Computational StructureBased Redesign of Enzyme Activity ChengYu Chen

Computational Structure-Based Redesign of Enzyme Activity Cheng-Yu Chen, Ivelin Georgiev, Amy C. Anderson, Bruce R. Donald A Different computational redesign strategy Yizhou Yin Mar 06, 2009

- Protein design: straightforward design vs. Directed mutation De Novo vs. redesign - Computational structure-based redesign GMEC (global minimum energy confirmation) - ROSETTA (Rosetta. Design, …) 1) Energy Function 2) Conformational Sampling
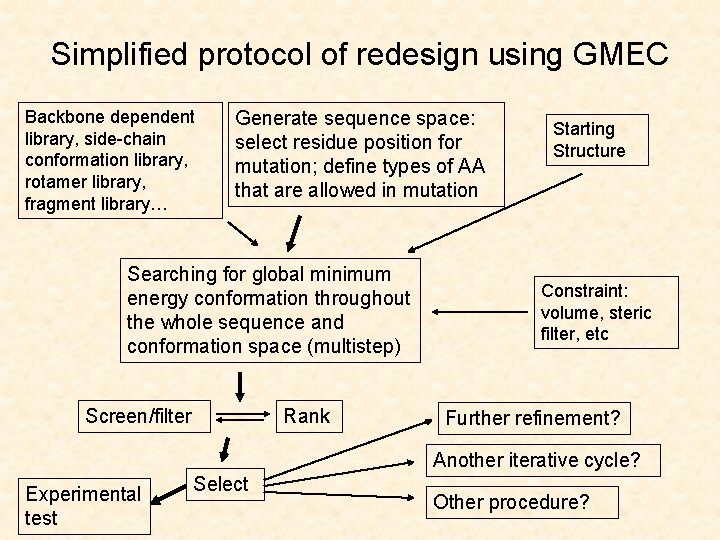
Simplified protocol of redesign using GMEC Backbone dependent library, side-chain conformation library, rotamer library, fragment library… Generate sequence space: select residue position for mutation; define types of AA that are allowed in mutation Searching for global minimum energy conformation throughout the whole sequence and conformation space (multistep) Screen/filter Rank Starting Structure Constraint: volume, steric filter, etc Further refinement? Another iterative cycle? Experimental test Select Other procedure?

Ensemble-based protein redesign Backbone dependent library, side-chain conformation library, rotamer library, fragment library… Generate sequence space: select residue position for mutation (steric shell); define types of AA that are allowed in mutation Starting Structure, targeted substrate, cofactor Filters: sequence-space filter, k-point, volume filter Active site mutation K* algorithm: search and score Rank + Select Multiple pruning methods Experimental verification Self-Consistent Mean Field entropy-based method Bolstering Mutation Min. DEE/A* algorithm: search and score Experimental verification

K* algorithm - For a given protein-substrate complex, K* computes a provably-accurate ε-approximation to the binding constant KA - K*= [Σexp(-Eb/RT)] / [Σexp(-El/RT)·Σexp(-Ef/RT)] b∈B l∈L f∈F B, L, F are rotamer-based ensembles; E is the conformation energy - Several algorithms are used to prune the candidate sequences at different steps so that the searching in the sequence space will be more efficient.

For each allowable mutated sequence: Step 1 Molecular ensemble is generated, then pruned by steric, volume filters. Step 2 After constrained energy minimization, the conformation is enumerated by A*. Step 3 The scores from step 2 are used to compute there separate partition functions, which is then combined to

SCMF entropy-based method Si = - ∑p(a︱i) ln p(a︱i) a∈Ai p(a︱i) = ∑ p(r︱i) r∈Ra - Ai is the set of AA types allowed at position i; p(a) is the probability of having AA type a at i. Ra is the set of rotamers for AA type a and p(r) is the probability of having rotamer r for AA type a at i. - Higher entropy implies higher probability of multiple AA types, hence higher tolerance to mutation at position i.

Example of Grs. A-Phe. A’s specificity switched from Phe to Leu - Grs. A-Phe. A is the phenylalanine adenylation domain of the nonribosomal peptide synthetase (NRPS) enzyme gramicidin S synthetase A, whose cognate substrate is Phe.

-7 residues at the active site are allowed to mutate to (G, A, V, L, I, W, F, Y, M) -only sequences with up to two mutations were considered, give the number candidates: 1450 (6. 44 x 10<7>) -After pruning, the number of sequences evaluated by K*: 505 (1. 12 x 10<7>) -Top ten sequences were experimentally verified. -7 residues were selected by SCMF and were allowed to mutate to different subset of AA. -Up to 3 -point mutations were considered.
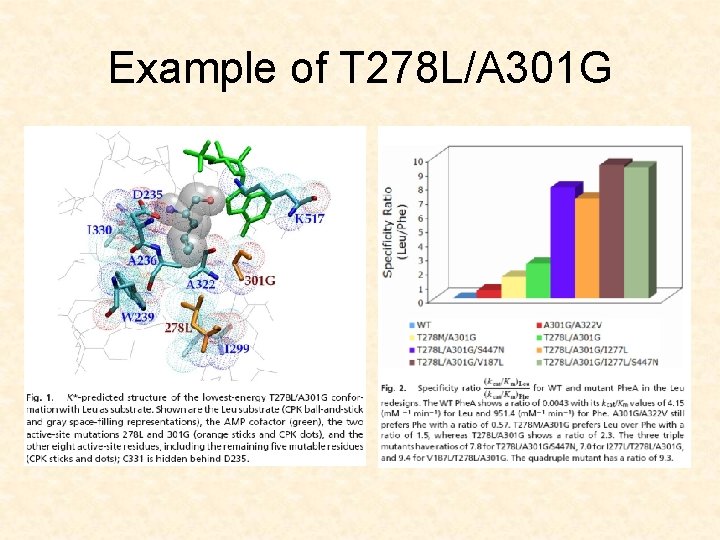
Example of T 278 L/A 301 G
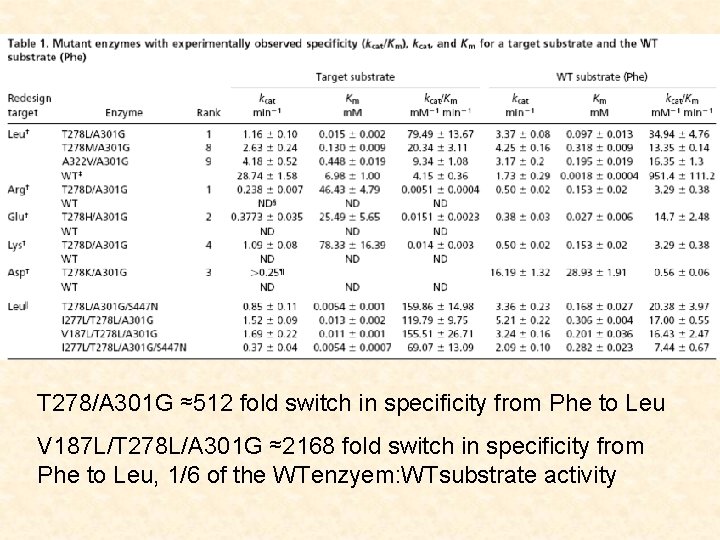
T 278/A 301 G ≈512 fold switch in specificity from Phe to Leu V 187 L/T 278 L/A 301 G ≈2168 fold switch in specificity from Phe to Leu, 1/6 of the WTenzyem: WTsubstrate activity

Comparison in efficiency, accuracy ensemble based vs. non-ensemble based 1) searching for best conformation 2) Searching for best mutation with best conformation 3) Other redesign 4) Other than redesign 5) structure-based design vs. other computational design/ evolution

- Will there be any better “hybrid” methods? - How to appropriately decide the sampling size based on the redesign methods? - Any other new strategy?
- Slides: 13