Computational Molecular Biology Protein Structure Introduction and Prediction

Computational Molecular Biology Protein Structure: Introduction and Prediction

Protein Folding Ø One of the most important problem in molecular biology Ø Given the one-dimensional amino-acid sequence that specifies the protein, what is the protein’s fold in three dimensions? My T. Thai mythai@cise. ufl. edu 2

Overview Ø Understand protein structures § Primary, secondary, tertiary Ø Why study protein folding: q Structure can reveal functional information which we cannot find from the sequence q Misfolding proteins can cause diseases: mad cow disease q Use in drug designs My T. Thai mythai@cise. ufl. edu 3

Overview of Protein Structure Ø Proteins make up about 50% of the mass of the average human Ø Play a vital role in keeping our bodies functioning properly Ø Biopolymers made up of amino acids Ø The order of the amino acids in a protein and the properties of their side chains determine three dimensional structure and function of the protein My T. Thai mythai@cise. ufl. edu 4
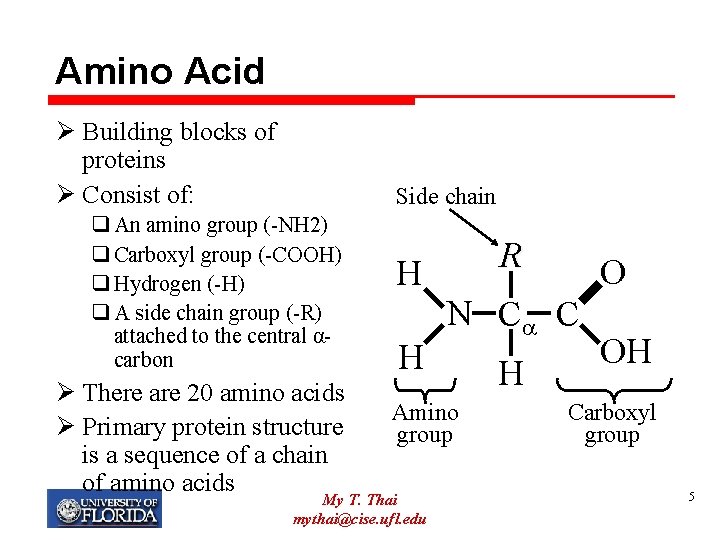
Amino Acid Ø Building blocks of proteins Ø Consist of: Side chain q An amino group (-NH 2) q Carboxyl group (-COOH) q Hydrogen (-H) q A side chain group (-R) attached to the central αcarbon Ø There are 20 amino acids Ø Primary protein structure is a sequence of a chain of amino acids R H H N C C Amino group My T. Thai mythai@cise. ufl. edu O H OH Carboxyl group 5

Side chains (Amino Acids) Ø 20 amino acids have side chains that vary in structure, size, hydrogen bonding ability, and charge. Ø R gives the amino acid its identity Ø R can be simple as hydrogen (glycine) or more complex such as an aromatic ring (tryptophan) My T. Thai mythai@cise. ufl. edu 6
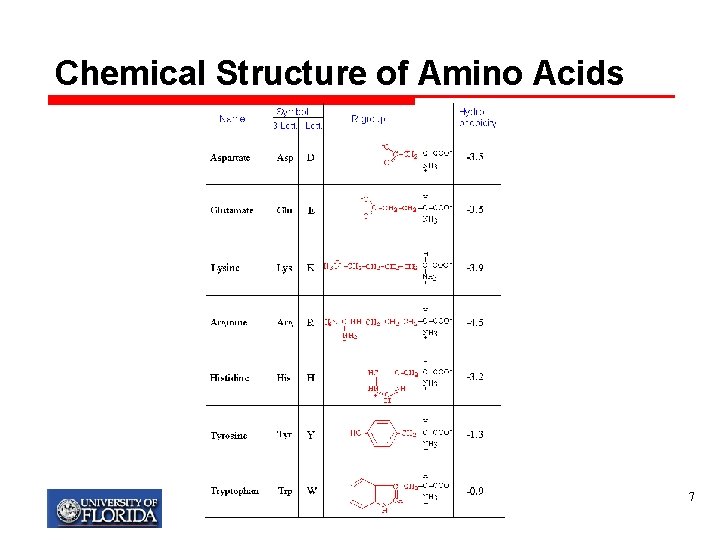
Chemical Structure of Amino Acids 7

How Amino Acids Become Proteins Peptide bonds My T. Thai mythai@cise. ufl. edu 8

Polypeptide Ø More than fifty amino acids in a chain are called a polypeptide. Ø A protein is usually composed of 50 to 400+ amino acids. Ø We call the units of a protein amino acid residues. amide nitrogen carbonyl carbon My T. Thai mythai@cise. ufl. edu 9

Side chain properties Ø Carbon does not make hydrogen bonds with water easily – hydrophobic. q These ‘water fearing’ side chains tend to sequester themselves in the interior of the protein Ø O and N are generally more likely than C to h-bond to water – hydrophilic q Ten to turn outward to the exterior of the protein My T. Thai mythai@cise. ufl. edu 10
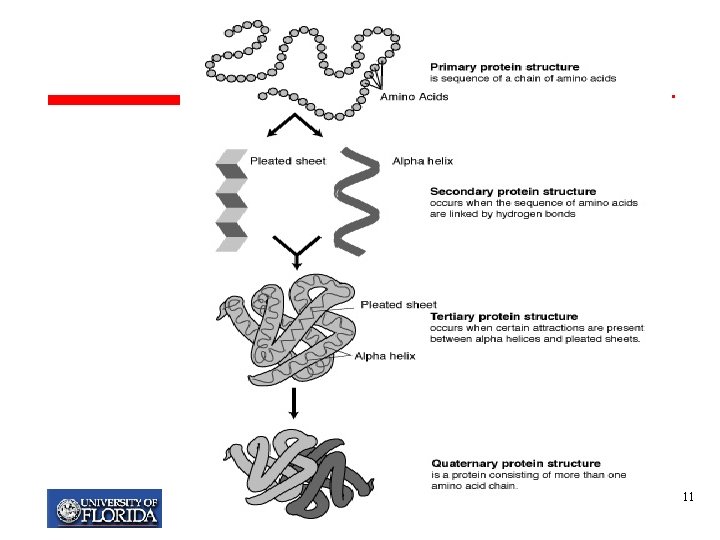
My T. Thai mythai@cise. ufl. edu 11

Primary Structure Primary structure: Linear String of Amino Acids Side-chain Backbone. . . ALA PHE LEU ILE LEU ARG. . . Each amino acid within a protein is referred to as residues Each different protein has a unique sequence of amino acid residues, this is its primary structure My T. Thai mythai@cise. ufl. edu 12

Secondary Structure Ø Refers to the spatial arrangement of contiguous amino acid residues Ø Regularly repeating local structures stabilized by hydrogen bonds q A hydrogen atom attached to a relatively electronegative atom Ø Examples of secondary structure are the α–helix and β–pleated-sheet My T. Thai mythai@cise. ufl. edu 13

Alpha-Helix Ø Amino acids adopt the form of a right handed spiral Ø The polypeptide backbone forms the inner part of the spiral Ø The side chains project outward Ø every backbone N-H group donates a hydrogen bond to the backbone C = O group My T. Thai mythai@cise. ufl. edu 14

Beta-Pleated-Sheet Ø Consists of long polypeptide chains called beta-strands, aligned adjacent to each other in parallel or anti-parallel orientation Ø Hydrogen bonding between the strands keeps them together, forming the sheet Ø Hydrogen bonding occurs between amino and carboxyl groups of different strands My T. Thai mythai@cise. ufl. edu 15

Parallel Beta Sheets My T. Thai mythai@cise. ufl. edu 16
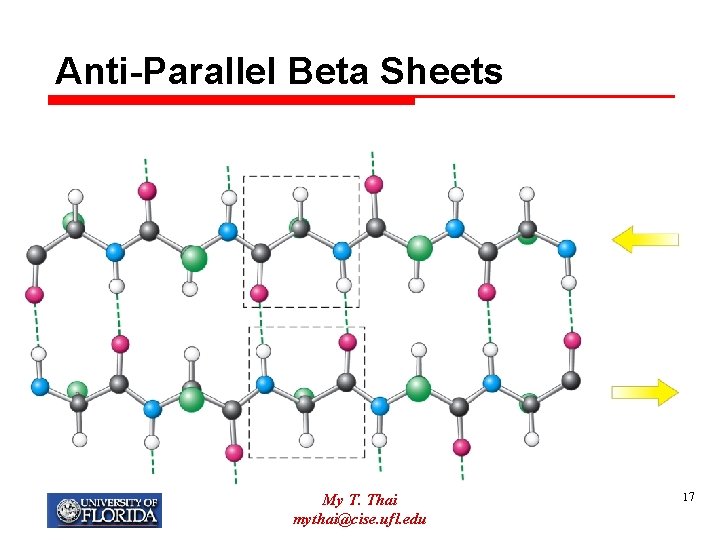
Anti-Parallel Beta Sheets My T. Thai mythai@cise. ufl. edu 17
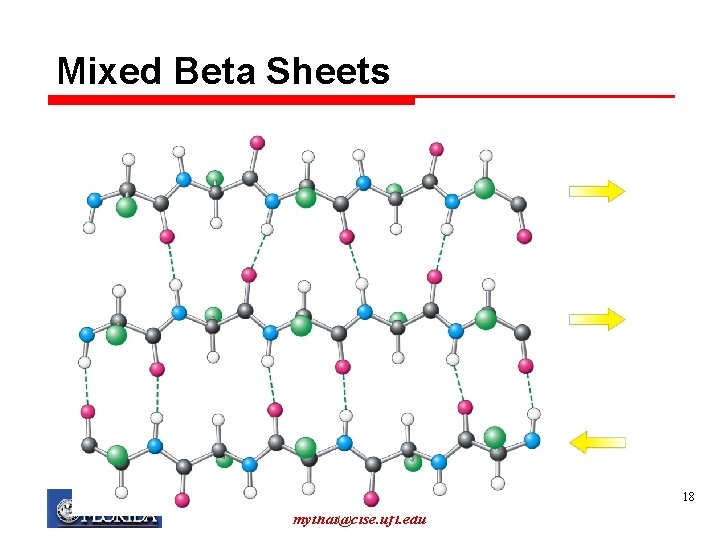
Mixed Beta Sheets My T. Thai mythai@cise. ufl. edu 18

Tertiary Structure Ø The full dimensional structure, describing the overall shape of a protein Ø Also known as its fold My T. Thai mythai@cise. ufl. edu 19

Quaternary Structure Ø Proteins are made up of multiple polypeptide chains, each called a subunit Ø The spatial arrangement of these subunits is referred to as the quaternary structure Ø Sometimes distinct proteins must combine together in order to form the correct 3 -dimensional structure for a particular protein to function properly. Ø Example: the protein hemoglobin, which carries oxygen in blood. Hemoglobin is made of four similar proteins that combine to form its quaternary structure. My T. Thai mythai@cise. ufl. edu 20

Other Units of Structure Ø Motifs (super-secondary structure): q Frequently occurring combinations of secondary structure units q A pattern of alpha-helices and beta-strands Ø Domains: A protein chain often consists of different regions, or domains q Domains within a protein often perform different functions q Can have completely different structures and folds q Typically a 100 to 400 residues long My T. Thai mythai@cise. ufl. edu 21

What Determines Structure Ø What causes a protein to fold in a particular way? Ø At a fundamental level, chemical interactions between all the amino acids in the sequence contribute to a protein’s final conformation Ø There are four fundamental chemical forces: q q Hydrogen bonds Hydrophobic effect Van der Waal Forces Electrostatic forces My T. Thai mythai@cise. ufl. edu 22

Hydrogen Bonds Ø Occurs when a pair of nucliophilic atoms such as oxygen and nitrogen share a hydrogen between them Ø Pattern of hydrogen bounding is essential in stabilizing basic secondary structures My T. Thai mythai@cise. ufl. edu 23

Van der Waal Forces Ø Interactions between immediately adjacent atoms Ø Result from the attraction between an atom’s nucleus and it neighbor’s electrons My T. Thai mythai@cise. ufl. edu 24

Electrostatic Forces Ø Oppositely charged side chains con form salt-bridges, which pulls chains together My T. Thai mythai@cise. ufl. edu 25

Experimental Determination Ø Centralized database (to deposit protein structures) called the protein Databank (PDB), accessible at http: //www. rcsb. org/pdb/index. html Ø Two main techniques are used to determine/verify the structure of a given protein: q X-ray crystallography q Nuclear Magnetic Resonance (NMR) Ø Both are slow, labor intensive, expensive (sometimes longer than a year!) My T. Thai mythai@cise. ufl. edu 26

X-ray Crystallography Ø A technique that can reveal the precise three dimensional positions of most of the atoms in a protein molecule Ø The protein is first isolated to yield a high concentration solution of the protein Ø This solution is then used to grow crystals Ø The resulting crystal is then exposed to an Xray beam My T. Thai mythai@cise. ufl. edu 27

Disadvantages Ø Not all proteins can be crystallized Ø Crystalline structure of a protein may be different from its structure Ø Multiple maps may be needed to get a consensus My T. Thai mythai@cise. ufl. edu 28

NMR Ø The spinning of certain atomic nuclei generates a magnetic moment Ø NMR measures the energy levels of such magnetic nuclei (radio frequency) Ø These levels are sensitive to the environment of the atom: q What they are bonded to, which atoms they are close to spatially, what distances are between different atoms… Ø Thus by carefully measurement, the structure of the protein can be constructed My T. Thai mythai@cise. ufl. edu 29

Disadvantages Ø Constraint of the size of the protein – an upper bound is 200 residues Ø Protein structure is very sensitive to p. H. My T. Thai mythai@cise. ufl. edu 30

Computational Methods Ø Given a long and painful experimental methods, need computational approaches to predict the structure from its sequence. My T. Thai mythai@cise. ufl. edu 31

Functional Region Prediction My T. Thai mythai@cise. ufl. edu 32

Protein Secondary Structure My T. Thai mythai@cise. ufl. edu 33

Tertiary Structure Prediction My T. Thai mythai@cise. ufl. edu 34

More Details on X-ray Crystallography My T. Thai mythai@cise. ufl. edu 35
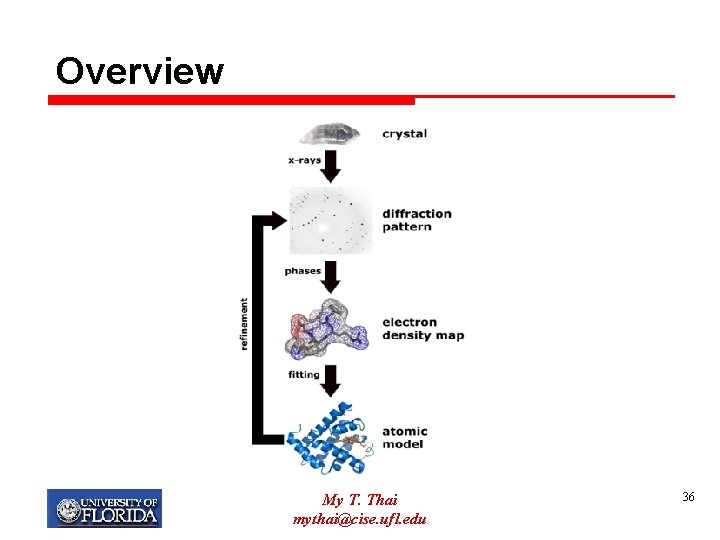
Overview My T. Thai mythai@cise. ufl. edu 36

Overview My T. Thai mythai@cise. ufl. edu 37

Crystal Ø A crystal can be defined as an arrangement of building blocks which is periodic in three dimensions My T. Thai mythai@cise. ufl. edu 38

Crystallize a Protein Ø Have to find the right combination of all the different influences to get the protein to crystallize Ø This can take a couple hundred or even thousand experiments Ø Most popular way to conduct these experiments q Hanging-drop method My T. Thai mythai@cise. ufl. edu 39

Hanging drop method Ø The reservoir contains a precipitant concentration twice as high as the protein solution Ø The protein solutions is made up of 50% of stock protein solution and 50% of reservoir solution Ø Overtime, water will diffuse from the protein drop into the reservoir Ø Both the protein concentration and precipitant concentration will increase Ø Crystals will appear after days, weeks, months My T. Thai mythai@cise. ufl. edu 40

Properties of protein crystal Ø Very soft Ø Mechanically fragile Ø Large solvent areas (30 -70%) My T. Thai mythai@cise. ufl. edu 41

A Schematic Diffraction Experiment My T. Thai mythai@cise. ufl. edu 42

Why do we need Crystals Ø A single molecule could never be oriented and handled properly for a diffraction experiment Ø In a crystal, we have about 1015 molecules in the same orientation so that we get a tremendous amplification of the diffraction Ø Crystals produce much simpler diffraction patterns than single molecules My T. Thai mythai@cise. ufl. edu 43

Why do we need X-rays Ø X-rays are electromagnetic waves with a wavelength close to the distance of atoms in the protein molecules Ø To get information about where the atoms are, we need to resolve them -> thus we need radiation My T. Thai mythai@cise. ufl. edu 44
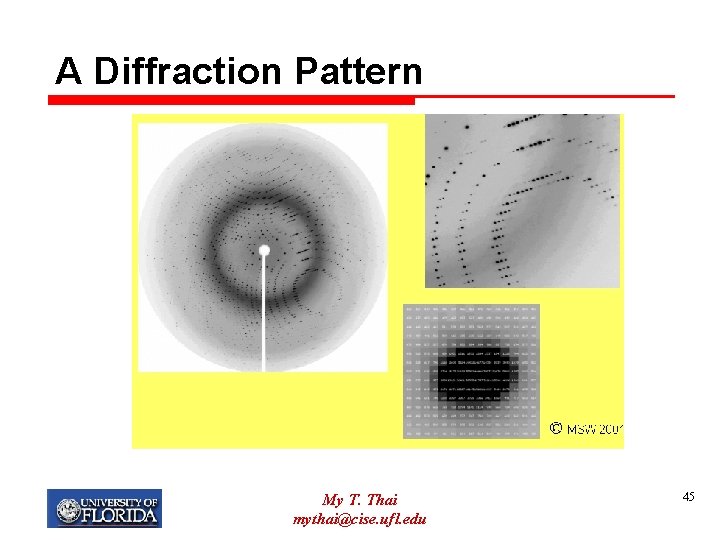
A Diffraction Pattern My T. Thai mythai@cise. ufl. edu 45

My T. Thai mythai@cise. ufl. edu 46

Resolution Ø The primary measure of crystal order/quality of the model Ø Ranges of resolution: q Low resolution (>3 -5 Ao) is difficult to see the side chains only the overall structural fold q Medium resolution (2. 5 -3 Ao) q High resolution (2. 0 Ao) My T. Thai mythai@cise. ufl. edu 47

Some Crystallographic Terms Ø h, k, l: Miller indices (like a name of the reflection) Ø I(h, k, l): intensity Ø 2θ: angle between the x-ray incident beam and reflect beam My T. Thai mythai@cise. ufl. edu 48

Diffraction by a Molecule in a Crystal Ø The electric vector of the X-ray wave forces the electrons in our sample to oscillate with the same wavelength as the incoming wave My T. Thai mythai@cise. ufl. edu 49

Description of Waves My T. Thai mythai@cise. ufl. edu 50

Structure Factor Equation Ø fj: proportional to the number of electrons this atom j has Ø One of the fundamental equations in X-ray Crystallography My T. Thai mythai@cise. ufl. edu 51

The Phase Ø From the measurement, we can only obtain the intensity I(hkl) of any given reflection (hkl) Ø The phase α(hkl) cannot be measured My T. Thai mythai@cise. ufl. edu 52
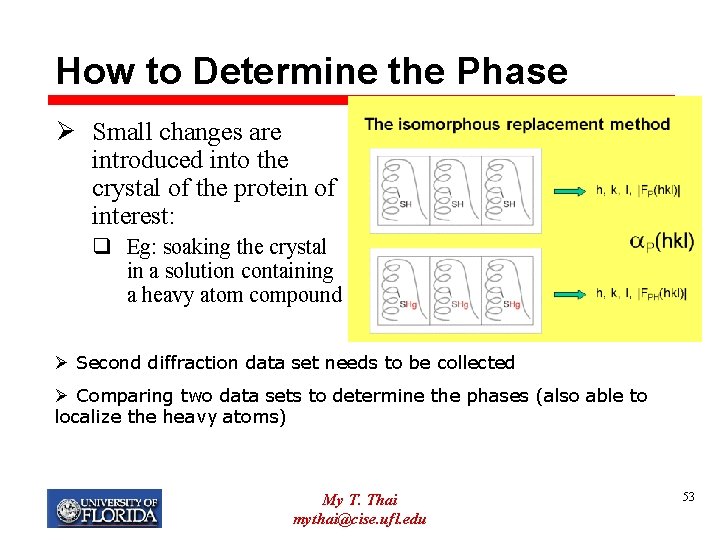
How to Determine the Phase Ø Small changes are introduced into the crystal of the protein of interest: q Eg: soaking the crystal in a solution containing a heavy atom compound Ø Second diffraction data set needs to be collected Ø Comparing two data sets to determine the phases (also able to localize the heavy atoms) My T. Thai mythai@cise. ufl. edu 53

Other Phase Determination Methods My T. Thai mythai@cise. ufl. edu 54

Electron Density Map Ø Once we know the complete diffraction pattern (amplitudes and phases), need to calculate an image of the structure Ø The above equation returns the electron density (so we get a map of where the electrons are their concentration) My T. Thai mythai@cise. ufl. edu 55

Interpretation of Electron Density Ø Now, the electron density has to be interpreted in terms of atom identities and positions. Ø (1): packing of the whole molecules is shown in the crystal Ø (2): a chain of seven amino acids in shown with the resulting structure superimposed Ø (3): the electron density of a trypophan side chain is shown My T. Thai mythai@cise. ufl. edu 56

Refinement and the R-Factor My T. Thai mythai@cise. ufl. edu 57

Nuclear Magnetic Resonance Ø Concentrated protein solution (very purified) Ø Magnetic field Ø Effect of radio frequencies on the resonance of different atoms is measured. My T. Thai mythai@cise. ufl. edu 58
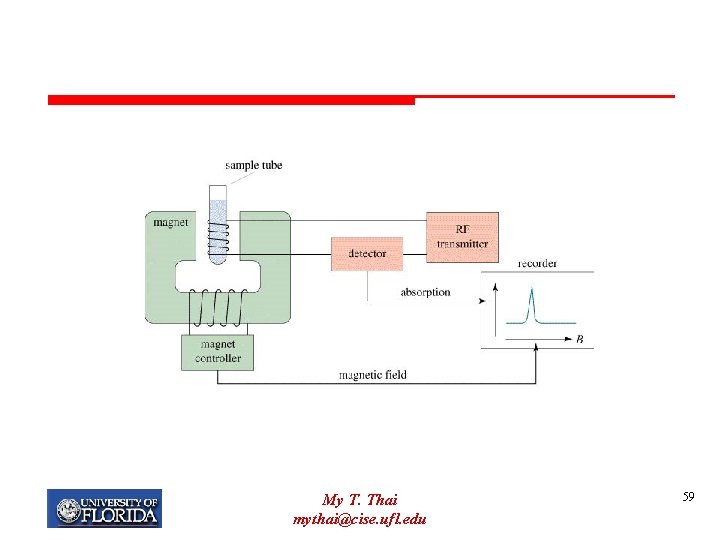
My T. Thai mythai@cise. ufl. edu 59

NMR Ø Behavior of any atom is influenced by neighboring atoms Ø more closely spaced residues are more perturbed than distant residues Ø can calculate distances based on perturbation My T. Thai mythai@cise. ufl. edu 60

NMR spectrum of a protein My T. Thai mythai@cise. ufl. edu 61

Computational Molecular Biology Protein Structure: Secondary Prediction

Primary Structure: Symbolic Definition Ø A = {A, C, D, E, F, G, H, I, J, K, L, M, N, P, Q, R, S. T, V, W, Y } – set of symbols denoting all amino acids Ø A* - set of all finite sequences formed out of elements of A, called protein sequences Ø Elements of A* are denoted by x, y, z …. . i. e. we write x A*, y A*, z A*, … etc Ø PROTEIN PRIMARY STRUCTURE: any x A* is also called a protein sequence or protein sub-unit My T. Thai mythai@cise. ufl. edu 63

Protein Secondary Structure (PSS) Ø Secondary structure: the arrangement of the peptide backbone in space. It is produced by hydrogen bondings between amino acids Ø PROTEIN SECONDARY STRUCTURE consists of: protein sequence and its hydrogen bonding patterns called SS categories My T. Thai mythai@cise. ufl. edu 64

Protein Secondary Structure Ø Databases for protein sequences are expanding rapidly Ø The number of determined protein structures (PSS – protein secondary structures) and the number of known protein sequences is still limited ØPSSP (Protein Secondary Structure Prediction) research is trying to breach this gap. My T. Thai mythai@cise. ufl. edu 65

Protein Secondary Structure Ø The most commonly observed conformations in secondary structure are: q. Alpha Helix q. Beta Sheets/Strands q. Loops/Turns My T. Thai mythai@cise. ufl. edu 66

Turns and Loops Ø Secondary structure elements are connected by regions of turns and loops Ø Turns – short regions of non- , non- conformation Ø Loops – larger stretches with no secondary structure. My T. Thai mythai@cise. ufl. edu 67

Three secondary structure states Ø Prediction methods are normally assessed for 3 states: q H (helix) q E (strands) q L (others (loop or turn)) My T. Thai mythai@cise. ufl. edu 68
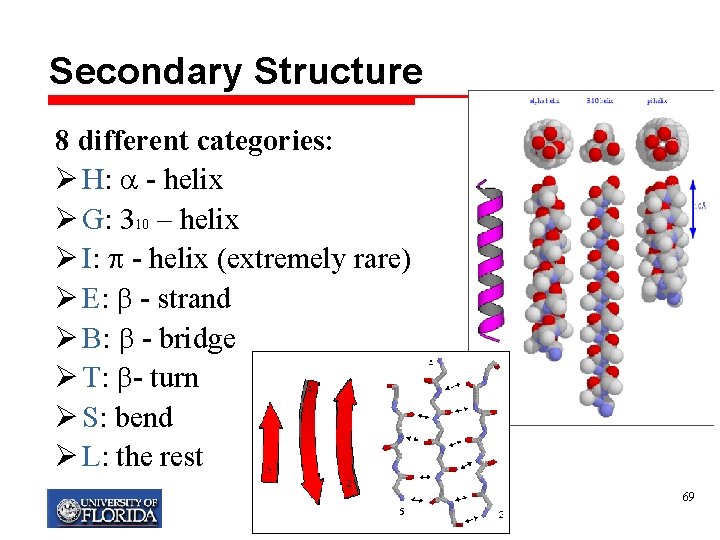
Secondary Structure 8 different categories: Ø H: - helix Ø G: 310 – helix Ø I: - helix (extremely rare) Ø E: - strand Ø B: - bridge Ø T: - turn Ø S: bend Ø L: the rest My T. Thai mythai@cise. ufl. edu 69

Three SS states: Reduction methods Ø Method 1, used by DSSP program: q H(helix) ={ G (310 – helix), H ( - helix)} q E (strands) = {E ( -strand), B ( -bridge)} , q L = all the rest • Shortly: E, B => E; G, H => H; Rest => C Ø Method 2, used by STRIDE program: q H as in Method 1 q E = {E ( -strand), b (isolated -bridge)}, q L = all the rest My T. Thai mythai@cise. ufl. edu 70

Three SS states: Reduction methods ØMethod 3, used by DEFINE program: q H(helix) as in Method 1 q E (strands) = {E ( -strand)}, q. L = all the rest My T. Thai mythai@cise. ufl. edu 71

Example of typical PSS Data ØExample: q Sequence § KELVLALYDYQEKSPREVTHKKGDILTLLNSTNKDWWKYEY NDRQGFVP q Observed SS § HHHHHLLLLEEEHHHLLLEEEEEELLLHHHHLLLEEEE EELLLHHH My T. Thai mythai@cise. ufl. edu 72

PSS: Symbolic Definition ØGiven A = {A, C, D, E, F, G, H, I, J, K, L, M, N, P, Q, R, S. T, V, W, Y } – set of symbols denoting amino acids and a protein sequence x A* ØLet S ={ H, E, L} be the set of symbols of 3 states: H (helix), E (strands) and L (loop) and S* be the set of all finite sequences of elements of S. Ø We denote elements of S* by e, e S* My T. Thai mythai@cise. ufl. edu 73

PSS: Symbolic Definition Ø Any one-to-one function f : A* S* i. e. f A* x S* is called a protein secondary structure (PSS) identification function ØAn element (x, e) f is a called protein secondary structure (of the protein sequence x) ØThe element e S* (of (x, e) f ) is called secondary structure. My T. Thai mythai@cise. ufl. edu 74

PSSP Ø If a protein sequence shows clear similarity to a protein of known three dimensional structure q then the most accurate method of predicting the secondary structure is to align the sequences by standard dynamic programming algorithms q. Why? § homology modelling is much more accurate than secondary structure prediction for high levels of sequence identity. My T. Thai mythai@cise. ufl. edu 75

PSSP Ø Secondary structure prediction methods are of most use when sequence similarity to a protein of known structure is undetectable. Ø It is important that there is no detectable sequence similarity between sequences used to train and test secondary structure prediction methods. My T. Thai mythai@cise. ufl. edu 76

Classification and Classifiers Ø Given a database table DB with a special atribute C, called a class attribute (or decision attribute). The values: C 1, C 2, . . . Cn of the class atrribute are called class labels. Ø Example: A 1 1 0 1 A 2 1 1 0 A 3 m v m My T. Thai mythai@cise. ufl. edu A 4 g g b C c 1 c 2 c 1 77

Classification and Classifiers Ø The attribute C partitions the records in the DB: q divides the records into disjoint subsets defined by the attributes C values, CLASSIFIES the records. q It means we use the attributre C and its values to divide the set R of records of DB into n disjoint classes: C 1={ r DB: C=c 1} . . . Cn={r DB: C=cn} Ø Example (from our table) C 1 = { (1, 1, m, g), (1, 0, m, b)} = {r 1, r 3} C 2 = { (0, 1, v, g)} ={r 2} My T. Thai mythai@cise. ufl. edu 78

Classification and Classifiers Ø An algorithm is called a classification algorithm if it uses the data and its classification to build a set of patterns. Ø Those patterns are structured in such a way that we can use them to classify unknown sets of objects- unknown records. Ø For that reason (because of the goal) the classification algorithm is often called shortly a classifier. Ø The name classifier implies more then just classification algorithm. A classifier is final product of a data set and a classification algorithm. My T. Thai mythai@cise. ufl. edu 79

Classification and Classifiers Ø Building a classifier consists of two phases: training and testing. Ø In both phases we use data (training data set and disjoint with it test data set) for which the class labels are known for ALL of the records. Ø We use the training data set to create patterns Ø We evaluate created patterns with the use of of test data, which classification is known. Ø The measure for a trained classifier accuracy is called predictive accuracy. Ø The classifier is build i. e. we terminate the process if it has been trained and tested and predictive accuracy was on an acceptable level. My T. Thai mythai@cise. ufl. edu 80

Classifiers Predictive Accuracy Ø PREDICTIVE ACCURACY of a classifier is a percentage of well classified data in the testing data set. ØPredictive accuracy depends heavily on a choice of the test and training data. Ø There are many methods of choosing test and training sets and hence evaluating the predictive accuracy. This is a separate field of research. My T. Thai mythai@cise. ufl. edu 81

Accuracy Evaluation Ø Use training data to adjust parameters of method until it gives the best agreement between its predictions and the known classes Ø Use the testing data to evaluate how well the method works (without adjusting parameters!) Ø How do we report the performance? Ø Average accuracy = fraction of all test examples that were classified correctly My T. Thai mythai@cise. ufl. edu 82

Accuracy Evaluation Ø Multiple cross-validation test has to be performed to exclude a potential dependency of the evaluated accuracy on the particular test set chosen Ø Jack-Knife: q Use 129 chains for setting up the tool (training set) q 1 for estimating the performance (testing) q This has to be repeated 130 times until each protein has been used once for testing q The average over all 130 tests gives an estimate of the prediction accuracy My T. Thai mythai@cise. ufl. edu 83

PSSP Datasets Ø Historic RS 126 dataset. Contains 126 sub-units with known secondary structure selected by Rost and Sander. Today is not used anymore Ø CB 513 dataset. Contains 513 sub-units with known secondary structure selected by Cuff and Barton in 1999. Used quite frencently in PSSP research Ø HS 17771 dataset. Created by Hobohm and Scharf. In March-2002 it contained 1771 sub-units Ø Lots of authors has their own and “secret” datasets My T. Thai mythai@cise. ufl. edu 84

Measures for PSSP accuracy Ø http: //cubic. bioc. columbia. edu/eva/doc/measur e_sec. html (for more information) Ø Q 3 : Three-state prediction accuracy (percent of succesful classified) Ø Qi %obs: How many of the observed residues were correctly predicted? Ø Qi %prd: How many of the predicted residues were correctly predicted? My T. Thai mythai@cise. ufl. edu 85

Measures for PSSP Accuracy Ø Aij = number of residues predicted to be in structure type j and observed to be in type i Ø Number of residues predicted to be in structure i: Ø Number of residues observed to be in structure i: My T. Thai mythai@cise. ufl. edu 86

Measures for SSP Accuracy Ø The percentage of residues correctly predicted to be in class i relative to those observed to be in class i Ø The percentages of residues correctly predicted to be in class i from all residues predicted to be in i Ø Overall 3 -state accuracy My T. Thai mythai@cise. ufl. edu 87

PSSP Algorithms ØThere are three generations in PSSP algorithms • First Generation: based on statistical information of single amino acids (1960 s and 1970 s) • Second Generation: based on windows (segments) of amino acids. Typically a window containes 11 -21 amino acids (dominating the filed until early 1990 s) • Third Generation: based on the use of windows on evolutionary information My T. Thai mythai@cise. ufl. edu 88

PSSP: First Generation Ø First generation PSSP systems are based on statistical information on a single amino acid Ø The most relevant algorithms: q. Chow-Fasman, 1974 q. GOR, 1978 Ø Both algorithms claimed 74 -78% of predictive accuracy, but tested with better constructed datasets were proved to have the predictive accuracy ~50% (Nishikawa, 1983) My T. Thai mythai@cise. ufl. edu 89

Chou-Fasman method Ø Uses table of conformational parameters determined primarily from measurements of the known structure (from experimental methods) Ø Table consists of one “likelihood” for each structure for each amino acid Ø Based on frequencies of residues in -helices, -sheets and turns Ø Notation: P(H): propensity to form alpha helices Ø f(i): probability of being in position 1 (of a turn) My T. Thai mythai@cise. ufl. edu 90
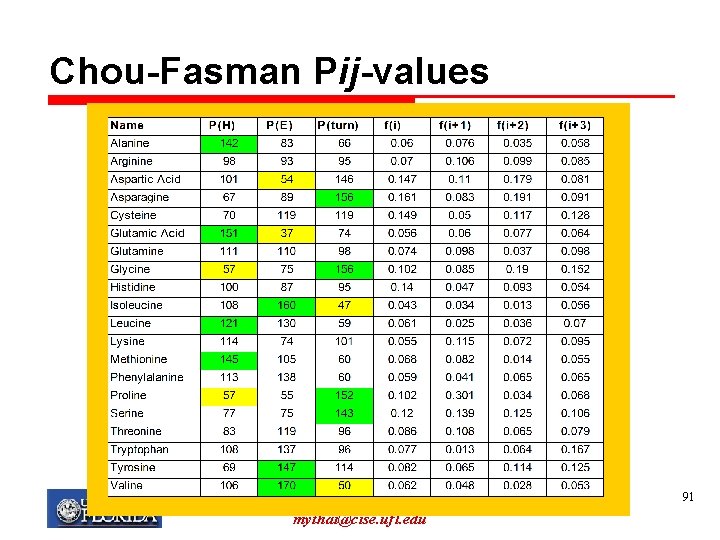
Chou-Fasman Pij-values My T. Thai mythai@cise. ufl. edu 91

Chou-Fasman Ø A prediction is made for each type of structure for each amino acid q Can result in ambiguity if a region has high propensities for both helix and sheet (higher value usually chosen) My T. Thai mythai@cise. ufl. edu 92

Chou-Fasman How it works: 1. Assign all of the residues the appropriate set of parameters 2. Identify -helix and -sheet regions. Extend the regions in both directions. 3. If structures overlap compare average values for P(H) and P(E) and assign secondary structure based on best scores. 4. Turns are calculated using 2 different probability values. My T. Thai mythai@cise. ufl. edu 93

Assign Pij values 1. Assign all of the residues the appropriate set of parameters My T. Thai mythai@cise. ufl. edu 94

Scan peptide for a-helix regions 2. Identify regions where 4 out of 6 have a P(H) >100 “alpha-helix nucleus” My T. Thai mythai@cise. ufl. edu 95

Extend a-helix nucleus 3. Extend helix in both directions until a set of four consecutive residues with P(H) <100. Find sum of P(H) and sum of P(E) in the extended region If region is long enough ( >= 5 letters) and sum P(H) > sum P(E) then declare the extended region as alpha helix My T. Thai mythai@cise. ufl. edu 96

Scan peptide for b-sheet regions 4. Identify regions where 3 out of 5 have a P(E) >100 “ -sheet nucleus” 5. Extend -sheet until 4 continuous residues with an average P(E) < 100 6. If region average > 100 and the average P(E) > average P(H) then “ -sheet” My T. Thai mythai@cise. ufl. edu 97

Overlapping Ø Resolving overlapping alpha helix & beta sheet q. Compute sum of P(H) and sum of P(E) in the overlap. q. If sum P(H) > sum P(E) => alpha helix q. If sum P(E) > sum P(H) => beta sheet My T. Thai mythai@cise. ufl. edu 98

Turn Prediction Ø An amino acid is predicted as turn if all of the following holds: qf(i)*f(i+1)*f(i+2)*f(i+3) > 0. 000075 q. Avg(P(i+k)) > 100, for k=0, 1, 2, 3 q. Sum(P(t)) > Sum(P(H)) and Sum(P(E)) for i+k, (k=0, 1, 2, 3) My T. Thai mythai@cise. ufl. edu 99

PSSP: Second Generation ØBased on the information contained in a window of amino acids (11 -21 aa. ) ØThe most systems use algorithms based on: q q q q Statistical information Physico-chemical properties Sequence patterns Graph-theory Multivariante statistics Expert rules Nearest-neighbour algorithms My T. Thai mythai@cise. ufl. edu 100

PSSP: First & Second Generation ØMain problems: q. Prediction accuracy <70% § SS assigments differ even between crystals of the same protein § SS formation is partially determined by longrange interactions, i. e. , by contacts between residues that are not visible by any method based on windows of 11 -21 adjacent residues My T. Thai mythai@cise. ufl. edu 101

PSSP: First & Second Generation ØMain problems: q. Prediction accuracy for -strand 28 -48%, only slightly better than random § beta-sheet formation is determined by more nonlocal contacts than in alpha-helix formation q. Predicted helices and strands are usually too short § Overlooked by most developers My T. Thai mythai@cise. ufl. edu 102

Example of Second Generation Ø Example for typical secondary structure prediction of the 2 nd generation. Ø The protein sequence (SEQ ) given was the SH 3 structure. Ø The observed secondary structure (OBS ) was assigned by DSSP (H = helix; E = strand; blank = non-regular structure; the dashes indicate the continuation). Ø The typical prediction of too short segments (TYP ) poses the following problems in practice. q (i) Are the residues predicted to be strand in segments 1, 5, and 6 errors, or should the helices be elongated? q (ii) Should the 2 nd and 3 rd strand be joined, or should one of them be ignored, or does the prediction indicate two strands, here? Note: the three-state per-residue accuracy is 60% for the prediction given. My T. Thai mythai@cise. ufl. edu 103

PSSP: Third Generation Ø PHD: First algorithm in this generation (1994) Ø Evolutionary information improves the prediction accuracy to 72% Ø Use of evolutionary information: 1. Scan a database with known sequences with alignment methods for finding similar sequences 2. Filter the previous list with a threshold to identify the most significant sequences 3. Build amino acid exchange profiles based on the probable homologs (most significant sequences) 4. The profiles are used in the prediction, i. e. in building the classifier My T. Thai mythai@cise. ufl. edu 104

PSSP: Third Generation Ø Many of the second generation algorithms have been updated to the third generation My T. Thai mythai@cise. ufl. edu 105

PSSP: Third Generation Ø Due to the improvement of protein information in databases i. e. better evolutionary information, today’s predictive accuracy is ~80% Ø It is believed that maximum reachable accuracy is 88%. Why such conjecture? My T. Thai mythai@cise. ufl. edu 106

Why 88% Ø SS assignments may vary for two versions of the same structure q Dynamic objects with some regions being more mobile than others q Assignment differ by 5 -15% between different Xray (NMR) versions of the same protein q Assignment diff. by about 12% between structural homologues Ø B. Rost, C. Sander, and R. Schneider, Redefining the goals of protein secondary structure predictions, J. Mol. Bio. My T. Thai mythai@cise. ufl. edu 107

PSSP Data Preparation Ø Public Protein Data Sets used in PSSP research contain protein secondary structure sequences. In order to use classification algorithms we must transform secondary structure sequences into classification data tables. Ø Records in the classification data tables are called, in PSSP literature (learning) instances. Ø The mechanism used in this transformation process is called window. Ø A window algorithm has a secondary structure as input and returns a classification table: set of instances for the classification algorithm. My T. Thai mythai@cise. ufl. edu 108

Window Ø Consider a secondary structure (x, e). where (x, e)= (x 1 x 2 …xn, e 1 e 2…en) Ø Window of the length w chooses a subsequence of length w of x 1 x 2 …xn, and an element ei from e 1 e 2…en, corresponding to a special position in the window, usually the middle Ø Window moves along the sequences x = x 1 x 2 …xn and e= e 1 e 2…en simultaneously, starting at the beginning moving to the right one letter at the time at each step of the process. My T. Thai mythai@cise. ufl. edu 109

Window: Sequence to Structure Ø Such window is called sequence to structure window. We will call it for short a window. Ø The process terminates when the window or its middle position reaches the end of the sequence x. Ø The pair: (subsequence, element of e ) is often written in a form Ø subsequence H, E or L is called an instance, or a rule. My T. Thai mythai@cise. ufl. edu 110

Example: Window Ø Consider a secondary structure (x, e) and the window of length 5 with the special position in the middle (bold letters) Ø Fist position of the window is: x = A R N S T V V S T A A …. e = H H L L L E E E Ø Window returns instance: Ø A R N S T H My T. Thai mythai@cise. ufl. edu 111

Example: Window Ø Second position of the window is: x = A R N S T V V S T A A …. e = H H L L L E E E Ø Windows returns instance: q. R N S T V H Ø Next instances are: q. N S T V V L q. S T V V S L q. T V V S T L My T. Thai mythai@cise. ufl. edu 112

Symbolic Notation ØLet f be a protein secondary structure (PSS) identification function: Øf : A* S* i. e. f A* x S* Ø Let x= x 1 x 2…xn, e= e 1 e 2…en, f(x)= e, we define Øf(x 1 x 2…xn)|{xi}= ei, i. e. f(x)|{xi}= ei My T. Thai mythai@cise. ufl. edu 113

Example: Semantics of Instances Let • x = A R N S T V V S T A A …. • e = H H L L L E E E And assume that the windows returns an instance: ARNST H • Semantics of the instance is: f(x)|{N}=H, where f is the identification function and N is preceded by A R and followed by S T and the window has the length 5 My T. Thai mythai@cise. ufl. edu 114

Classification Data Base (Table) Ø We build the classification table with attributes being the positions p 1, p 2, p 3, p 4, p 5. . pw in the window, where w is length of the window. The corresponding values of attributes are elements of of the subsequent on the given position. Ø Classification attribute is S with values in the set {H, E, L} assigned by the window operation (instance, rule). Ø The classification table for our example (first few records) is the following. My T. Thai mythai@cise. ufl. edu 115

Classification Table (Example) Ø x = A R N S T V V S T A A …. Ø e = H H L L L E E E p 1 p 2 p 3 p 4 p 5 S A R N S T H R N S T S T V H T V V L V V S L Semantics of record r= r(p 1, p 2, p 3, p 4, p 5, S) is : f(x)|{Vp 3} = Vs where Va denotes a value of the attribute a. My T. Thai mythai@cise. ufl. edu 116

Size of classification datasets (tables) ØThe window mechanism produces very large datasets ØFor example window of size 13 applied to the CB 513 dataset of 513 protein subunits produces about 70, 000 records (instances) My T. Thai mythai@cise. ufl. edu 117

Window Ø Window has the following parameters: Ø PARAMETER 1 : i N+, the starting point of the window as it moves along the sequence x= x 1 x 2 …. xn. The value i=1 means that window starts at x 1, i=5 means that window starts at x 5 Ø PARAMETER 2: w N+ denotes the size (length) of the window. Ø For example: the PHD system of Rost and Sander (1994) uses two window sizes: 13 and 17. My T. Thai mythai@cise. ufl. edu 118

Window Ø PARAMETER 3: p {1, 2, …, w} where p is a special position of the window that returns the classification attribute values from S ={H, E, L} and w is the size (length) of the window Ø PSSP PROBLEM: find optimal size w, optimal special position p for the best prediction accuracy My T. Thai mythai@cise. ufl. edu 119

Window: Symbolic Definition Ø Window Arguments: window parameters and secondary structure (x, e) Ø Window Value: (subsequence of x, element of e) Ø OPERATION (sequence – to –structure window) W is a partial function W: N+ {1, …, k} (A* S* ) A* S W(i, k, p, (x, e)) = (xi x(i+1)…. x(i+k-1), f(x)|{x(i+p)}) where (x, e)= (x 1 x 2. . xn, e 1 e 2…en) My T. Thai mythai@cise. ufl. edu 120

Neural network models Ø machine learning approach Ø provide training sets of structures (e. g. -helices, non helices) Ø are trained to recognize patterns in known secondary structures Ø provide test set (proteins with known structures) Ø accuracy ~ 70 – 75% My T. Thai mythai@cise. ufl. edu 121

Reasons for improved accuracy Ø Align sequence with other related proteins of the same protein family Ø Find members that has a known structure Ø If significant matches between structure and sequence assign secondary structures to corresponding residues My T. Thai mythai@cise. ufl. edu 122

3 State Neural Network My T. Thai mythai@cise. ufl. edu 123

Neural Network My T. Thai mythai@cise. ufl. edu 124

Input Layer Ø Most of approach set w = 17. Why? q Based on evidence of statistical correlation with secondary structure as far as 8 residues on either side of the prediction point Ø The input layer consists of: q 17 blocks, each represent a position of window q Each block has 21 units: § The first 20 units represent the 20 aa § One to provide a null input used when the moving window overlaps the amino- or carboxyl-terminal end of the protein My T. Thai mythai@cise. ufl. edu 125

Binary Encoding Scheme Ø Example: Ø Let w = 5, and let say we have the sequence: A E G K Q…. Ø Then the input layer is: Ø A, C, D, E, F, G, …, N, P, Q, R, S. T, V, W, Y 1 0 0 0 0 …. 0 0… 1 0 …. . 0 … 0 1 0 …. . My T. Thai mythai@cise. ufl. edu 126

Hidden Layer Ø Represent the structure of the central aa Ø Encoding scheme: q Can use two units to present: § (1, 0) = H, (0, 1) = E, (0, 0) = L q Some uses three units: § (1, 0, 0) = H, (0, 1, 0) = E, (0, 0, 1) = L Ø For each connection, we can assign some weight value. Ø This weight value can be adjusted to best fit the data (training) My T. Thai mythai@cise. ufl. edu 127

Output Level Ø Based on the hidden level and some function f, calculate the output. Ø Helix is assigned to any group of 4 or more contiguous residues q Having helix output values greater than sheet outputs and greater than some threshold t Ø Strand (E) is assigned to any group of two or more contiguous resides, having sheet output values greater than helix outputs and greater than t Ø Otherwise, assigned to L Ø Note that t can be adjusted as well (training) My T. Thai mythai@cise. ufl. edu 128

How PHD works Step 1. BLAST search with input sequence Step 2. Perform multiple seq. alignment and calculate aa frequencies for each position My T. Thai mythai@cise. ufl. edu 129

How PHD works Step 3. First Level: “Sequence to structure net” Input: alignment profile, Output: units for H, E, L Calculate “occurrences” of any of the residues to be present in either an a-helix, b-strand, or loop. N=0. 2, S=0. 4, A=0. 4 1 2 3 4 5 6 7 H = 0. 05 E = 0. 18 L= 0. 67 My T. Thai mythai@cise. ufl. edu 130

How PHD works Step 3. Second Level: “Structure to structure net” Input: First Level values, Output: units for H, E, L Window size = 17 H = 0. 59 E = 0. 09 L= 0. 31 E=0. 18 Step 4. Decision level My T. Thai mythai@cise. ufl. edu 131
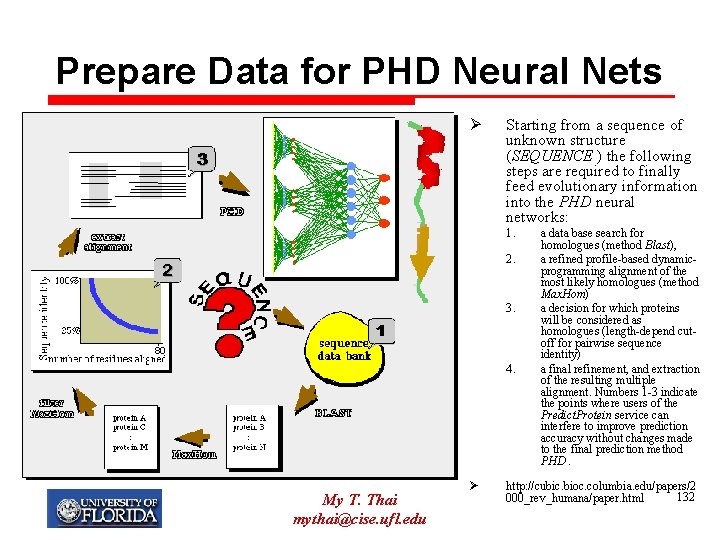
Prepare Data for PHD Neural Nets Ø Starting from a sequence of unknown structure (SEQUENCE ) the following steps are required to finally feed evolutionary information into the PHD neural networks: 1. 2. 3. 4. My T. Thai mythai@cise. ufl. edu Ø a data base search for homologues (method Blast), a refined profile-based dynamicprogramming alignment of the most likely homologues (method Max. Hom) a decision for which proteins will be considered as homologues (length-depend cutoff for pairwise sequence identity) a final refinement, and extraction of the resulting multiple alignment. Numbers 1 -3 indicate the points where users of the Predict. Protein service can interfere to improve prediction accuracy without changes made to the final prediction method PHD. http: //cubic. bioc. columbia. edu/papers/2 132 000_rev_humana/paper. html

PHD Neural Network My T. Thai mythai@cise. ufl. edu 133

Prediction Accuracy My T. Thai mythai@cise. ufl. edu 134

Where can I learn more? Protein Structure Prediction Center Biology and Biotechnology Research Program Lawrence Livermore National Laboratory, Livermore, CA http: //predictioncenter. llnl. gov/Center. html DSSP Database of Secondary Structure Prediction http: //www. sander. ebi. ac. uk/dssp/ My T. Thai mythai@cise. ufl. edu 135

Computational Molecular Biology Protein Structure: Tertiary Prediction via Threading

Objective Ø Study the problem of predicting the tertiary structure of a given protein sequence My T. Thai mythai@cise. ufl. edu 137
A Few Examples actual predicted My T. Thai mythai@cise. ufl. edu actual predicted 138

Two Comparative Modeling Ø Homology modeling – identification of homologous proteins through sequence alignment; structure prediction through placing residues into “corresponding” positions of homologous structure models Ø Protein threading – make structure prediction through identification of “good” sequence-structure fit Ø We will focus on the Protein Threading. My T. Thai mythai@cise. ufl. edu 139

Why it Works? Ø Observations: q Many protein structures in the PDB are very similar § Eg: many 4 -helical bundles, globins… in the set of solved structure Ø Conjecture: q There are only a limited number of “unique” protein folds in nature My T. Thai mythai@cise. ufl. edu 140

Threading Method Ø General Idea: q Try to determine the structure of a new sequence by finding its best ‘fit’ to some fold in library of structures Ø Sequence-Structure Alignment Problem: q Given a solved structure T for a sequence t 1 t 2…tn and a new sequence S = s 1 s 2… sm, we need to find the “best match” between S and T My T. Thai mythai@cise. ufl. edu 141

What to Consider Ø How to evaluate (score) a given alignment of s with a structure T? Ø How to efficiently search over all possible alignments? My T. Thai mythai@cise. ufl. edu 142

Three Main Approaches Ø Protein Sequence Alignment Ø 3 D Profile Method Ø Contact Potentials My T. Thai mythai@cise. ufl. edu 143

Protein Sequence Alignment Method Ø Align two sequences S and T Ø If in the alignment, si aligns with tj, assign si to the position pj in the structure Ø Advantages: q Simple Ø Disadvantages: q Similar structures have lots of sequence variability, thus sequence alignment may not be very helpful My T. Thai mythai@cise. ufl. edu 144

3 D Profile Method Ø Actually uses structural information Ø Main idea: q Reduce the 3 D structure to a 1 D string describing the environment of each position in the protein. (called the 3 D profile (of the fold)) q To determine if a new sequence S belongs to a given fold T, we align the sequence with the fold’s 3 D profile Ø First question: How to create the 3 D profile? My T. Thai mythai@cise. ufl. edu 145

Create the 3 D Profile Ø For a given fold, do: 1. For each residue, determine: q How buried is it? q Fraction of surrounding environment that is polar q What secondary structure is it in (alpha-helix, betasheet, or neither) My T. Thai mythai@cise. ufl. edu 146

Create the 3 D profile 2. Assign an environment class to each position: Six classes describe the burial and polarity criteria (exposed, partially buried, very buried, different fractions of polar environment) My T. Thai mythai@cise. ufl. edu 147

Create the 3 D Profile Ø These environment classes depend on the number of surrounding polar residues and how buried the position is. Ø There are 3 SS for each of these, thus have 18 environment classes My T. Thai mythai@cise. ufl. edu 148

Create the 3 D Profile 3. Convert the known structure T to a string of environment descriptors: 4. Align the new sequence S with E using dynamic programming My T. Thai mythai@cise. ufl. edu 149

Scores for Alignment Ø Need scores for aligning individual residues with environments. Ø Key: Different aa prefer diff. environment. Thus determine scores by looking at the statistical data My T. Thai mythai@cise. ufl. edu 150

Scores for Alignment 1. Choose a database of known structures 2. Tabulate the number of times we see a particular residue in a particular environment class -> compute the score for each env class and each aa pair 3. Choose gap penalties, eg. may charge more for gaps in alpha and beta environments… My T. Thai mythai@cise. ufl. edu 151

Alignment Ø This gives us a table of scores for aligning an aa sequence with an environment string Ø Using this scoring and Dynamic Programming, we can find an optimal alignment and score for each fold in our library Ø The fold with the highest score is the best fold for the new sequence 152 My T. Thai mythai@cise. ufl. edu

Contact Potentials Method Ø Take 3 D structure into account more carefully Ø Include information about how residues interact with each other q Consider pairwise interactions between the position pi, pj in the fold q For a given alignment, produce a score which is the sum over these interactions: My T. Thai mythai@cise. ufl. edu 153

Problem Ø Have a sequence from the database T = t 1…tn with known positions p 1…pn, and a new sequence S = s 1…sm. Ø Find 1 <= r 1 < r 2 < … < rn < m which maximize where ri is the index of the aa in S which occupies position pi Ø This problem is NP-complete for pairwise interactions My T. Thai mythai@cise. ufl. edu 154

How to Define that Score? Ø Use so-called “knowledge-based potentials”, which comes from databases of observed interactions. Ø The general form: My T. Thai mythai@cise. ufl. edu 155

How to Define the Score Ø General Idea: q Define cutoff parameter for “contact” (e. g. up to 6 Angstroms) q Use the PDB to count up the number of times aa i and j are in contact Ø Several method for normalization. Eg. Normalization is by hypothetical random frequencies My T. Thai mythai@cise. ufl. edu 156

Other Variations Ø Many other variations in defining the potentials Ø In addition to pairwise potentials, consider single residue potentials Ø Distance-dependent intervals: q Counting up pairwise contacts separately for intervals within 1 Angstrom, between 1 and 2 Angstroms… My T. Thai mythai@cise. ufl. edu 157

Threading via Tree-Decomposition My T. Thai mythai@cise. ufl. edu 158
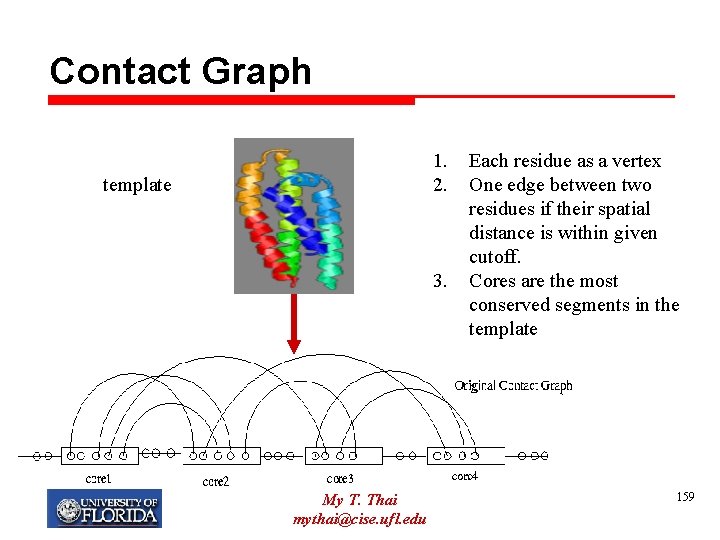
Contact Graph 1. 2. template 3. My T. Thai mythai@cise. ufl. edu Each residue as a vertex One edge between two residues if their spatial distance is within given cutoff. Cores are the most conserved segments in the template 159

Simplified Contact Graph My T. Thai mythai@cise. ufl. edu 160
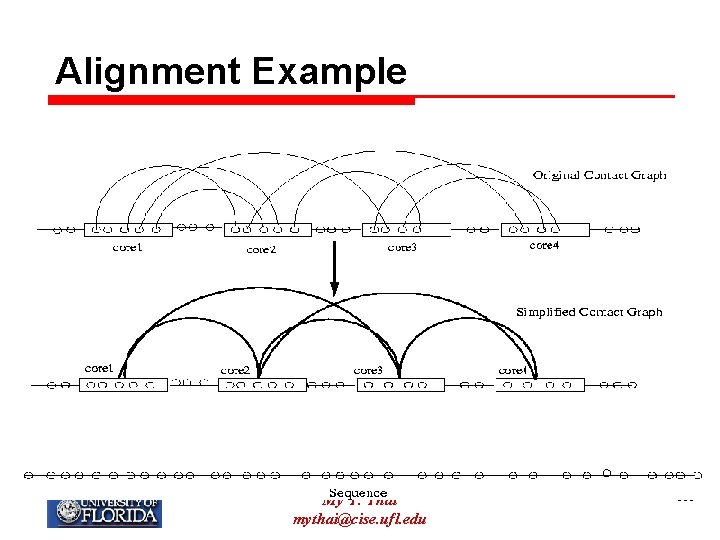
Alignment Example My T. Thai mythai@cise. ufl. edu 161
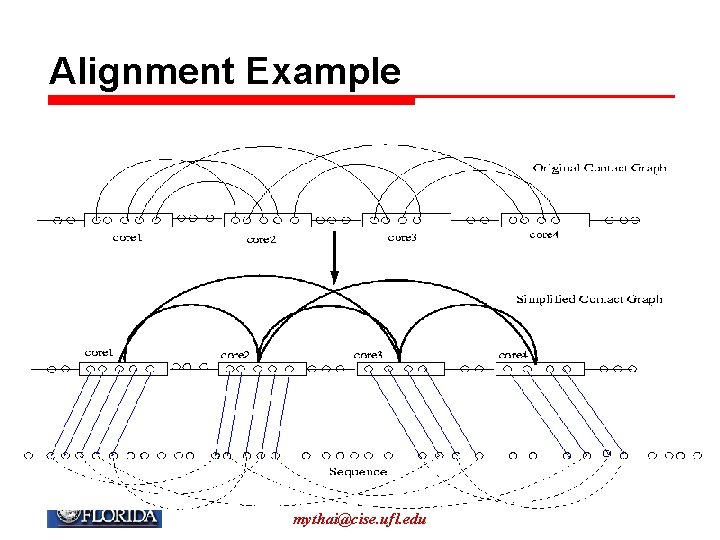
Alignment Example My T. Thai mythai@cise. ufl. edu 162

Calculation of Alignment Score My T. Thai mythai@cise. ufl. edu 163
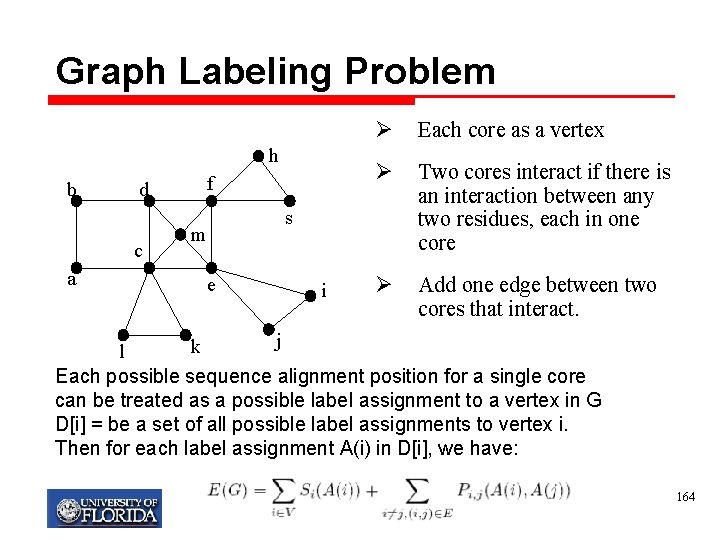
Graph Labeling Problem h b d c a f Ø Each core as a vertex Ø Two cores interact if there is an interaction between any two residues, each in one core Ø Add one edge between two cores that interact. s m e i j k l Each possible sequence alignment position for a single core can be treated as a possible label assignment to a vertex in G D[i] = be a set of all possible label assignments to vertex i. Then for each label assignment A(i) in D[i], we have: My T. Thai mythai@cise. ufl. edu 164

Tree Decomposition My T. Thai mythai@cise. ufl. edu 165
![Tree Decomposition [Robertson & Seymour, 1986] Greedy: minimum degree heuristic b f d c Tree Decomposition [Robertson & Seymour, 1986] Greedy: minimum degree heuristic b f d c](http://slidetodoc.com/presentation_image/2aba40f3319561958832b0387e61918f/image-166.jpg)
Tree Decomposition [Robertson & Seymour, 1986] Greedy: minimum degree heuristic b f d c h c e l 1. 2. 3. 4. 5. k i f d abd g m a h m a e l j g k i j Choose the vertex with minimum degree The chosen vertex and its neighbors form a component Add one edge to any two neighbors of the chosen vertex Remove the chosen vertex Repeat the above steps until the graph is empty My T. Thai mythai@cise. ufl. edu 166
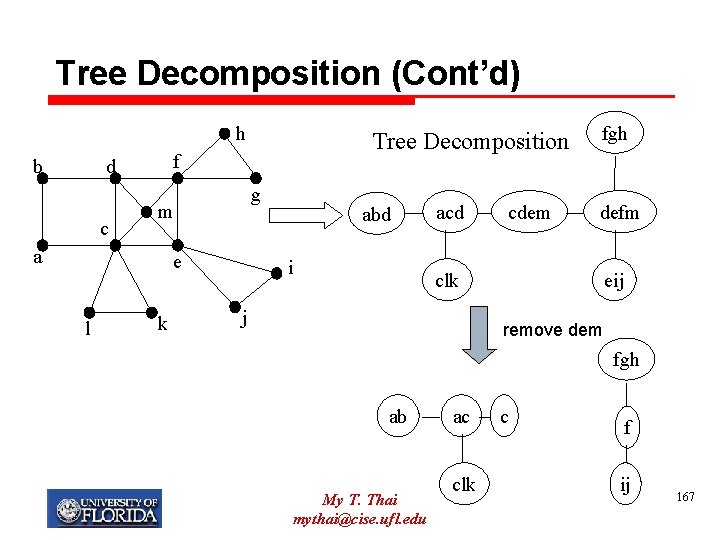
Tree Decomposition (Cont’d) h b f d c g m a e l Tree Decomposition k abd i acd cdem fgh defm clk j eij remove dem fgh ab My T. Thai mythai@cise. ufl. edu ac clk c f ij 167

Tree Decomposition-Based Algorithms Xir Xr Xi Xp Xq 1. Bottom-to-Top: Calculate the minimal F function Xji Xj Xl Xli 2. Top-to-Bottom: Extract the optimal assignment A tree decomposition rooted at Xr The score of subtree rooted at Xi The score of component Xi The scores of subtree rooted at Xl The scores of subtree rooted at Xj My T. Thai mythai@cise. ufl. edu 168
- Slides: 168