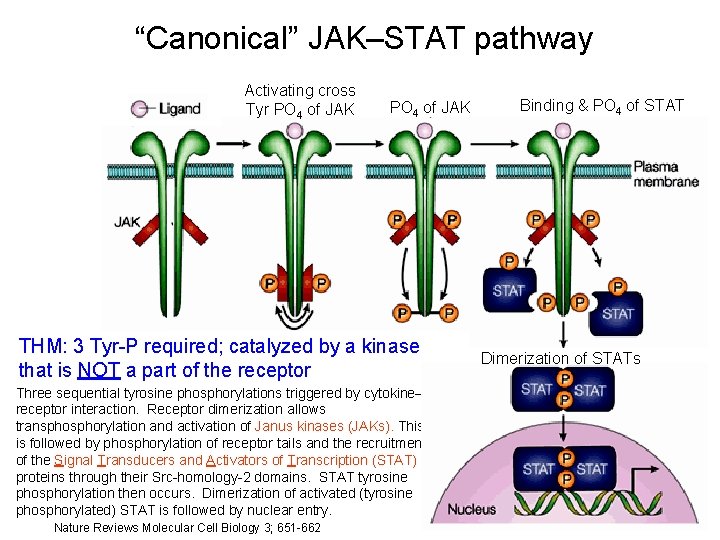
“Canonical” JAK–STAT pathway Activating cross Tyr PO 4 of JAK THM: 3 Tyr-P required; catalyzed by a kinase that is NOT a part of the receptor Three sequential tyrosine phosphorylations triggered by cytokine– receptor interaction. Receptor dimerization allows transphorylation and activation of Janus kinases (JAKs). This is followed by phosphorylation of receptor tails and the recruitment of the Signal Transducers and Activators of Transcription (STAT) proteins through their Src-homology-2 domains. STAT tyrosine phosphorylation then occurs. Dimerization of activated (tyrosine phosphorylated) STAT is followed by nuclear entry. Nature Reviews Molecular Cell Biology 3; 651 -662 Binding & PO 4 of STAT Dimerization of STATs

So, lets go into more detail about each of these players
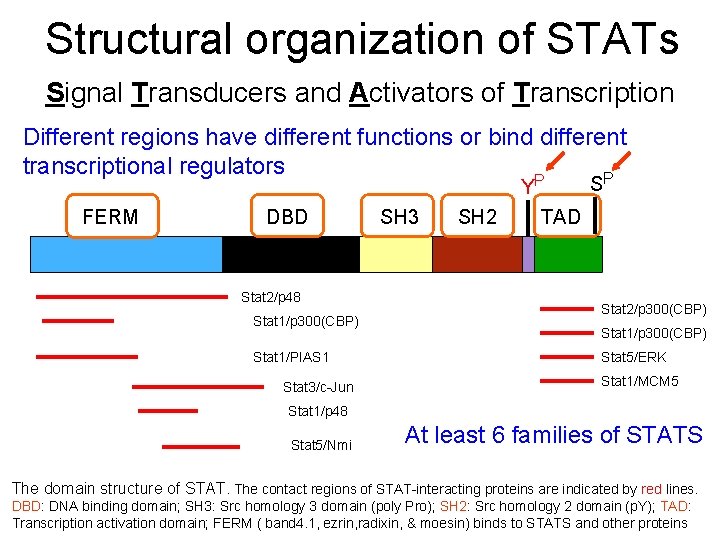
Structural organization of STATs Signal Transducers and Activators of Transcription Different regions have different functions or bind different transcriptional regulators P YP S FERM DBD SH 3 SH 2 TAD Stat 2/p 48 Stat 1/p 300(CBP) Stat 1/PIAS 1 Stat 3/c-Jun Stat 2/p 300(CBP) Stat 1/p 300(CBP) Stat 5/ERK Stat 1/MCM 5 Stat 1/p 48 Stat 5/Nmi At least 6 families of STATS The domain structure of STAT. The contact regions of STAT-interacting proteins are indicated by red lines. 8 DBD: DNA binding domain; SH 3: Src homology 3 domain (poly Pro); SH 2: Src homology 2 domain (p. Y); TAD: Transcription activation domain; FERM ( band 4. 1, ezrin, radixin, & moesin) binds to STATS and other proteins
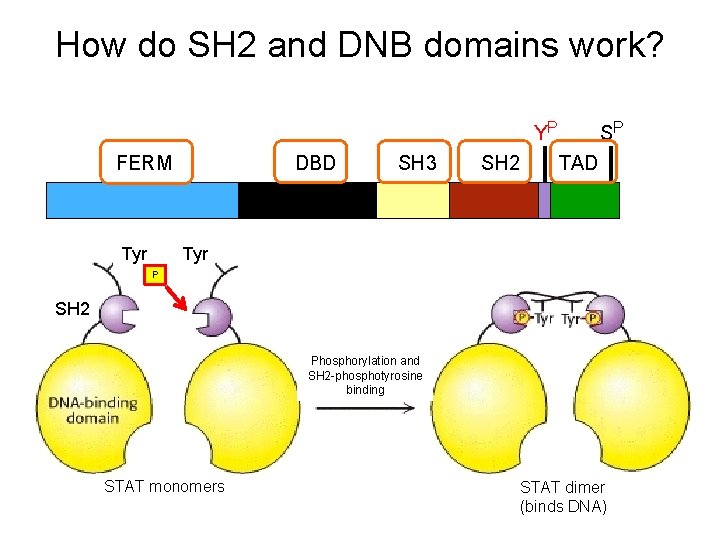
How do SH 2 and DNB domains work? YP SP FERM DBD SH 3 SH 2 TAD Tyr P SH 2 Phosphorylation and SH 2 -phosphotyrosine binding STAT monomers STAT dimer (binds DNA)
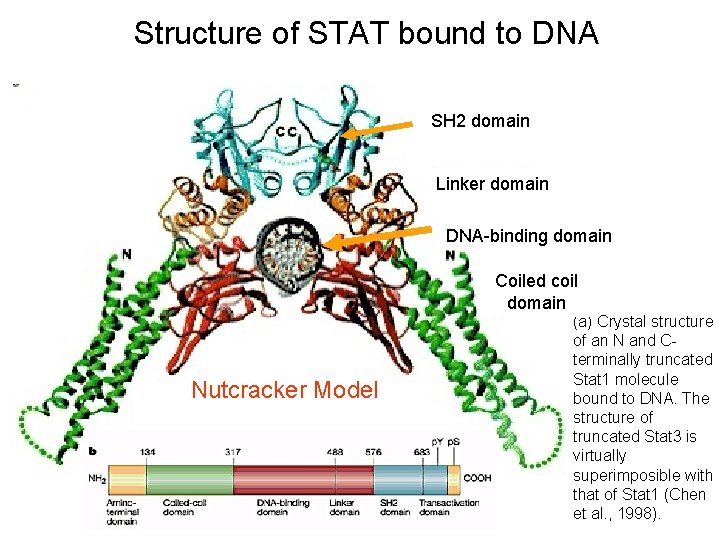
Structure of STAT bound to DNA SH 2 domain Linker domain DNA-binding domain Coiled coil domain Nutcracker Model (a) Crystal structure of an N and Cterminally truncated Stat 1 molecule bound to DNA. The structure of truncated Stat 3 is virtually superimposible with that of Stat 1 (Chen et al. , 1998).
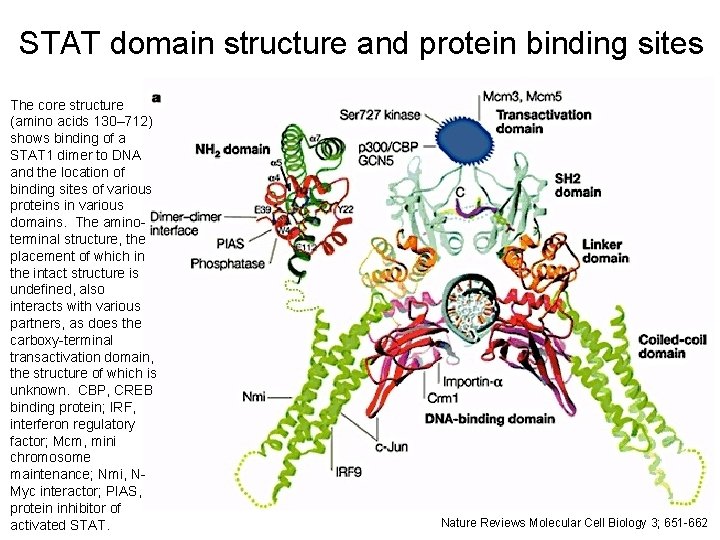
STAT domain structure and protein binding sites The core structure (amino acids 130– 712) shows binding of a STAT 1 dimer to DNA and the location of binding sites of various proteins in various domains. The aminoterminal structure, the placement of which in the intact structure is undefined, also interacts with various partners, as does the carboxy-terminal transactivation domain, the structure of which is unknown. CBP, CREB binding protein; IRF, interferon regulatory factor; Mcm, mini chromosome maintenance; Nmi, NMyc interactor; PIAS, protein inhibitor of activated STAT. 11 Nature Reviews Molecular Cell Biology 3; 651 -662
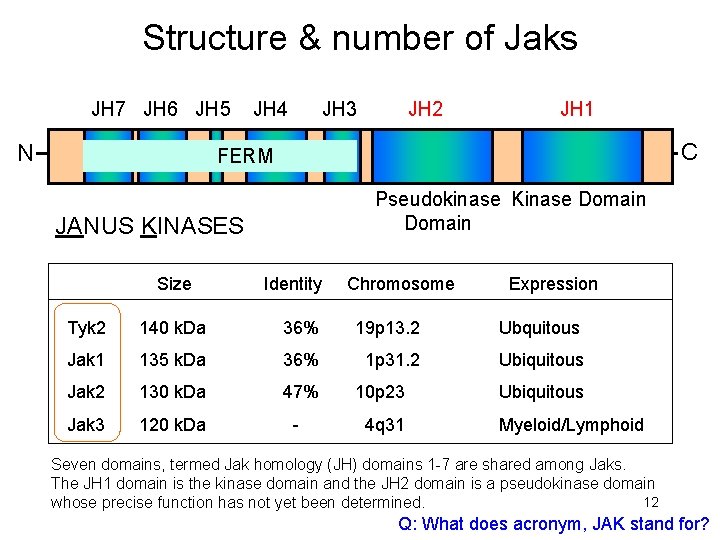
Structure & number of Jaks JH 7 JH 6 JH 5 JH 4 JH 3 JH 2 JH 1 N C FERM Pseudokinase Kinase Domain JANUS KINASES Size Identity Chromosome Expression Tyk 2 140 k. Da 36% 19 p 13. 2 Ubquitous Jak 1 135 k. Da 36% 1 p 31. 2 Ubiquitous Jak 2 130 k. Da 47% 10 p 23 Ubiquitous Jak 3 120 k. Da - 4 q 31 Myeloid/Lymphoid Seven domains, termed Jak homology (JH) domains 1 -7 are shared among Jaks. The JH 1 domain is the kinase domain and the JH 2 domain is a pseudokinase domain 12 whose precise function has not yet been determined. Q: What does acronym, JAK stand for?

Janus - the “two-faced” god, keeper of the gate The Janus kinases, were thought to contain 2 types of phosphate-transferring domains. Thus, it is named after “Janus”, the Roman two-faced gatekeeper of 13 the heavens.

Different cytokine receptors bind different combinations of Jaks TYPE HORMONE PHOSPHORYLATION Single specificity Growth Hormone Homo-phosphorylation Promiscuous (all JAKs) IL-6 Multi- phosphorylation Obligate Hetero (2 different JAKs) INFa, INFg Hetero-phosphorylation INFa INFg Jak 1 How determined? Tyk 2 (Mutate Jak 1 ---> no Tyk 2 PO 4 Jak 1 Why important? (Mutate Jak 1 ---> no Jak 2 PO 4 14 PO O 4 Different Jaks PO 4 different STATS Jak 2
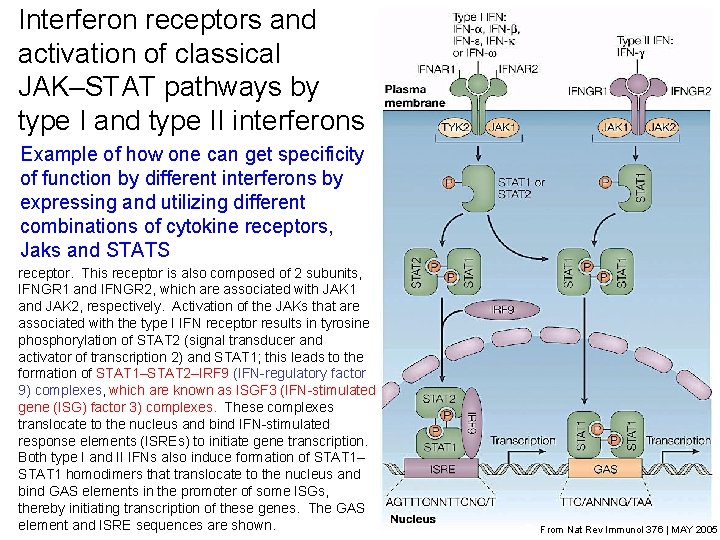
Interferon receptors and activation of classical JAK–STAT pathways by type I and type II interferons All type I interferons (IFNs) bind a common receptor Example of how one can get specificity which is known as the type I IFN receptor. The type I of function by different interferons by IFN receptor is composed of two subunits, IFNAR 1 and expressing and utilizing different IFNAR 2, which are associated with the Janus activated kinases (JAKs), tyrosine kinase 2 (TYK 2) and JAK 1, combinations of cytokine receptors, respectively. A single type II IFN, IFN-g, binds a distinct Jaks and STATS cell-surface receptor, which is known as the type II IFN receptor. This receptor is also composed of 2 subunits, IFNGR 1 and IFNGR 2, which are associated with JAK 1 and JAK 2, respectively. Activation of the JAKs that are associated with the type I IFN receptor results in tyrosine phosphorylation of STAT 2 (signal transducer and activator of transcription 2) and STAT 1; this leads to the formation of STAT 1–STAT 2–IRF 9 (IFN-regulatory factor 9) complexes, which are known as ISGF 3 (IFN-stimulated gene (ISG) factor 3) complexes. These complexes translocate to the nucleus and bind IFN-stimulated response elements (ISREs) to initiate gene transcription. Both type I and II IFNs also induce formation of STAT 1– STAT 1 homodimers that translocate to the nucleus and bind GAS elements in the promoter of some ISGs, thereby initiating transcription of these genes. The GAS element and ISRE sequences are shown. 15 From Nat Rev Immunol 376 | MAY 2005

How is Cytokine Function Regulated? A: Several types of negative feedback
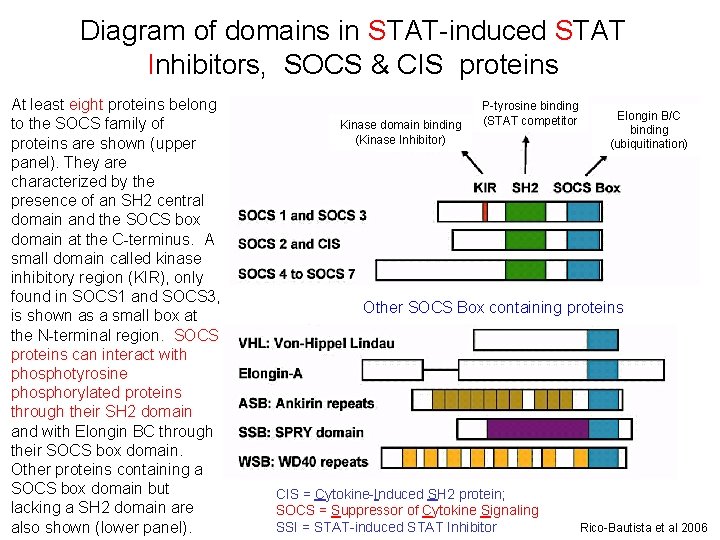
Diagram of domains in STAT-induced STAT Inhibitors, SOCS & CIS proteins At least eight proteins belong to the SOCS family of proteins are shown (upper panel). They are characterized by the presence of an SH 2 central domain and the SOCS box domain at the C-terminus. A small domain called kinase inhibitory region (KIR), only found in SOCS 1 and SOCS 3, is shown as a small box at the N-terminal region. SOCS proteins can interact with phosphotyrosine phosphorylated proteins through their SH 2 domain and with Elongin BC through their SOCS box domain. Other proteins containing a SOCS box domain but lacking a SH 2 domain are also shown (lower panel). Kinase domain binding (Kinase Inhibitor) P-tyrosine binding (STAT competitor Elongin B/C binding (ubiquitination) Other SOCS Box containing proteins CIS = Cytokine-Induced SH 2 protein; SOCS = Suppressor of Cytokine Signaling SSI = STAT-induced STAT Inhibitor Rico-Bautista et al 2006
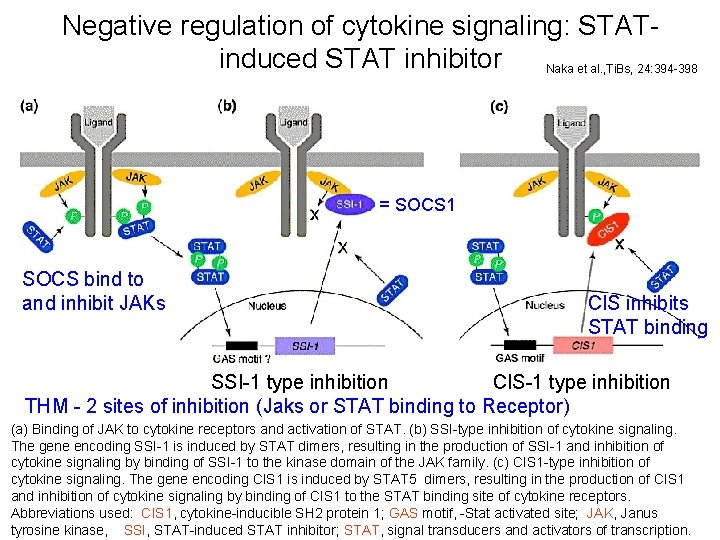
Negative regulation of cytokine signaling: STATinduced STAT inhibitor Naka et al. , Ti. Bs, 24: 394 -398 = SOCS 1 SOCS bind to and inhibit JAKs CIS inhibits STAT binding SSI-1 type inhibition CIS-1 type inhibition THM - 2 sites of inhibition (Jaks or STAT binding to Receptor) (a) Binding of JAK to cytokine receptors and activation of STAT. (b) SSI-type inhibition of cytokine signaling. The gene encoding SSI-1 is induced by STAT dimers, resulting in the production of SSI-1 and inhibition of cytokine signaling by binding of SSI-1 to the kinase domain of the JAK family. (c) CIS 1 -type inhibition of cytokine signaling. The gene encoding CIS 1 is induced by STAT 5 dimers, resulting in the production of CIS 1 and inhibition of cytokine signaling by binding of CIS 1 to the STAT binding site of cytokine receptors. 18 Abbreviations used: CIS 1, cytokine-inducible SH 2 protein 1; GAS motif, -Stat activated site; JAK, Janus tyrosine kinase, SSI, STAT-induced STAT inhibitor; STAT, signal transducers and activators of transcription.
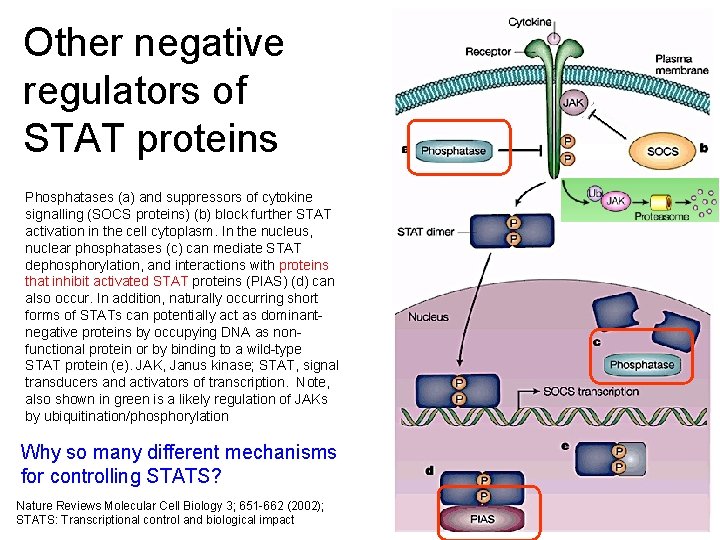
Other negative regulators of STAT proteins Phosphatases (a) and suppressors of cytokine signalling (SOCS proteins) (b) block further STAT activation in the cell cytoplasm. In the nucleus, nuclear phosphatases (c) can mediate STAT dephosphorylation, and interactions with proteins that inhibit activated STAT proteins (PIAS) (d) can also occur. In addition, naturally occurring short forms of STATs can potentially act as dominantnegative proteins by occupying DNA as nonfunctional protein or by binding to a wild-type STAT protein (e). JAK, Janus kinase; STAT, signal transducers and activators of transcription. Note, also shown in green is a likely regulation of JAKs by ubiquitination/phosphorylation Why so many different mechanisms for controlling STATS? Nature Reviews Molecular Cell Biology 3; 651 -662 (2002); STATS: Transcriptional control and biological impact 19
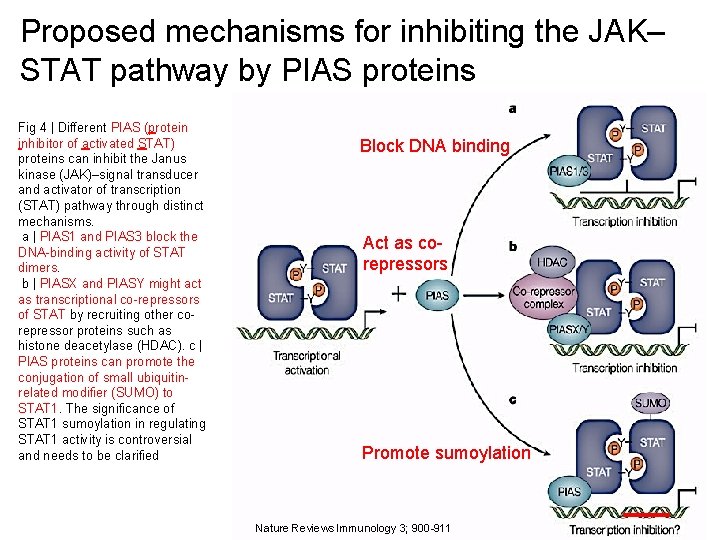
Proposed mechanisms for inhibiting the JAK– STAT pathway by PIAS proteins Fig 4 | Different PIAS (protein inhibitor of activated STAT) proteins can inhibit the Janus kinase (JAK)–signal transducer and activator of transcription (STAT) pathway through distinct mechanisms. a | PIAS 1 and PIAS 3 block the DNA-binding activity of STAT dimers. b | PIASX and PIASY might act as transcriptional co-repressors of STAT by recruiting other corepressor proteins such as histone deacetylase (HDAC). c | PIAS proteins can promote the conjugation of small ubiquitinrelated modifier (SUMO) to STAT 1. The significance of STAT 1 sumoylation in regulating STAT 1 activity is controversial and needs to be clarified Block DNA binding Act as corepressors Promote sumoylation 20 Nature Reviews Immunology 3; 900 -911
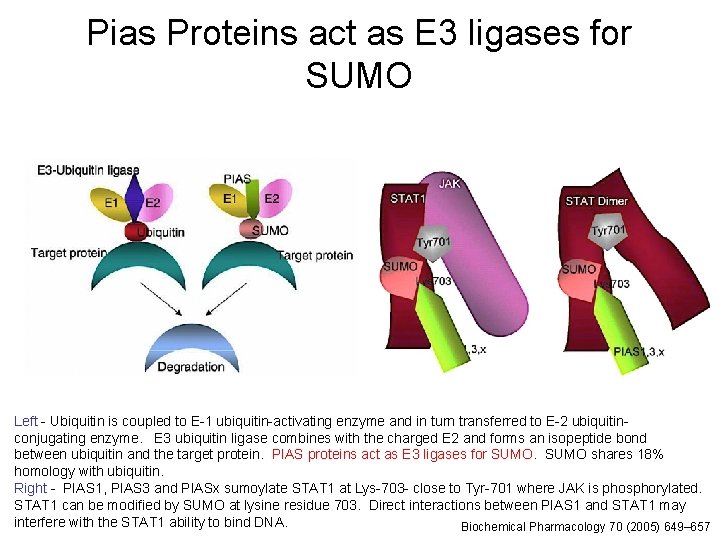
Pias Proteins act as E 3 ligases for SUMO Left - Ubiquitin is coupled to E-1 ubiquitin-activating enzyme and in turn transferred to E-2 ubiquitinconjugating enzyme. E 3 ubiquitin ligase combines with the charged E 2 and forms an isopeptide bond between ubiquitin and the target protein. PIAS proteins act as E 3 ligases for SUMO shares 18% homology with ubiquitin. Right - PIAS 1, PIAS 3 and PIASx sumoylate STAT 1 at Lys-703 - close to Tyr-701 where JAK is phosphorylated. 21 STAT 1 can be modified by SUMO at lysine residue 703. Direct interactions between PIAS 1 and STAT 1 may interfere with the STAT 1 ability to bind DNA. Biochemical Pharmacology 70 (2005) 649– 657

Example of Feedback Regulation by SOCS
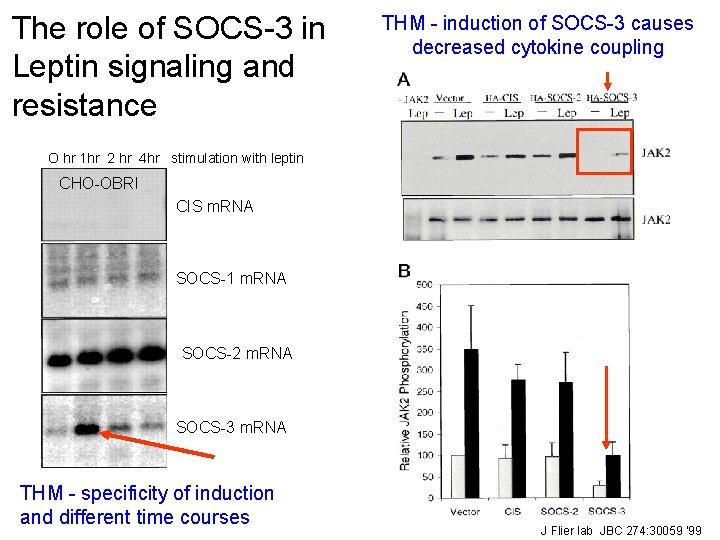
The role of SOCS-3 in Leptin signaling and resistance THM - induction of SOCS-3 causes decreased cytokine coupling O hr 1 hr 2 hr 4 hr stimulation with leptin CHO-OBRI CIS m. RNA SOCS-1 m. RNA SOCS-2 m. RNA SOCS-3 m. RNA Leptin induces SOCS-3, but not CIS, SOCS-1, THM - specificity of induction or SOCS-2, m. RNA in CHO cells expressing and different time courses the long form of the leptin receptor. 23 J Flier lab JBC 274: 30059 ‘ 99

How important are SSI proteins? Studies with SOCS 1 KO mice
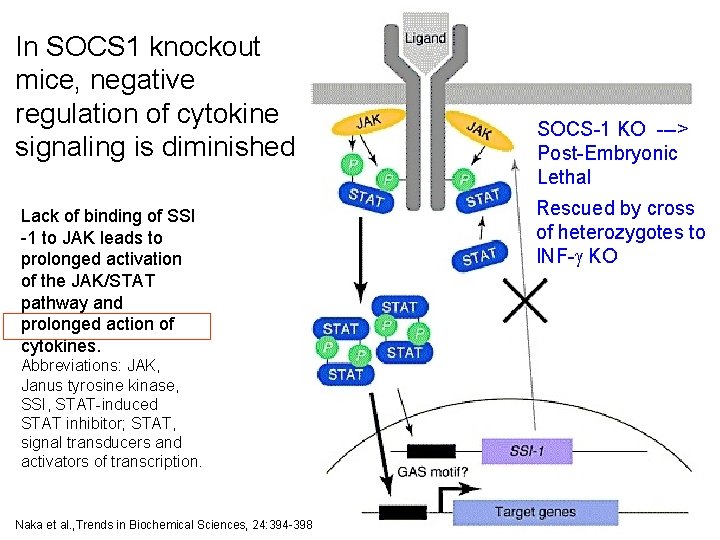
In SOCS 1 knockout mice, negative regulation of cytokine signaling is diminished Lack of binding of SSI -1 to JAK leads to prolonged activation of the JAK/STAT pathway and prolonged action of cytokines. SOCS-1 KO ---> Post-Embryonic Lethal Rescued by cross of heterozygotes to INF-g KO Abbreviations: JAK, Janus tyrosine kinase, SSI, STAT-induced STAT inhibitor; STAT, signal transducers and activators of transcription. 25 Naka et al. , Trends in Biochemical Sciences, 24: 394 -398

“Cross talk” between Jak/STAT and other signaling pathways THM 1: Other pathways can be activated by ligand binding to "cytokine" receptors THM 2: Jaks can bind & activate other tyrosine kinase pathways THM 3: Other serine kinase pathways can modulate Jak/STAT function THM 4: SOCS proteins can modulate other pathways
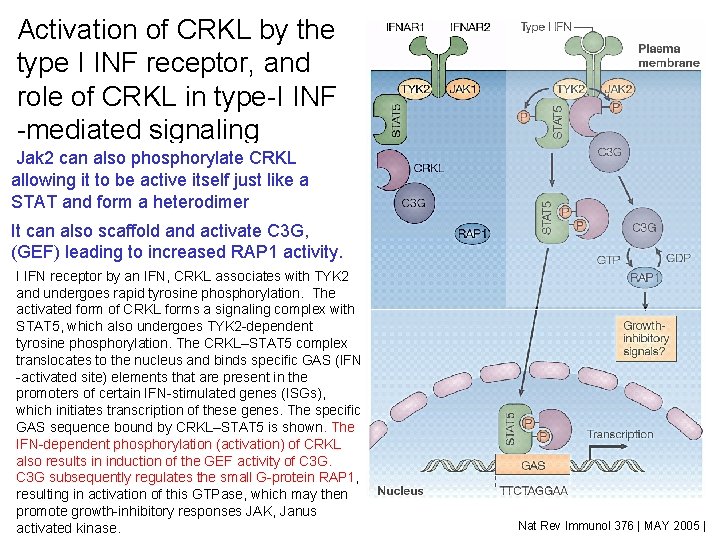
Activation of CRKL by the type I INF receptor, and role of CRKL in type-I INF -mediated signaling Fig 2 CRKL is present as a latent cytoplasmic form that Jak 2 can also phosphorylate CRKL constitutively associates with the guanine-nucleotideallowing it to be active itself just like a exchange factor (GEF) C 3 G. A member of the STAT and form a heterodimer (signal transducer and activator of transcription) family of proteins, STAT 5, is associated with tyrosine kinase 2 It can also scaffold and activate C 3 G, (TYK 2) that is bound to the type I interferon (IFN) (GEF) leading to increased RAP 1 activity. receptor subunit IFNAR 1. After engagement of the type I IFN receptor by an IFN, CRKL associates with TYK 2 and undergoes rapid tyrosine phosphorylation. The activated form of CRKL forms a signaling complex with STAT 5, which also undergoes TYK 2 -dependent tyrosine phosphorylation. The CRKL–STAT 5 complex translocates to the nucleus and binds specific GAS (IFN -activated site) elements that are present in the promoters of certain IFN-stimulated genes (ISGs), which initiates transcription of these genes. The specific GAS sequence bound by CRKL–STAT 5 is shown. The IFN-dependent phosphorylation (activation) of CRKL also results in induction of the GEF activity of C 3 G subsequently regulates the small G-protein RAP 1, resulting in activation of this GTPase, which may then promote growth-inhibitory responses JAK, Janus activated kinase. Nat Rev Immunol 376 | MAY 2005 |
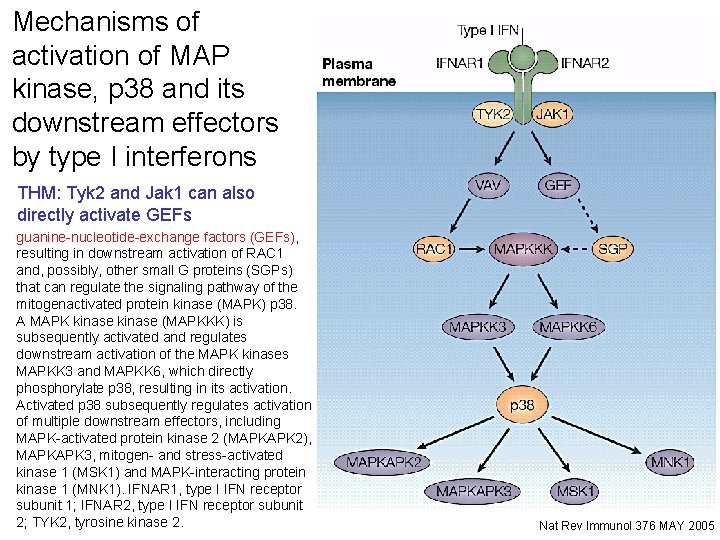
Mechanisms of activation of MAP kinase, p 38 and its downstream effectors by type I interferons THM: Tyk 2 and Jak 1 can also Interferon (IFN)-activated JAKs regulate the directly activate GEFs phosphorylation (activation) of VAV or other guanine-nucleotide-exchange factors (GEFs), resulting in downstream activation of RAC 1 and, possibly, other small G proteins (SGPs) that can regulate the signaling pathway of the mitogenactivated protein kinase (MAPK) p 38. A MAPK kinase (MAPKKK) is subsequently activated and regulates downstream activation of the MAPK kinases MAPKK 3 and MAPKK 6, which directly phosphorylate p 38, resulting in its activation. Activated p 38 subsequently regulates activation of multiple downstream effectors, including MAPK-activated protein kinase 2 (MAPKAPK 2), MAPKAPK 3, mitogen- and stress-activated kinase 1 (MSK 1) and MAPK-interacting protein kinase 1 (MNK 1). IFNAR 1, type I IFN receptor subunit 1; IFNAR 2, type I IFN receptor subunit 2; TYK 2, tyrosine kinase 2. 28 Nat Rev Immunol 376 MAY 2005
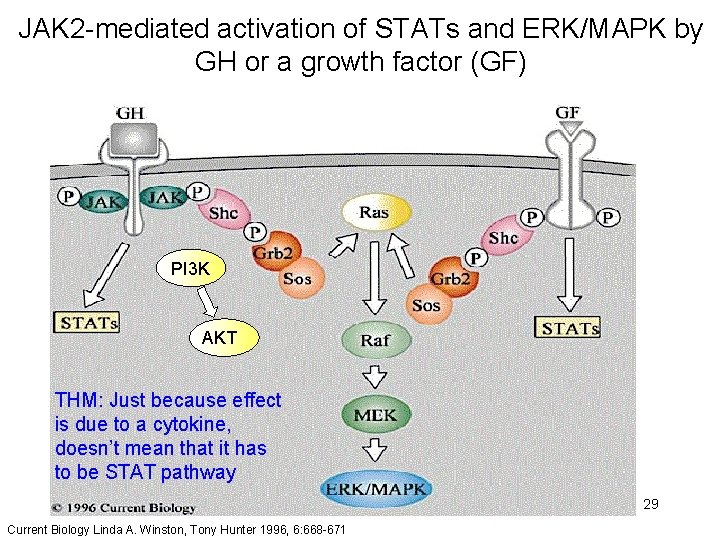
JAK 2 -mediated activation of STATs and ERK/MAPK by GH or a growth factor (GF) PI 3 K AKT THM: Just because effect is due to a cytokine, doesn’t mean that it has to be STAT pathway 29 Current Biology Linda A. Winston, Tony Hunter 1996, 6: 668 -671
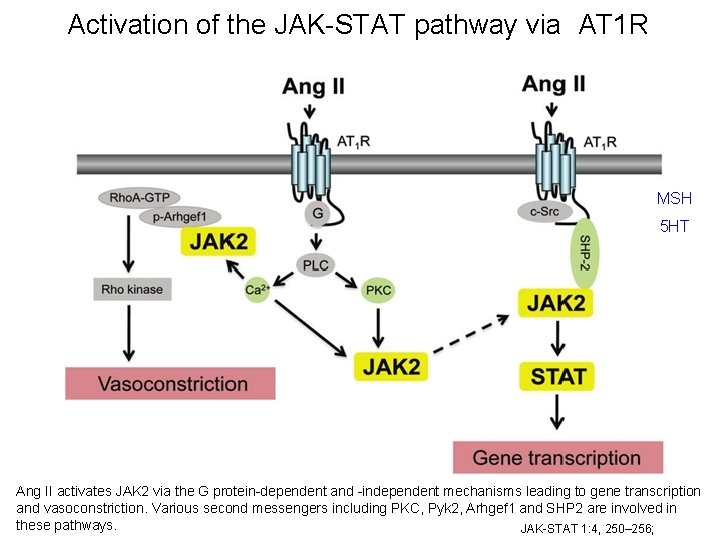
Activation of the JAK-STAT pathway via AT 1 R MSH 5 HT Ang II activates JAK 2 via the G protein-dependent and -independent mechanisms leading to gene transcription and vasoconstriction. Various second messengers including PKC, Pyk 2, Arhgef 1 and SHP 2 are involved in these pathways. JAK-STAT 1: 4, 250– 256;

Please - Don’t get behind on reading. You can be sure that some of exam questions will come from the readings. For example, in the research paper I assigned that came out just last week, Do you think that the authors are correct when they say that these inflamasomes that contain NLRP 3 are direct binders and effectors for c. AMP action on immune system? ? 31
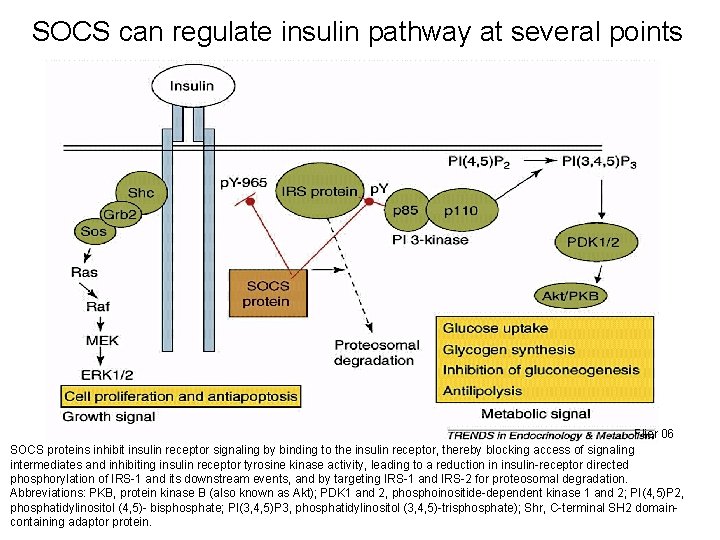
SOCS can regulate insulin pathway at several points Flier 06 SOCS proteins inhibit insulin receptor signaling by binding to the insulin receptor, thereby blocking access of signaling intermediates and inhibiting insulin receptor tyrosine kinase activity, leading to a reduction in insulin-receptor directed phosphorylation of IRS-1 and its downstream events, and by targeting IRS-1 and IRS-2 for proteosomal degradation. Abbreviations: PKB, protein kinase B (also known as Akt); PDK 1 and 2, phosphoinositide-dependent kinase 1 and 2; PI(4, 5)P 2, 32 phosphatidylinositol (4, 5)- bisphosphate; PI(3, 4, 5)P 3, phosphatidylinositol (3, 4, 5)-trisphosphate); Shr, C-terminal SH 2 domaincontaining adaptor protein.
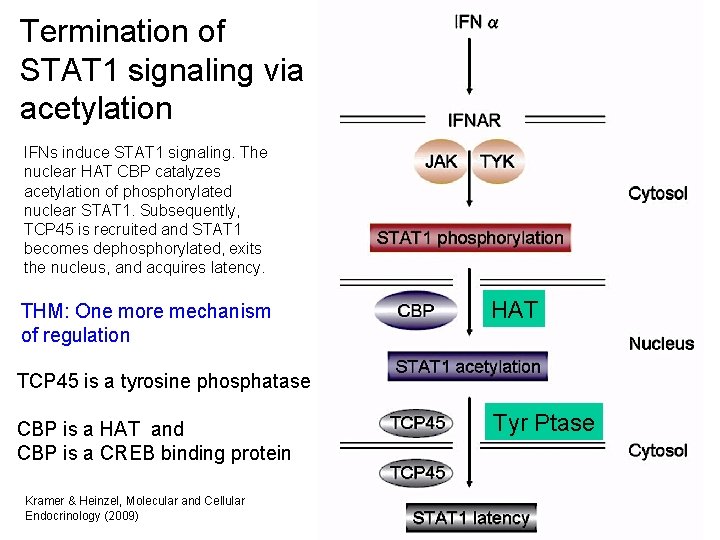
Termination of STAT 1 signaling via acetylation IFNs induce STAT 1 signaling. The nuclear HAT CBP catalyzes acetylation of phosphorylated nuclear STAT 1. Subsequently, TCP 45 is recruited and STAT 1 becomes dephosphorylated, exits the nucleus, and acquires latency. THM: One more mechanism of regulation HAT TCP 45 is a tyrosine phosphatase CBP is a HAT and CBP is a CREB binding protein Kramer & Heinzel, Molecular and Cellular Endocrinology (2009) Tyr Ptase 33
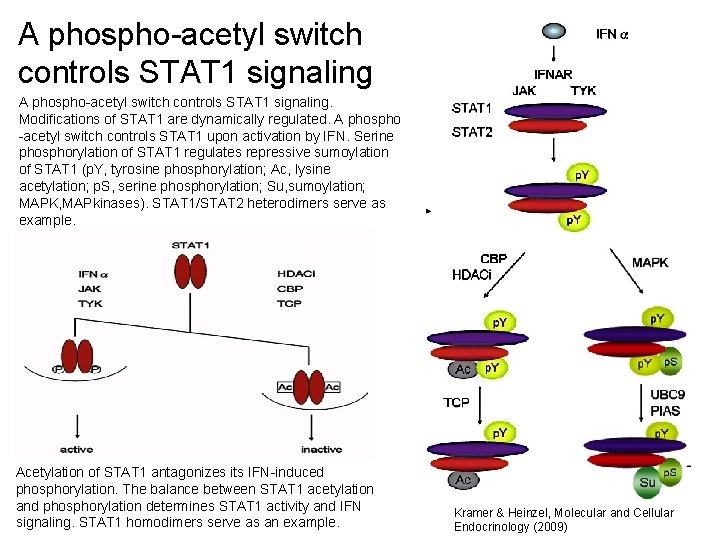
A phospho-acetyl switch controls STAT 1 signaling. Modifications of STAT 1 are dynamically regulated. A phospho -acetyl switch controls STAT 1 upon activation by IFN. Serine phosphorylation of STAT 1 regulates repressive sumoylation of STAT 1 (p. Y, tyrosine phosphorylation; Ac, lysine acetylation; p. S, serine phosphorylation; Su, sumoylation; MAPK, MAPkinases). STAT 1/STAT 2 heterodimers serve as example. Acetylation of STAT 1 antagonizes its IFN-induced phosphorylation. The balance between STAT 1 acetylation and phosphorylation determines STAT 1 activity and IFN signaling. STAT 1 homodimers serve as an example. 34 Kramer & Heinzel, Molecular and Cellular Endocrinology (2009)
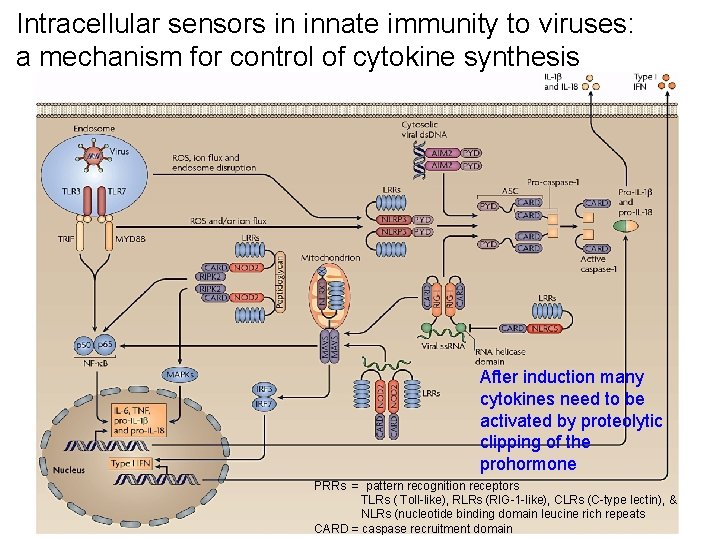
Intracellular sensors in innate immunity to viruses: a mechanism for control of cytokine synthesis After induction many cytokines need to be activated by proteolytic clipping of the prohormone PRRs = pattern recognition receptors TLRs ( Toll-like), RLRs (RIG-1 -like), CLRs (C-type lectin), & NLRs (nucleotide binding domain leucine rich repeats CARD = caspase recruitment domain

Figure 1 | Intracellular sensors in innate immunity to viruses. Viral pathogen-associated molecular patterns (PAMPs) activate nucleotide-binding oligomerization domain (NOD)-like receptors (NLRs) and inflammasomes to initiate signalling cascades that lead to the production of pro-inflammatory cytokines, thereby amplifying antiviral innate immune responses. In the presence of viral PAMPs, NLR family PYD-containing protein 3 (NLRP 3) and absent in melanoma 2 (AIM 2) oligomerize and recruit the adaptor protein apoptosis-associated speck-like protein containing a CARD (ASC) through homotypic pyrin domain (PYD) interactions. The caspase-recruitment domain (CARD) of ASC binds the CARD of pro-caspase‑ 1, leading to caspase‑ 1 activation and the production of interleukin‑ 1β (IL‑ 1β) and IL‑ 18 through cleavage of pro-IL‑ 1β and pro-IL‑ 18. Retinoic acid inducible gene‑I (RIG‑I) contains an RNA helicase domain and an amino-terminal CARD. The helicase domain of RIG‑I senses the 5ʹ-triphosphate moiety of single-stranded (ss)RNA virus genomes and then signals through CARD–CARD interactions with the adaptor molecule mitochondrial antiviral signalling protein (MAVS). This results in the phosphorylation and activation of interferon (IFN) response factor 3 (IRF 3) and IRF 7 to turn on the transcription of type I IFN (IFNα/β) genes. RIG‑I also regulates IL‑ 1β production transcriptionally and post-translationally following recognition of 5ʹ-triphosphate double-stranded (ds)RNA. Whereas RIG‑I-triggered transcription of pro-IL‑ 1β depends on nuclear factor-κB (NF-κB) activation and is mediated by MAVS, inflammasome formation, caspase‑ 1 activation, and IL‑ 1β and IL‑ 18 production in response to RIG‑I activation involve ASC. The NLRs NOD 2, NLR family member X 1 (NLRX 1) and NLR family CARD-containing protein 5 (NLRC 5) associate with MAVS. Whereas NOD 2 mediates the induction of type I IFNs, NLRX 1 and NLRC 5 inhibit RIG‑I–MAVS interactions and thereby negatively regulate type I IFN production. LRR, leucine-rich repeat; MAPK, mitogen-activated protein kinase; MYD 88, myeloid differentiation primary-response protein 88; RIPK 2, receptor-interacting serine-threonine protein kinase 2; ROS, reactive oxygen species; TLR, Toll-like receptor; TNF, tumour necrosis factor; 36 TRIF, TIR-domain-containing adaptor protein inducing IFNβ.
- Slides: 36