Computer simulations of proteins allatom and coarsegrained models












- Slides: 29

Computer simulations of proteins: all-atom and coarse-grained models Valeri Barsegov Department of Chemistry University of Massachusetts Lowell YITP, Kyoto University, Japan (2008)

Outline: I. Introduction: • single molecule spectroscopy of protein unfolding: biological relevance; pulling experiments (AFM, laser/optical tweezers, force protocols) single molecule spectroscopy of unbinding: biological relevance; experimental probes; resolution of forces, lifetimes, and extension • II. Molecular simulations of proteins: • • • proteins: structure, fold types, examples all-atom Molecular Dynamics (MD) simulations: force fields, examples, simulations of IR spectra coarse-grained description of proteins: approximations, examples III. New direction - computer simulations using graphics cards: • • basic facts, computer architecture, algorithms applications

I. 1 Single-molecule dynamic force spectroscopy of forced unfolding of proteins: biological relevance Fact 1: “mechanically active” proteins perform their biological function in linear tandems of “head-to-tail” (C-terminal-to-N-terminal) connected protein domains Examples: • Titin contains tandems of immunoglobulin (Ig) domains, separated by short linkers sequences (muscle function) • Actin-crosslinking filamins contain rod-like tandem of dd. FLN domains (cellular locomotion) • Fibronectin tandems consist of nonidentical Fn domains (extracellular matrix, cell elasticity) • Ubiquitin is a multimeric protein (Ub)n of n=9 identical Ub repeats (protein degradation, signaling pathways)

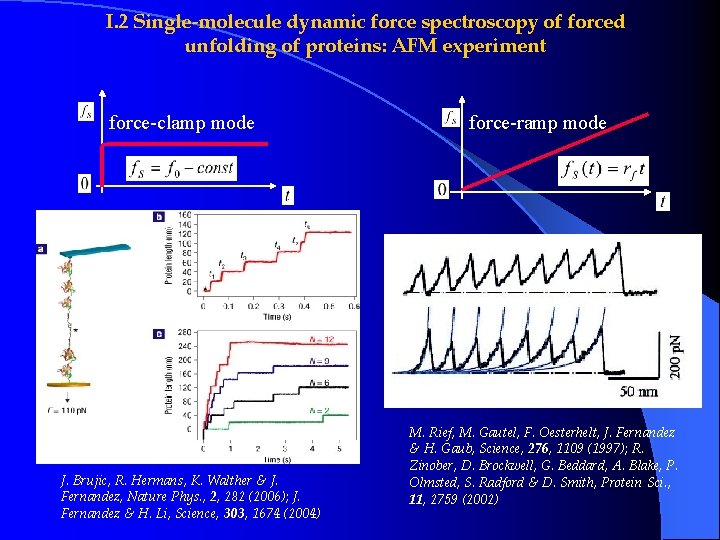
I. 2 Single-molecule dynamic force spectroscopy of forced unfolding of proteins: AFM experiment force-clamp mode J. Brujic, R. Hermans, K. Walther & J. Fernandez, Nature Phys. , 2, 282 (2006); J. Fernandez & H. Li, Science, 303, 1674 (2004) force-ramp mode M. Rief, M. Gautel, F. Oesterhelt, J. Fernandez & H. Gaub, Science, 276, 1109 (1997); R. Zinober, D. Brockwell, G. Beddard, A. Blake, P. Olmsted, S. Radford & D. Smith, Protein Sci. , 11, 2759 (2002)

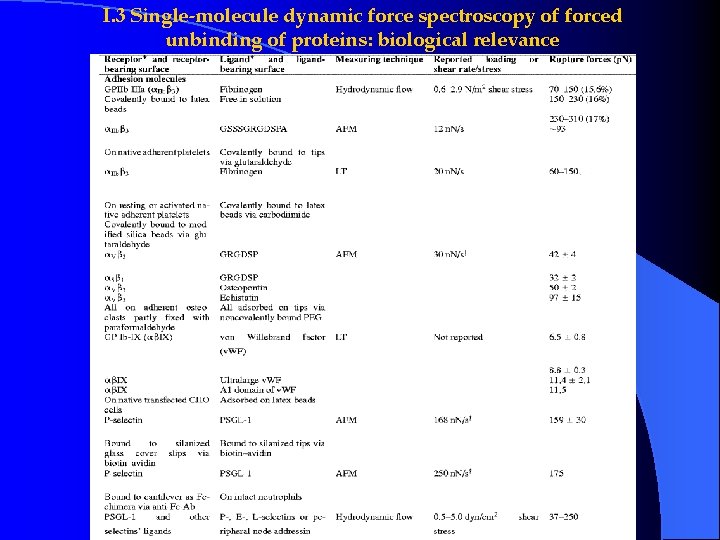
I. 3 Single-molecule dynamic force spectroscopy of forced unbinding of proteins: biological relevance

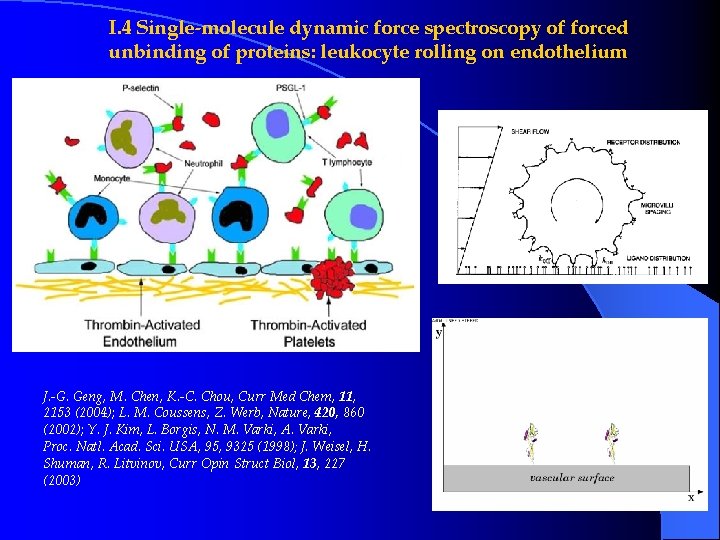
I. 4 Single-molecule dynamic force spectroscopy of forced unbinding of proteins: leukocyte rolling on endothelium J. -G. Geng, M. Chen, K. -C. Chou, Curr Med Chem, 11, 2153 (2004); L. M. Coussens, Z. Werb, Nature, 420, 860 (2002); Y. J. Kim, L. Borgis, N. M. Varki, A. Varki, Proc. Natl. Acad. Sci. USA, 95, 9325 (1998); J. Weisel, H. Shuman, R. Litvinov, Curr Opin Struct Biol, 13, 227 (2003)

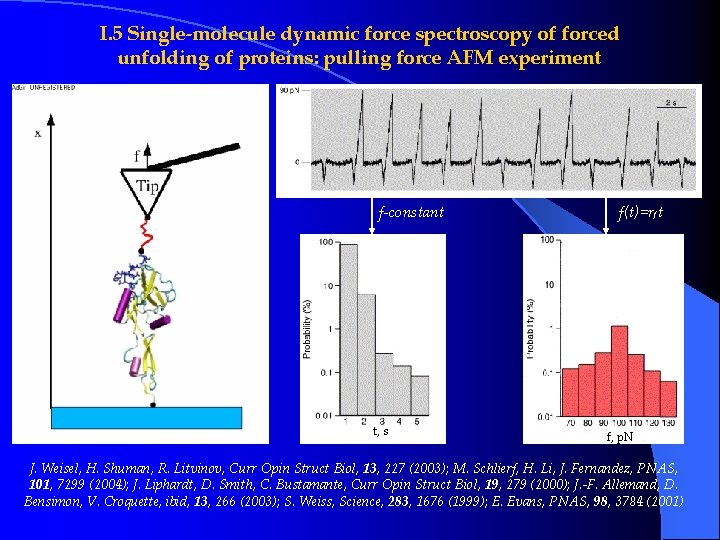
I. 5 Single-molecule dynamic force spectroscopy of forced unfolding of proteins: pulling force AFM experiment f-constant t, s f(t)=rf t f, p. N J. Weisel, H. Shuman, R. Litvinov, Curr Opin Struct Biol, 13, 227 (2003); M. Schlierf, H. Li, J. Fernandez, PNAS, 101, 7299 (2004); J. Liphardt, D. Smith, C. Bustamante, Curr Opin Struct Biol, 19, 279 (2000); J. -F. Allemand, D. Bensimon, V. Croquette, ibid, 13, 266 (2003); S. Weiss, Science, 283, 1676 (1999); E. Evans, PNAS, 98, 3784 (2001)

I. 6 Single-molecule dynamic force spectroscopy of proteins: experimental resolution of unfolding forces, times, and distances Experimental resolution: • protein extension X ~1 nm; • stretching force f. S 100 p. N • force-quench f. Q 5 -10 p. N • relaxation interval T 10 -100 s J. Fernandez & H. Li, Science, 303, 1674 (2004); I. Schwaiger, M. Schleicher, A. Noegel & M. Rief, EMBO Reports, 6, 46 (2005); J. Brujic, R. Hermans, K. Walther & J. Fernandez, Nature Phys. , 2, 282 (2006)

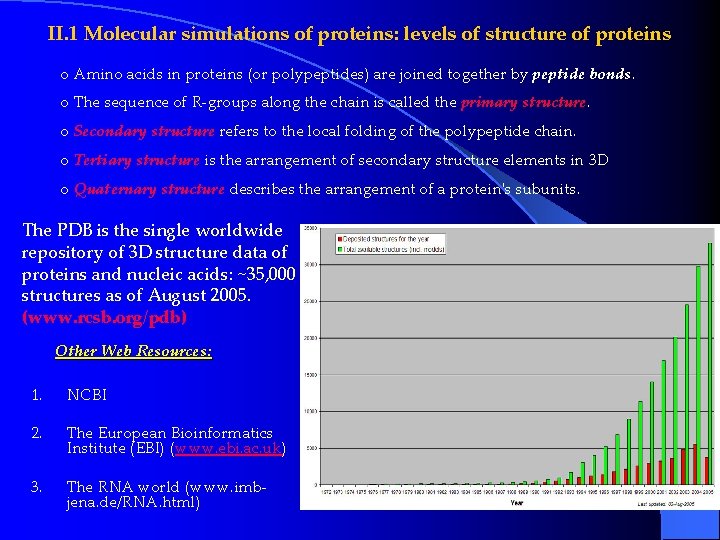
II. 1 Molecular simulations of proteins: levels of structure of proteins o Amino acids in proteins (or polypeptides) are joined together by peptide bonds. o The sequence of R-groups along the chain is called the primary structure. o Secondary structure refers to the local folding of the polypeptide chain. o Tertiary structure is the arrangement of secondary structure elements in 3 D o Quaternary structure describes the arrangement of a protein's subunits. The PDB is the single worldwide repository of 3 D structure data of proteins and nucleic acids: ~35, 000 structures as of August 2005. (www. rcsb. org/pdb) Other Web Resources: 1. NCBI 2. The European Bioinformatics Institute (EBI) (www. ebi. ac. uk) 3. The RNA world (www. imbjena. de/RNA. html)

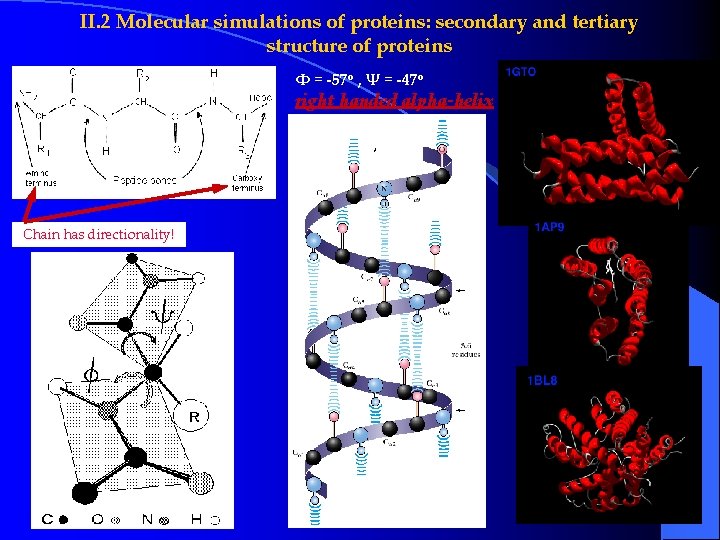
II. 2 Molecular simulations of proteins: secondary and tertiary structure of proteins Φ = -57 o , Ψ = -47 o right handed alpha-helix Chain has directionality!

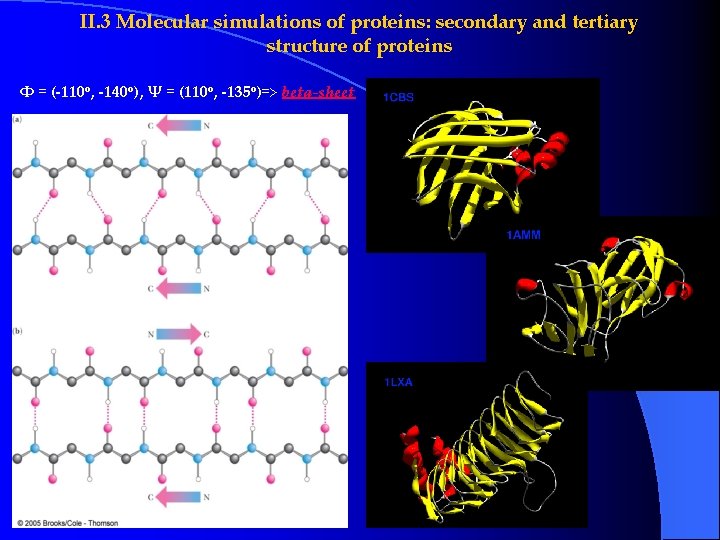
II. 3 Molecular simulations of proteins: secondary and tertiary structure of proteins Φ = (-110 o, -140 o), Ψ = (110 o, -135 o)=> beta-sheet

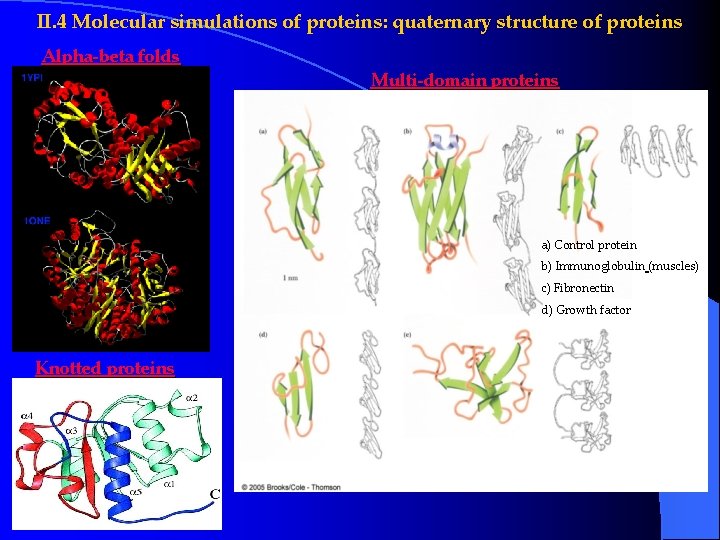
II. 4 Molecular simulations of proteins: quaternary structure of proteins Alpha-beta folds Multi-domain proteins a) Control protein b) Immunoglobulin (muscles) c) Fibronectin d) Growth factor Knotted proteins

II. 5 All-atom classical Molecular Dynamics (MD) simulations: force fields I. Potential for bonded interactions: VBL-bondlength potential, VBA-bond-angle potential, VDIH-dihedral angle potential, VSS – disulfide bond potential II. Potential for non-bonded interactions: VPP- protein-protein interaction potential, VWW- water-water potential, VWP - water-protein interaction potential III. Software (open-source): IV. Water models: • GROMACS (force field: OPLS and GROMOS ) • NAMD (force fields: CHARMM 22, CHARMM 27) • GROMACS (SPC, SPC/E, SPC-fw) • NAMD (TIP, TIP 3 P) GROMACS (Univ. of Groeningen, Netherlands): ftp: //ftp. gromacs. org/pub/ NAMD (Univ. of Illinois at Urbana Shampaign, USA): http: //www. ks. uiuc. edu/Research/namd/

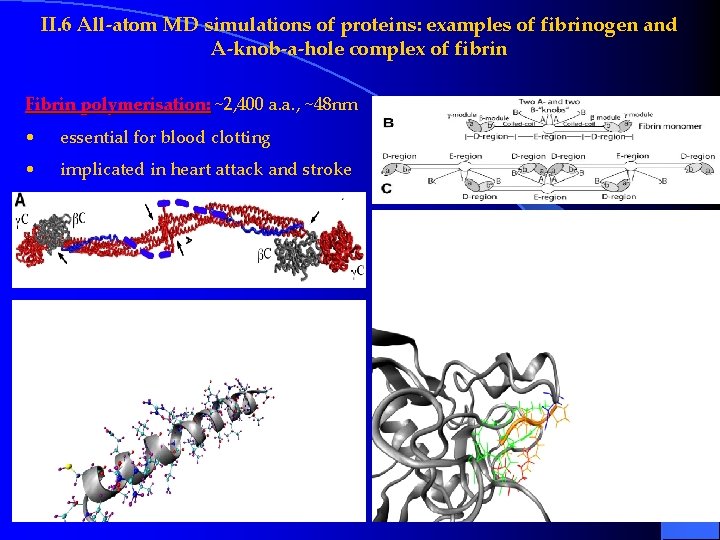
II. 6 All-atom MD simulations of proteins: examples of fibrinogen and A-knob-a-hole complex of fibrin Fibrin polymerisation: ~2, 400 a. a. , ~48 nm • essential for blood clotting • implicated in heart attack and stroke

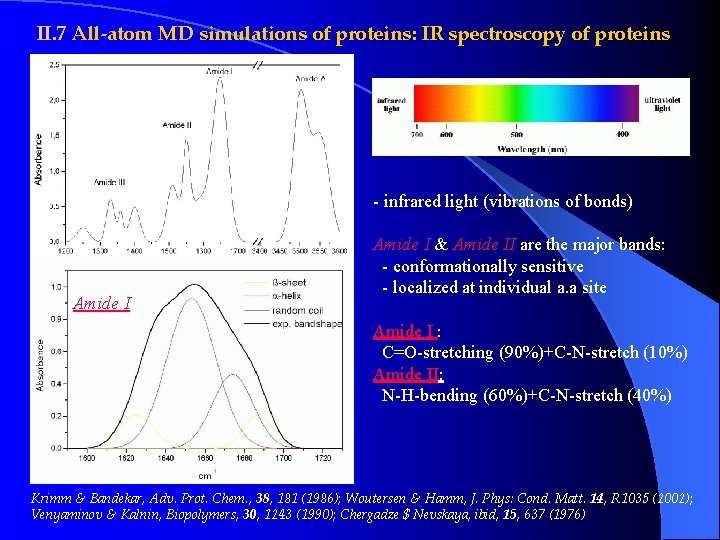
II. 7 All-atom MD simulations of proteins: IR spectroscopy of proteins - infrared light (vibrations of bonds) Amide I & Amide II are the major bands: - conformationally sensitive - localized at individual a. a site Amide I : C=O-stretching (90%)+C-N-stretch (10%) Amide II: N-H-bending (60%)+C-N-stretch (40%) Krimm & Bandekar, Adv. Prot. Chem. , 38, 181 (1986); Woutersen & Hamm, J. Phys: Cond. Matt. 14, R 1035 (2002); Venyaminov & Kalnin, Biopolymers, 30, 1243 (1990); Chergadze $ Nevskaya, ibid, 15, 637 (1976)

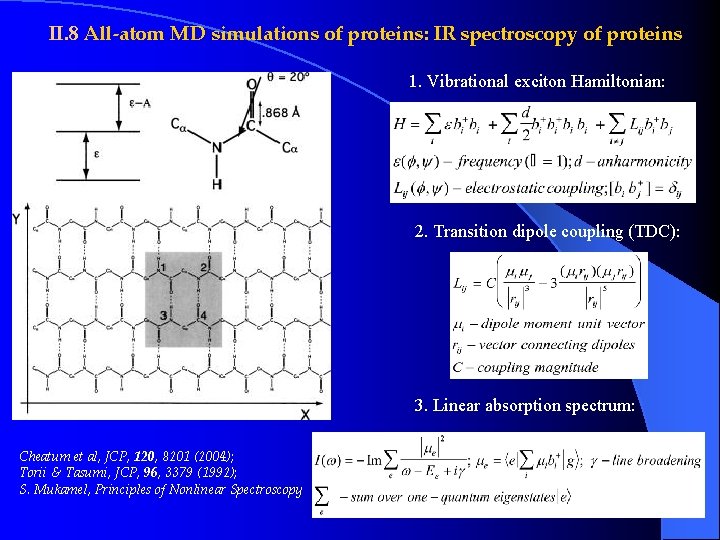
II. 8 All-atom MD simulations of proteins: IR spectroscopy of proteins 1. Vibrational exciton Hamiltonian: 2. Transition dipole coupling (TDC): 3. Linear absorption spectrum: Cheatum et al, JCP, 120, 8201 (2004); Torii & Tasumi, JCP, 96, 3379 (1992); S. Mukamel, Principles of Nonlinear Spectroscopy

II. 9 All-atom MD simulations of proteins: IR spectroscopy of proteins Assumptions used in the vibrational exciton Hamiltonian: - dynamics of in the near-equilibrium state - fast bath relaxation (fixed line broadening, ) - fitting parameters (diagonal energies, peak amplitudes, frequency splitting) - energies/amplitudes are from ab initio maps of N-methylacetamide, glycine dipeptide analogs - transferability of ab initio maps to larger proteins Direct calculation of IR spectra of Amide I from MD: - Amide I CO-vibration with - Correction Factor due to assumptions/harmonic force field Advantages of correlation functions: - IR obtained directly from classical MD - beyond ensemble average - far-from-equilibrium regime

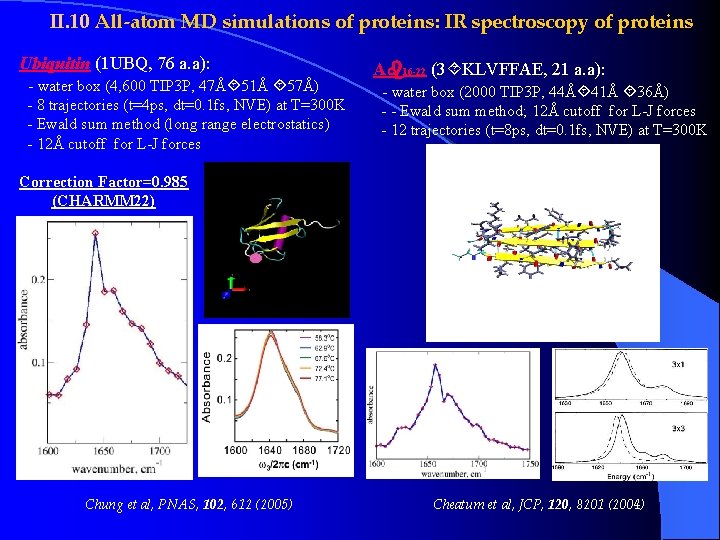
II. 10 All-atom MD simulations of proteins: IR spectroscopy of proteins Ubiquitin (1 UBQ, 76 a. a): - water box (4, 600 TIP 3 P, 47Å 51Å 57Å) - 8 trajectories (t=4 ps, dt=0. 1 fs, NVE) at T=300 K - Ewald sum method (long range electrostatics) - 12Å cutoff for L-J forces A 16 -22 (3 KLVFFAE, 21 a. a): - water box (2000 TIP 3 P, 44Å 41Å 36Å) - - Ewald sum method; 12Å cutoff for L-J forces - 12 trajectories (t=8 ps, dt=0. 1 fs, NVE) at T=300 K Correction Factor=0. 985 (CHARMM 22) Chung et al, PNAS, 102, 612 (2005) Cheatum et al, JCP, 120, 8201 (2004)

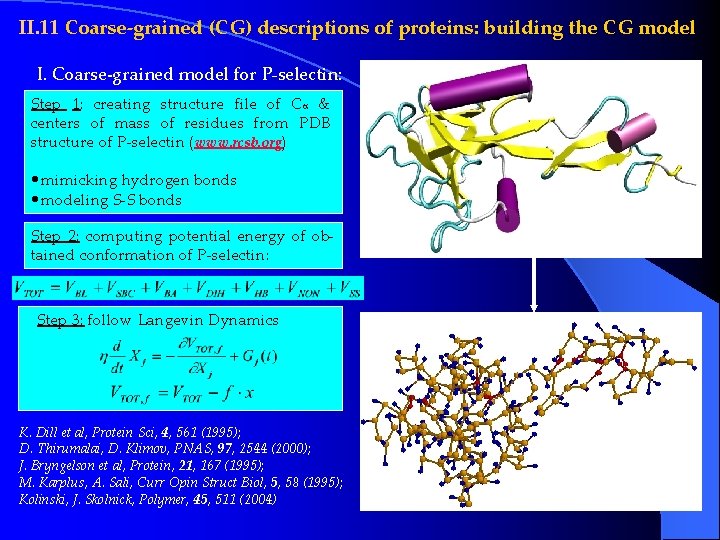
II. 11 Coarse-grained (CG) descriptions of proteins: building the CG model I. Coarse-grained model for P-selectin: Step 1: creating structure file of Ca & centers of mass of residues from PDB structure of P-selectin (www. rcsb. org) • mimicking hydrogen bonds • modeling S-S bonds Step 2: computing potential energy of obtained conformation of P-selectin: Step 3: follow Langevin Dynamics K. Dill et al, Protein Sci, 4, 561 (1995); D. Thirumalai, D. Klimov, PNAS, 97, 2544 (2000); J. Bryngelson et al, Protein, 21, 167 (1995); M. Karplus, A. Sali, Curr Opin Struct Biol, 5, 58 (1995); Kolinski, J. Skolnick, Polymer, 45, 511 (2004)

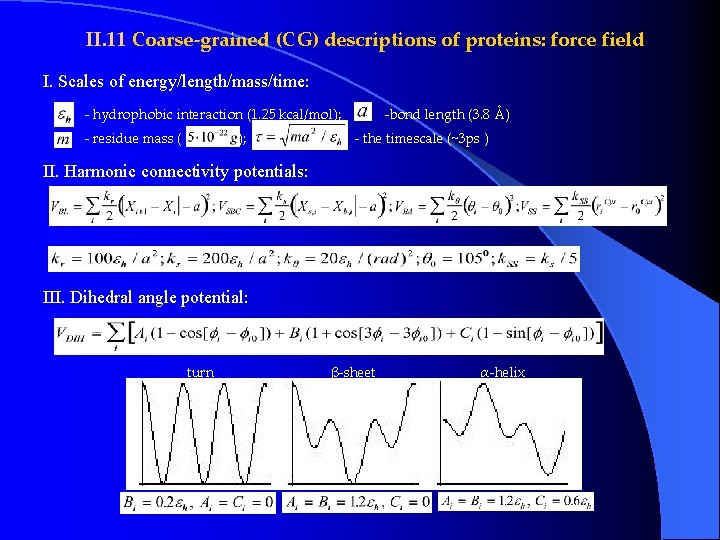
II. 11 Coarse-grained (CG) descriptions of proteins: force field I. Scales of energy/length/mass/time: - hydrophobic interaction (1. 25 kcal/mol); - residue mass ( ); -bond length (3. 8 Å) - the timescale (~3 ps ) II. Harmonic connectivity potentials: III. Dihedral angle potential: turn β-sheet α-helix

II. 11 Coarse-grained (CG) descriptions of proteins: force field IV. Hydrogen bond potential: V. Potential for native contacts: bij-contact interaction matrix; -contact distance (Kolinski et al, JCP, 98, 7420 (1993)) VI. Nonbonded potential: VII. Unfolding/unbinding trajectories:

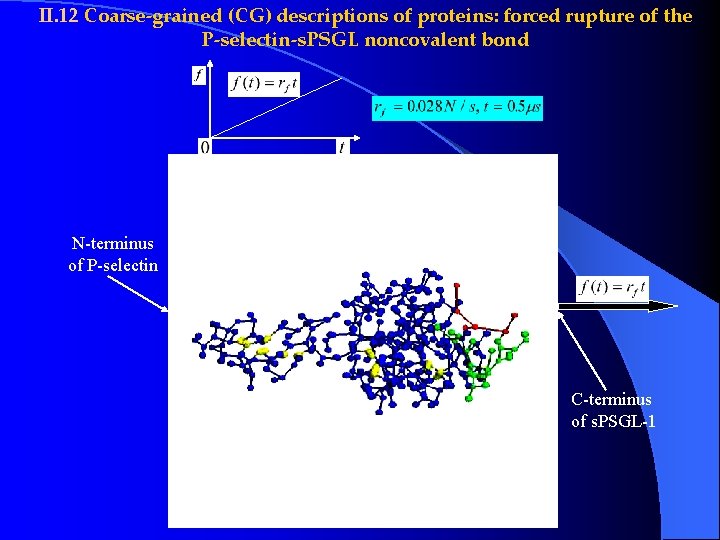
II. 12 Coarse-grained (CG) descriptions of proteins: forced rupture of the P-selectin-s. PSGL noncovalent bond N-terminus of P-selectin C-terminus of s. PSGL-1

III. 1 Computer simulations using graphics cards: basic facts CPU: Advantages: • can perform very sophisticated flow control (IF/THEN – cycles, conditionals, etc. ) • single CPU cores are faster (3. 0 GHz) or faster • a lot of well-tested (commercial) software is available Disadvantages: • has no more than 6 cores (today) • parallel programming on CPU is difficult • data exchange b/w nodes in a cluster occurs through relatively slow network GPU: Advantages: • up to 240 cores (Ge. Force 280, Tesla C 1060) • easy to write parallel codes with CUDA language (extension of C) • memory bandwidth is high because all cores are local Disadvantages: • single core clock is not as fast as CPU core (0. 5 GHz) • can’t be used for applications with sophisticated flow control • not many software available for GPU (started in ~2006)

III. 2 Computer simulations using graphics cards: hardware GPU: • highly parallel • multythreaded • manycore processor Historically, GPU • was designed for compute-intensive, highly parallel computation • more transistors are devoted to data processing rather than data caching and flow control • well-suited for problems that involve data-parallel computations, i. e. the same program is executed on many data elements in parallel (MD, coarse-grained simulations).

III. 3 Computer simulations using graphics cards: programming mode CUDA: • consist of a minimal extension to the C language • parallel programming model and software environment • designed to overcome the challenge of creating software that transparently scales on manycore processors Example: • vec. Add() function is called N times on GPU. • <<<1, N>>> means that the procedure runs in one 1 D block with N threads. • i = thread. Idx. x is a way for thread to identify, which element of the vector it should work with.

III. 4 Computer simulations using graphics cards: software organization Thread hierarchy: 1. thread index is a 3 D vector, so that threads can be identified using a 1 D, 2 D or 3 D index forming 1 D, 2 D or 3 D thread block 2. multiple blocks can be organized into 1 D or 2 D grid. Each block can be identified within grid using 1 D, 2 D or 3 D block index 3. all threads in one block are doing the same thing with different data 4. threads can synchronize and pass data to each other within block using shared memory 5. threads can pass the data to the CPU through the GPU global memory

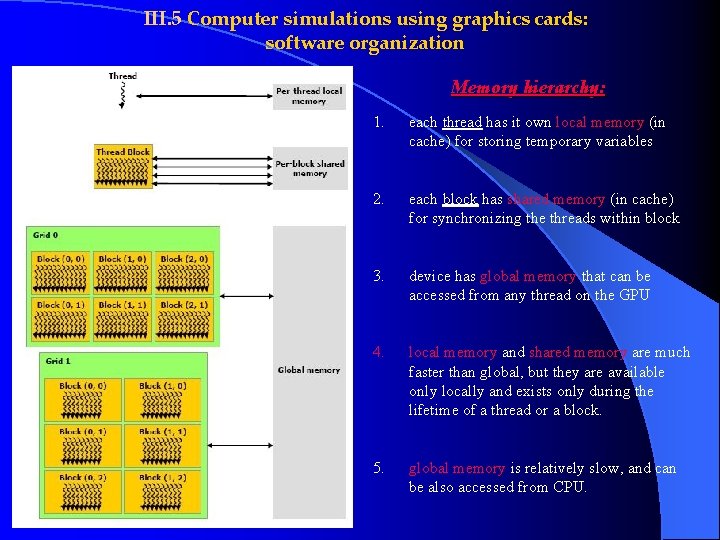
III. 5 Computer simulations using graphics cards: software organization Memory hierarchy: 1. each thread has it own local memory (in cache) for storing temporary variables 2. each block has shared memory (in cache) for synchronizing the threads within block 3. device has global memory that can be accessed from any thread on the GPU 4. local memory and shared memory are much faster than global, but they are available only locally and exists only during the lifetime of a thread or a block. 5. global memory is relatively slow, and can be also accessed from CPU.

III. 6 Computer simulations using graphics cards: hardware model Hardware organization: 1. each device have N multiprocessors 2. multiprocessors can share data only through device (global) memory 3. A multiprocessor have M processors (ALUs) 4. number of threads that can run at the same time is equal to Nx. M; for Ge. Force 8800 GT, M=8, N=14 (number of processors = 112!!!) 5. one block can run only on one multiprocessor so the number of blocks in program should be at least equal to the number of multiprocessors on the device.

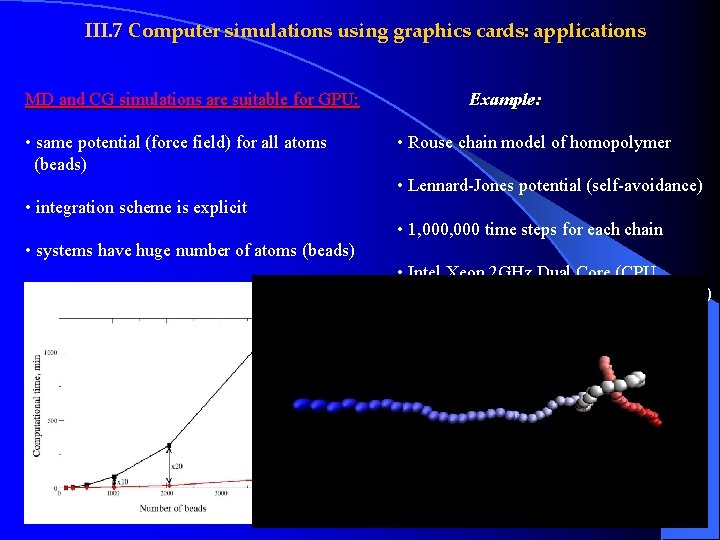
III. 7 Computer simulations using graphics cards: applications MD and CG simulations are suitable for GPU: • same potential (force field) for all atoms (beads) Example: • Rouse chain model of homopolymer • Lennard-Jones potential (self-avoidance) • integration scheme is explicit • 1, 000 time steps for each chain • systems have huge number of atoms (beads) • Intel Xeon 2 GHz Dual Core (CPU, ~$350) vs Ge. Force 8800 GT (GPU, ~$130)