CONFORMATIONS OF CATIONIZED PEPTIDES DETERMINATION OF LIGAND BINDING

CONFORMATIONS OF CATIONIZED PEPTIDES DETERMINATION OF LIGAND BINDING GEOMETRIES BY IRMPD SPECTROSCOPY ROBERT C. DUNBAR Case Western Reserve University JEFFREY STEILL, JOS OOMENS FOM Institute for Plasma Physics NICK C. POLFER University of Florida $$$ FOM, NSF - PIRE

How Metal Ions Bind to Peptides Metal-ion complexation – Aqueous solution environment – peptide surface – Hydrophobic condensed phase – peptide interior – Gas-phase – solvent free binding

An Interesting Choice – CS versus SB • CS (Charge-Solvated, Canonical) – Peptide wrapped around metal ion • SB (Salt-Bridge, Zwitterion) – Separated + and − charges
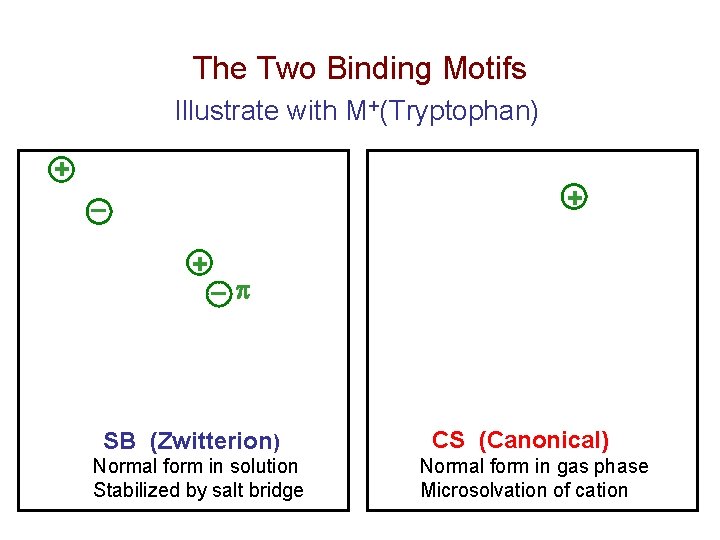
The Two Binding Motifs Illustrate with M+(Tryptophan) + + ¯ SB (Zwitterion) Normal form in solution Stabilized by salt bridge CS (Canonical) Normal form in gas phase Microsolvation of cation

Action Spectroscopy and IRMPD Infrared Multiple Photon Dissociation IR photon typically 0. 1 e. V Dissociation energy typically 3 e. V Many photons delivered by an intense, short laser pulse (IRMPD) M+Trp M+ + Trp Resonance with infrared absorptions gives peaks in IRMPD – Approximates IR spectrum
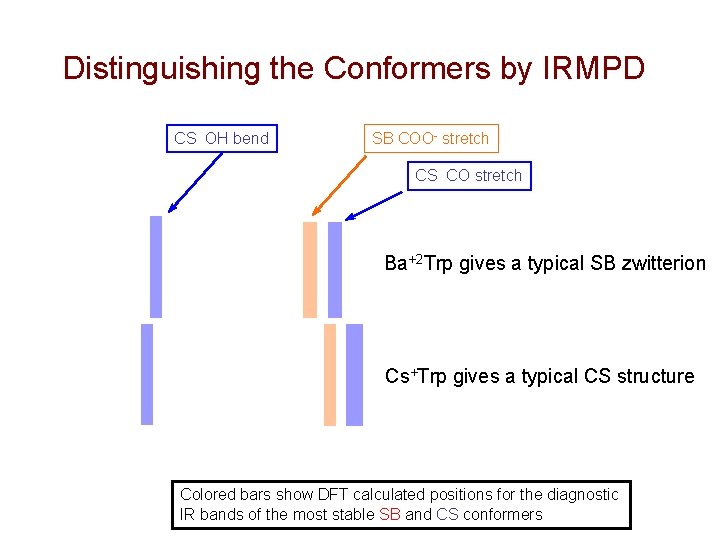
Distinguishing the Conformers by IRMPD CS OH bend SB COO- stretch CS CO stretch Ba+2 Trp gives a typical SB zwitterion Cs+Trp gives a typical CS structure Colored bars show DFT calculated positions for the diagnostic IR bands of the most stable SB and CS conformers

Light Source The Free Electron Laser (FELIX) gives v Convenient sweep across the chemically informative IR spectrum v High intensity and energy per pulse v Tight collimation of beam

Background: Mono-amino Acid Complexes • Complexes with M+1 cations usually CS • Higher charge, large ion size favors SB - Ba+2 complexes usually SB

Peptides • Going from amino acid to peptide: Longer chain favors CS by – Amide group coordination – More flexibility for chelating metal ion • Present study aimed at seeing how these effects play out in small peptide systems
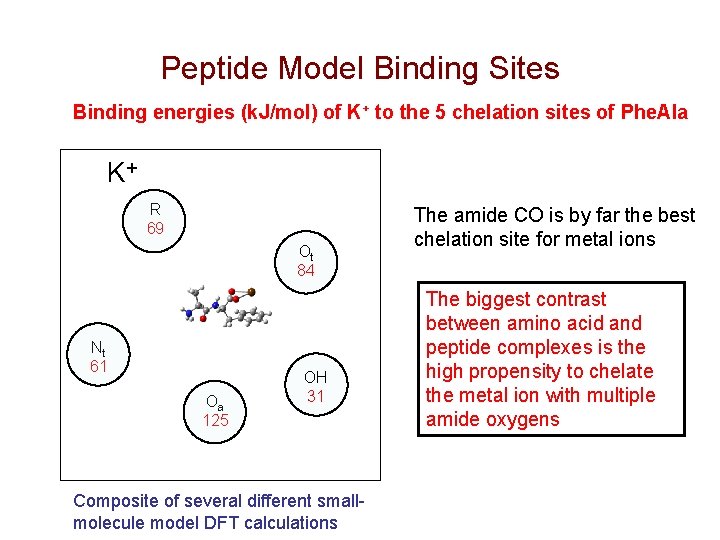
Peptide Model Binding Sites Binding energies (k. J/mol) of K+ to the 5 chelation sites of Phe. Ala K+ R 69 Ot 84 Nt 61 Oa 125 OH 31 Composite of several different smallmolecule model DFT calculations The amide CO is by far the best chelation site for metal ions The biggest contrast between amino acid and peptide complexes is the high propensity to chelate the metal ion with multiple amide oxygens
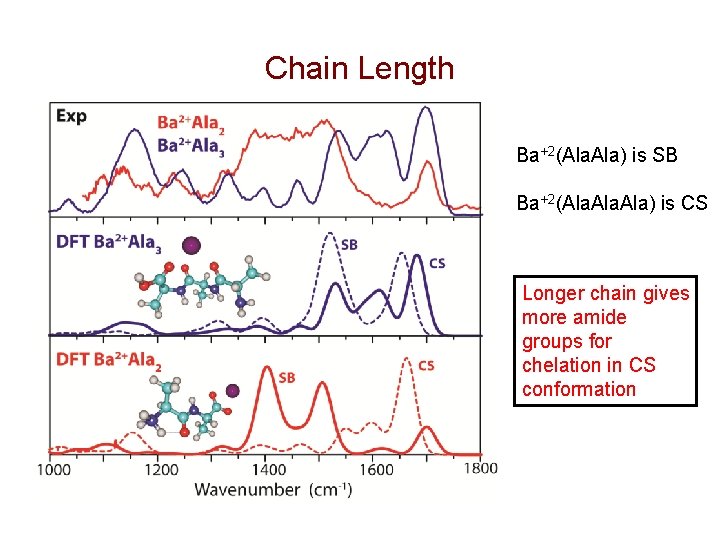
Chain Length Ba+2(Ala. Ala) is SB Ba+2(Ala. Ala) is CS Longer chain gives more amide groups for chelation in CS conformation
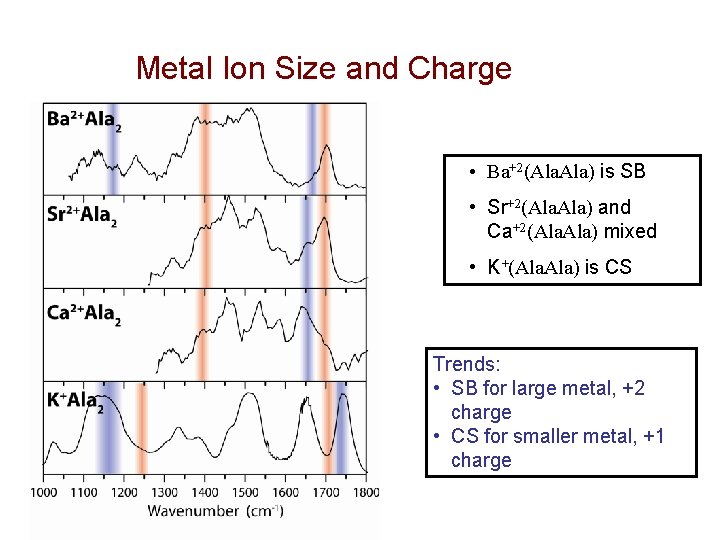
Metal Ion Size and Charge • Ba+2(Ala. Ala) is SB • Sr+2(Ala. Ala) and Ca+2(Ala. Ala) mixed • K+(Ala. Ala) is CS Trends: • SB for large metal, +2 charge • CS for smaller metal, +1 charge

Can peptide sequence influence SB stability? No encouragement from these alkalis: All CS conformation Polfer, Oomens, Dunbar, Chem. Phys. Chem 9, 579 (2008)
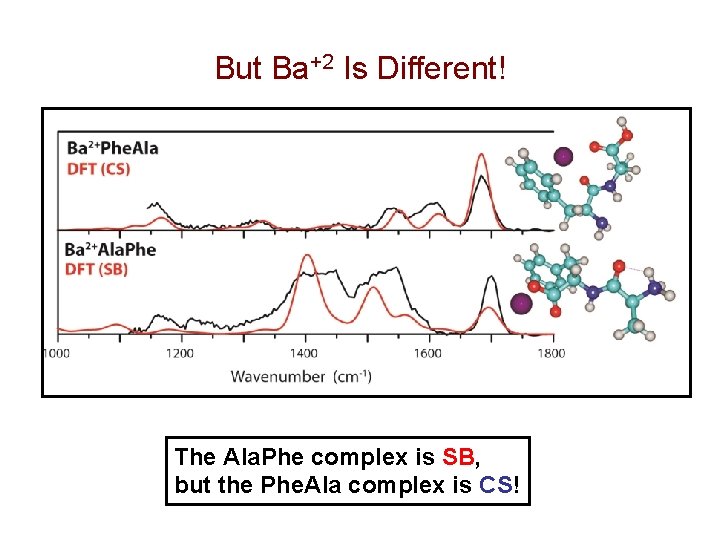
But Ba+2 Is Different! The Ala. Phe complex is SB, but the Phe. Ala complex is CS!

Conclusions • CS vs SB choice is closely balanced for small peptides, alkaline earth ion complexes • CS is favored by longer peptide chain (i. e. more amide chelation sites) • SB is favored by large metal ions, high metal charge • Sequence-dependent conformation switches correspond to varying side-chain chelation of the metal ion

From amino acid to protein Cyt-C from Oomens et al, PCCP, 2005


Metal-Ion Induced Peak Shifts All of these complexes are CS Shift of free CO and amide CO peaks gives a direct measure of the strength of the metaloxygen interaction The Na+(Ala)3 spectrum is taken from: Balaj, Kapota, Lemaire and Ohanessian, Int. J. Mass Spectrom. 269 (2008) 196.


Does the Phe/Ala order matter? With K+ the order makes almost no difference. With Ba+2 the order is crucial! What are these structures, and why are they so different? ?
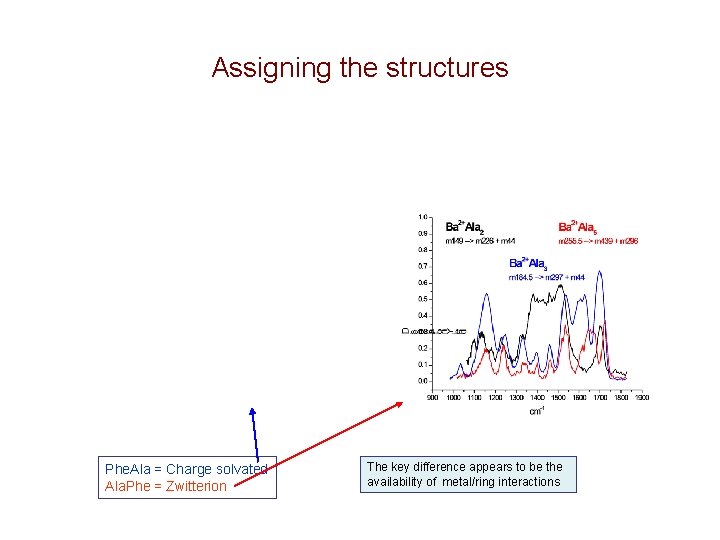
Assigning the structures Phe. Ala = Charge solvated Ala. Phe = Zwitterion The key difference appears to be the availability of metal/ring interactions

Proteins: Cytochrome C vs K+Ala. Phe The future: Maybe bigger and bigger biomolecules? OH Cyt-C from Oomens et al, PCCP, 2005

The Intensity Problem Conventional absorption spectroscopy Sample Photon detector Problem: Too little light absorbed by ions Heroic solution: cavity ringdown spectroscopy

The Infrared Photon Energy Problem The energy of an IR photon is typically 0. 1 e. V The energy of the dissociations of interest is typically 2 e. V Two solutions: v Rare gas tagging (supersonic jet very-cold-ion source) Ar+C 10 H 8 Ar + C 10 H 8+ v Many photons delivered by an intense, short laser pulse (IRMPD) C 10 H 8+ C 8 H 6 + + C 2 H 2 This is how we do it.

The alkali ion complexes of Ala 2 FELIX K+Ala 2 The K+ complex is very similar to the Na+ complex. Small blue shift of the C=O stretch due to weaker interaction with the metal ion. CLIO Na+Ala 2
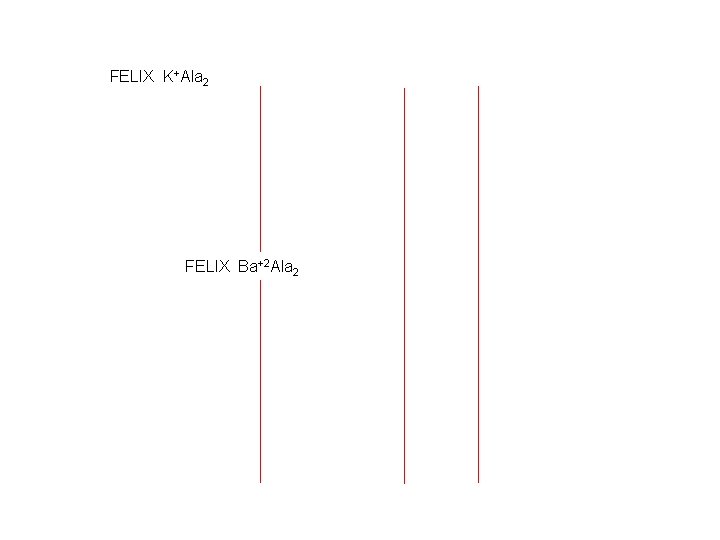
FELIX K+Ala 2 FELIX Ba+2 Ala 2

Ba+2 complexes of polyalanines

Conformation of the Cu. His. Tpy+2 complex Background: Siu, Hopkinson, Verkerk and others at York University (Canada) have worked out an efficient and promising synthetic route to amino acid radical cations. Histidine+ is an outstanding example. The procedure involves the collisional dissociation of a Cu(II) complex Cu. His. L+2, where L is an auxiliary ligand like terpyridine (Tpy). Cu. His. L+2 → His + + Cu. Tpy+. Features of the chemistry of the resulting His+ suggest a zwitterionic conformation of the His+. It was of interest to see if IRMPD spectroscopy can determine the conformation of this His ligand.

Conformation of the Cu. His. Tpy+2 complex The mixed dimer Cu. His. Tpy+2 can be rationalized well as a sum of one Tpy ligand one canonical His ligand. Presence of a zwitterionic His ligand is not suggested by the spectroscopy.
- Slides: 30