Computational identification and experimental validation of PPRE motifs














- Slides: 28

Computational identification and experimental validation of PPRE motifs in NHE 1 and Mn. SOD genes of Human Gireedhar V; Kumar AP; Loo SY; Pervaiz S; Clement MV; and Sakharkar MK Presenting author: Gireedhar Venkatachalam International Conference on Bioinformatics (In. Co. B)

Brief overview 1) Introduction to PPAR (Peroxisome Proliferator Activated Receptor) and PPRE (Peroxisome Proliferator Response elements) 2) Aims and PPRE prediction 3) PPAR , NHE 1 and Mn. SOD in Breast cancer 4) Computational Prediction of PPREs in NHE 1 and Mn. SOD 5) Experimental Validation of PPREs in NHE 1 and Mn. SOD 6) Conclusion

Brief overview 1) Introduction to PPAR (Peroximsome Proliferator Activated Receptor) and PPRE (Peroximsome Proliferator Response elements) 2) Aims and PPRE prediction 3) PPAR , NHE 1 and Mn. SOD in breast cancer 4) Computational Prediction of PPREs in NHE 1 and Mn. SOD 5) Experimental Validation of PPREs in NHE 1 and Mn. SOD 6) Conclusion

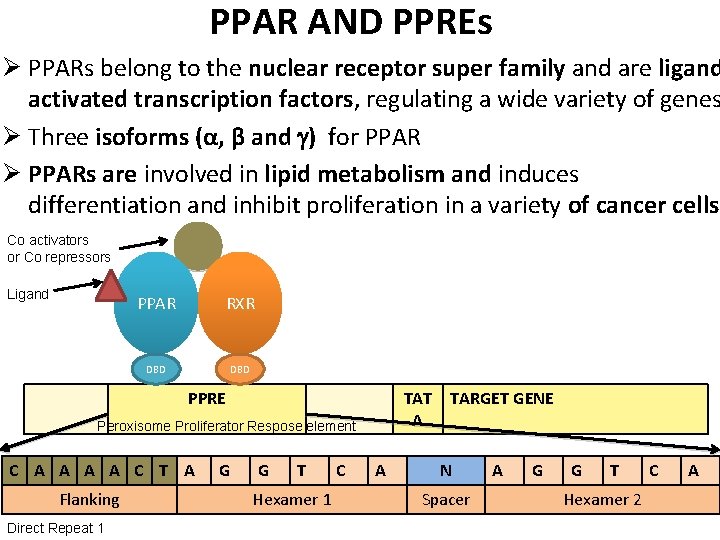
PPAR AND PPREs Ø PPARs belong to the nuclear receptor super family and are ligand activated transcription factors, regulating a wide variety of genes Ø Three isoforms (α, β and ) for PPAR Ø PPARs are involved in lipid metabolism and induces differentiation and inhibit proliferation in a variety of cancer cells Co activators or Co repressors Ligand PPAR RXR DBD PPRE TAT TARGET GENE A Peroxisome Proliferator Respose element C A A C T A Flanking Direct Repeat 1 G G T Hexamer 1 C A N Spacer A G G T Hexamer 2 C A

Brief overview 1 ) Introduction to PPAR (Peroximsome Proliferator Activated Receptor) and PPRE (Peroximsome Proliferator Response elements) 2) Aims and PPRE prediction 3) PPAR , NHE 1 and Mn. SOD-Breast cancer 4) Computational Prediction of PPREs in NHE 1 and Mn. SOD 5) Experimental Validation of PPREs in NHE 1 and Mn. SOD 6) Conclusion

Aim of the study • Recently it is shown that PPARs also bind to DR 2 repeats (AGGTCA NN AGGTCA) (Fontaine et al. , 2003; AP Kumar et al. , 2004) • Flanking sequence plays a significant role in PPARs specificity and binding (Palmer et al. , 1995) • Nuclear receptor- competitive binders (Harikrishna et al. , 1998)

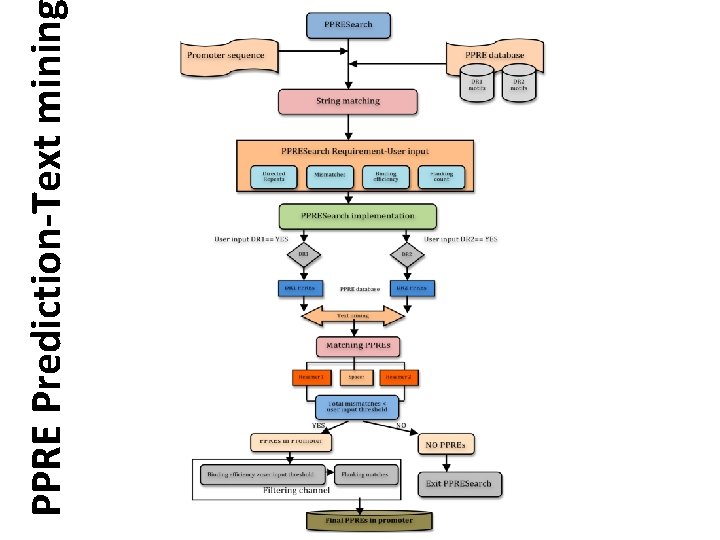
PPRE Prediction • Collection of PPRE database Ø Contains 414 reported PPRE motifs from literature. ØThe sequences reported only with experimental validation were added to this database. ØPPRE element in database - reported consensus, isoform specificity, in vivo and in vitro binding efficiencies and Pubmed IDs

PPRE Prediction-Text minin

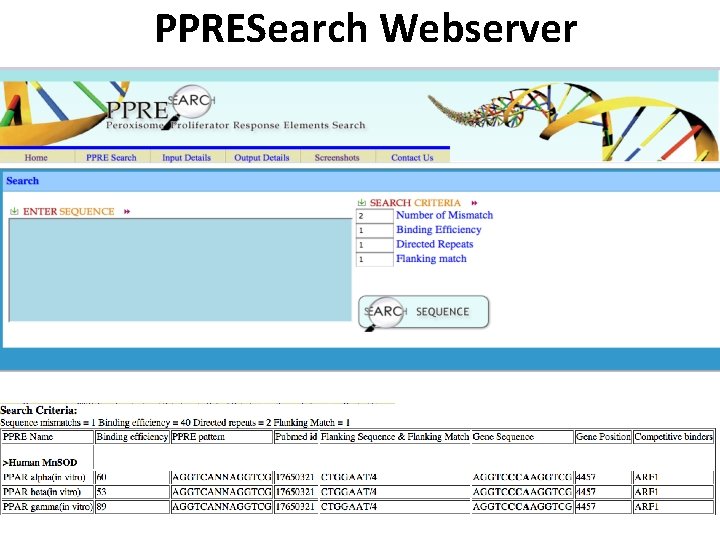
PPRESearch Webserver

Brief overview 1) Introduction to PPAR (Peroximsome Proliferator Activated Receptor) and PPRE (Peroximsome Proliferator Response elements) 2) PPRE prediction 3) PPAR , NHE 1 and Mn. SOD-Breast cancer 4) Computational Prediction of PPREs in NHE 1 and Mn. SOD 5) Experimental Validation of PPREs in NHE 1 and Mn. SOD 6) PPAR as novel therapeutic approach in breast cancer therapy

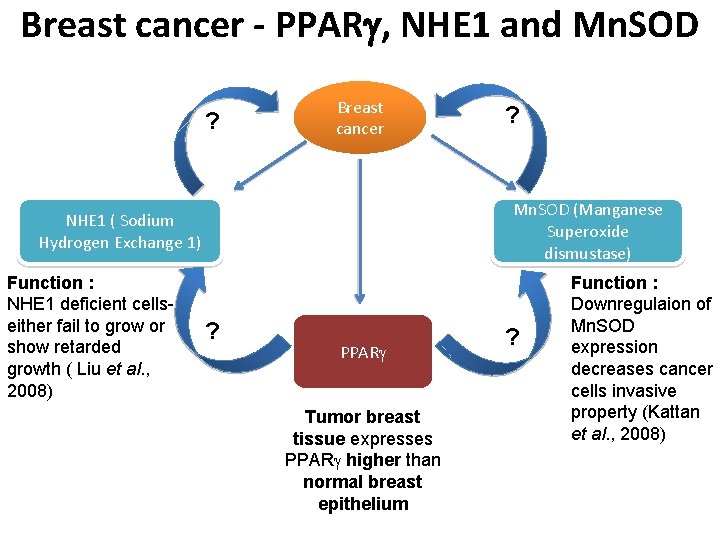
Breast cancer - PPAR , NHE 1 and Mn. SOD ? Breast cancer Mn. SOD (Manganese Superoxide dismustase) NHE 1 ( Sodium Hydrogen Exchange 1) Function : NHE 1 deficient cellseither fail to grow or show retarded growth ( Liu et al. , 2008) ? ? PPAR Tumor breast tissue expresses PPAR higher than normal breast epithelium ? Function : Downregulaion of Mn. SOD expression decreases cancer cells invasive property (Kattan et al. , 2008)

Brief overview 1) Introduction to PPAR (Peroximsome Proliferator Activated Receptor) and PPRE (Peroximsome Proliferator Response elements) 2) PPRE prediction 3) Breast cancer-PPAR , NHE 1 and Mn. SOD 4) Computational Prediction of PPREs in NHE 1 and Mn. SOD 5) Experimental Validation of PPREs in NHE 1 and Mn. SOD 6) Conclusion

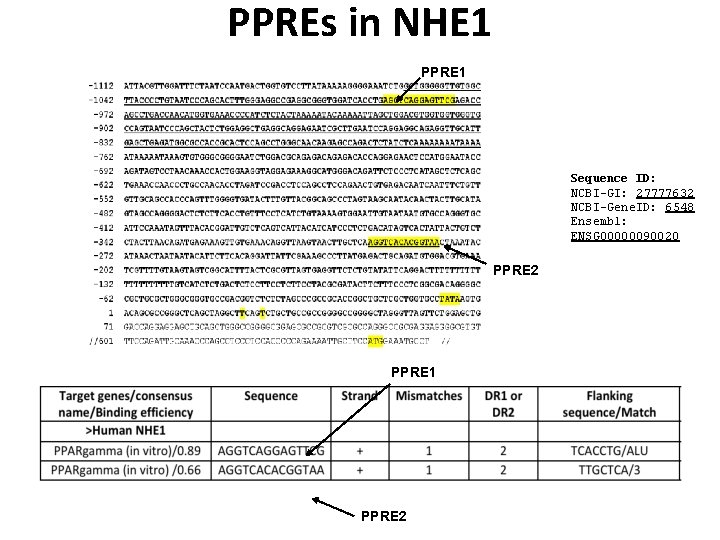
PPREs in NHE 1 PPRE 1 Sequence ID: NCBI-GI: 27777632 NCBI-Gene. ID: 6548 Ensembl: ENSG 00000090020 PPRE 2 PPRE 1 PPRE 2

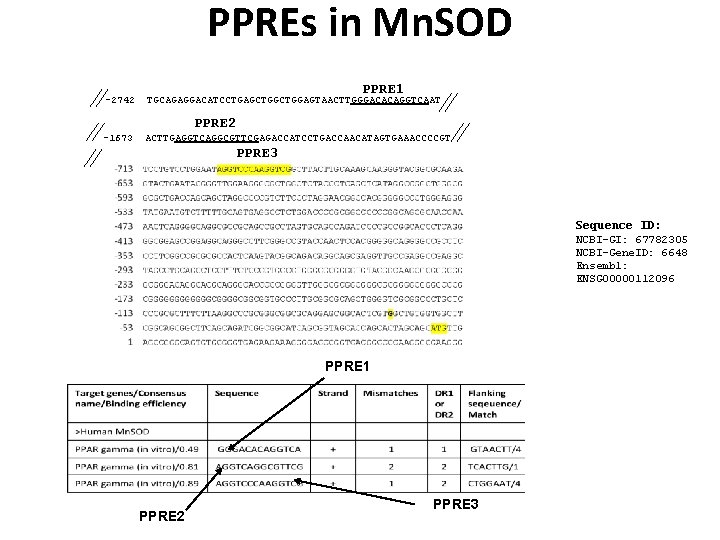
PPREs in Mn. SOD -2742 PPRE 1 TGCAGAGGACATCCTGAGCTGGAGTAACTTGGGACACAGGTCAAT PPRE 2 -1673 ACTTGAGGTCAGGCGTTCGAGACCATCCTGACCAACATAGTGAAACCCCGT PPRE 3 Sequence ID: NCBI-GI: 67782305 NCBI-Gene. ID: 6648 Ensembl: ENSG 00000112096 PPRE 1 PPRE 2 PPRE 3

Brief overview 1) Introduction to PPAR (Peroximsome Proliferator Activated Receptor) and PPRE (Peroximsome Proliferator Response elements) 2) PPRE prediction 3) Breast cancer-PPAR , NHE 1 and Mn. SOD 4) Computational Prediction of PPREs in NHE 1 and Mn. SOD 5) Experimental Validation of PPREs in NHE 1 and Mn. SOD 6) Conclusion

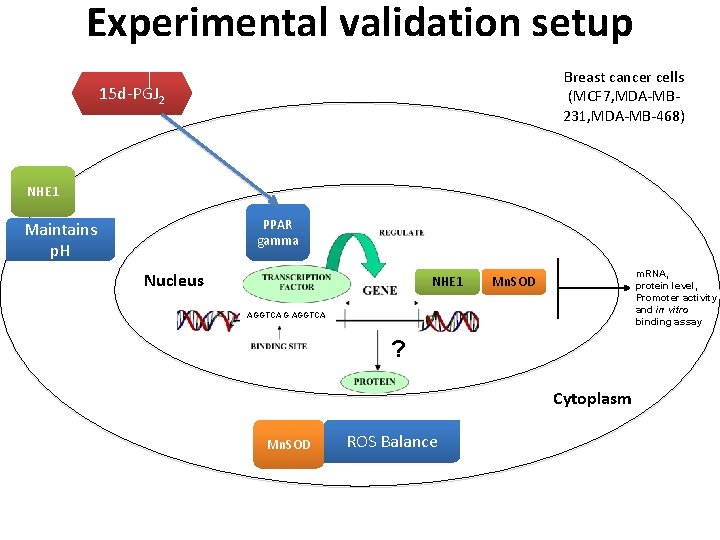
Experimental validation setup Breast cancer cells (MCF 7, MDA-MB 231, MDA-MB-468) 15 d-PGJ 2 NHE 1 PPAR gamma Maintains p. H Nucleus NHE 1 m. RNA, protein level, Promoter activity and in vitro binding assay Mn. SOD AGGTCA G AGGTCA ? Cytoplasm Mn. SOD ROS Balance

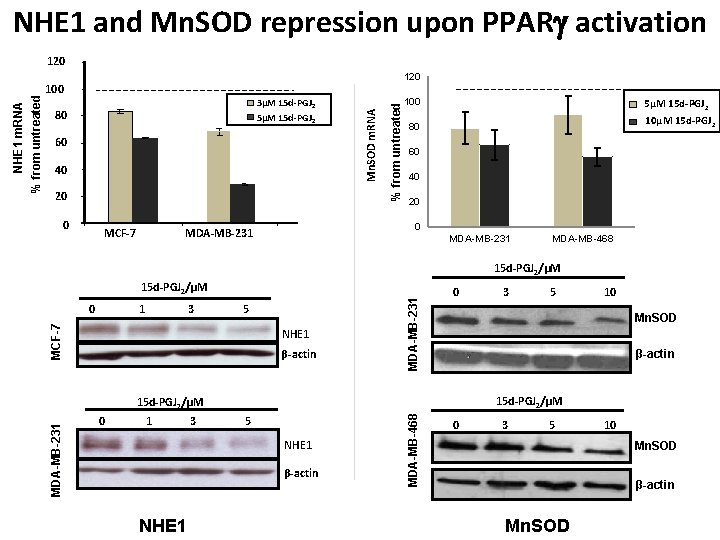
NHE 1 and Mn. SOD repression upon PPAR activation 120 60 40 20 0 MCF-7 % from untreated 3µM 15 d-PGJ 2 5µM 15 d-PGJ 2 80 Mn. SOD m. RNA 100 5µM 15 d-PGJ 2 10µM 15 d-PGJ 2 80 60 40 20 0 MDA-MB-231 MDA-MB-468 15 d-PGJ 2/µM 1 3 5 MCF-7 0 NHE 1 β-actin MDA-MB-231 15 d-PGJ 2/µM 0 1 3 5 NHE 1 β-actin NHE 1 5 10 β-actin 15 d-PGJ 2/µM MDA-MB-468 0 3 Mn. SOD 15 d-PGJ 2/µM MDA-MB-231 NHE 1 m. RNA % from untreated 120 0 3 5 10 Mn. SOD β-actin Mn. SOD

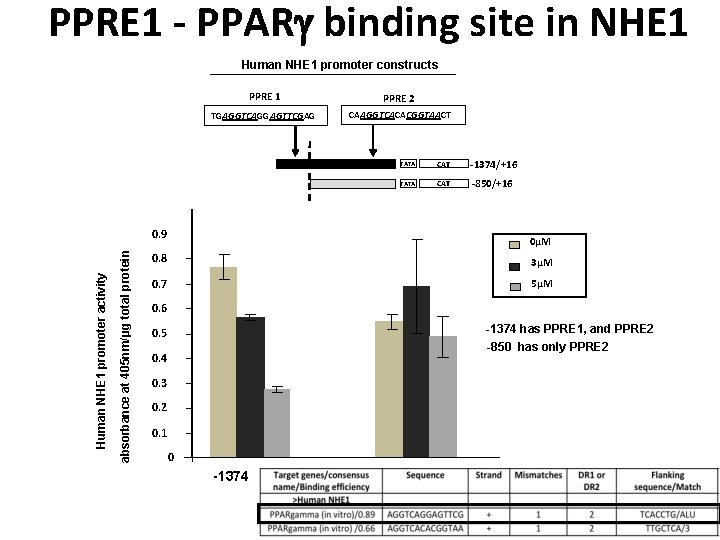
PPRE 1 - PPAR binding site in NHE 1 Human NHE 1 promoter constructs PPRE 1 PPRE 2 TGAGGTCAGGAGTTCGAG CAAGGTCACACGGTAACT TATA CAT -1374/+16 TATA CAT -850/+16 absorbance at 405 nm/μg total protein Human NHE 1 promoter activity 0. 9 0µM 0. 8 3µM 0. 7 5µM 0. 6 -1374 has PPRE 1, and PPRE 2 0. 5 -850 has only PPRE 2 0. 4 0. 3 0. 2 0. 1 0 -1374 -850

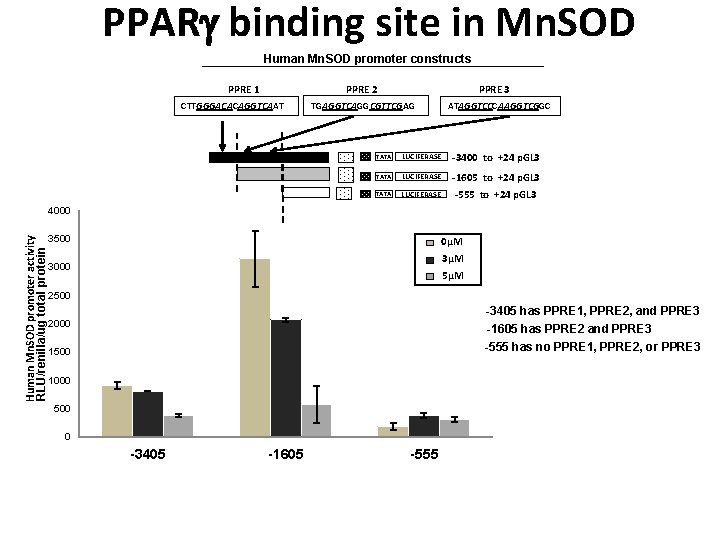
PPAR binding site in Mn. SOD Human Mn. SOD promoter constructs PPRE 1 PPRE 3 PPRE 2 CTTGGGACACAGGTCAAT TGAGGTCAGGCGTTCGAG ATAGGTCCCAAGGTCGGC TATA LUCIFERASE -3400 to +24 p. GL 3 TATA LUCIFERASE -1605 to +24 p. GL 3 TATA LUCIFERASE -555 to +24 p. GL 3 Human Mn. SOD promoter activity RLU/renilla/ug total protein 4000 3500 0µM 3000 3µM 5µM 2500 -3405 has PPRE 1, PPRE 2, and PPRE 3 2000 -1605 has PPRE 2 and PPRE 3 -555 has no PPRE 1, PPRE 2, or PPRE 3 1500 1000 500 0 -3405 -1605 -555

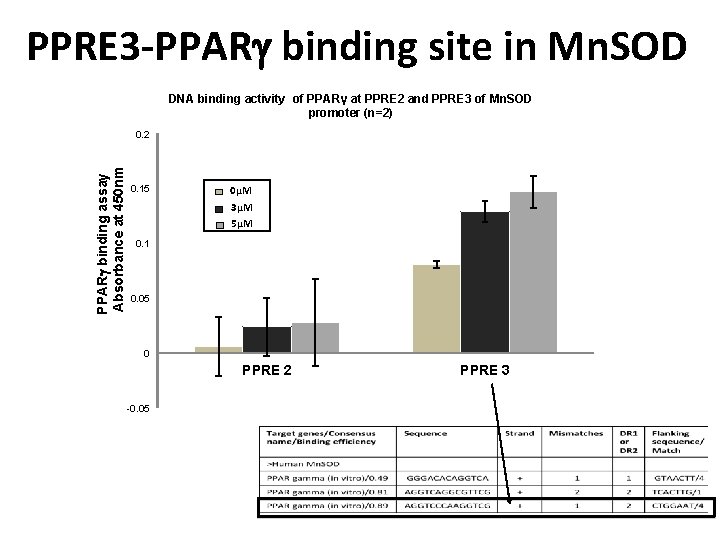
PPRE 3 -PPAR binding site in Mn. SOD DNA binding activity of PPARγ at PPRE 2 and PPRE 3 of Mn. SOD promoter (n=2) PPAR binding assay Absorbance at 450 nm 0. 2 0. 15 0µM 3µM 5µM 0. 1 0. 05 0 PPRE 2 -0. 05 PPRE 3

Brief overview 1) Introduction to PPAR (Peroximsome Proliferator Activated Receptor) and PPRE (Peroximsome Proliferator Response elements) 2) PPRE prediction 3) Breast cancer-PPAR , NHE 1 and Mn. SOD 4) Computational Prediction of PPREs in NHE 1 and Mn. SOD 5) Experimental Validation of PPREs in NHE 1 and Mn. SOD 6) Conclusion

Conclusion • We have constructed a better in silico approach to finding genes containing PPRE in their promoter region • Our approach helps us to identify both DR 1 and DR 2 sites • Importance of flanking sequence were incorporated. • It is our hope with this PPRESearch database, researchers in the field of PPARs would better identify new target genes which could then be translated into the clinic for intervention.

Thank you

Questions?

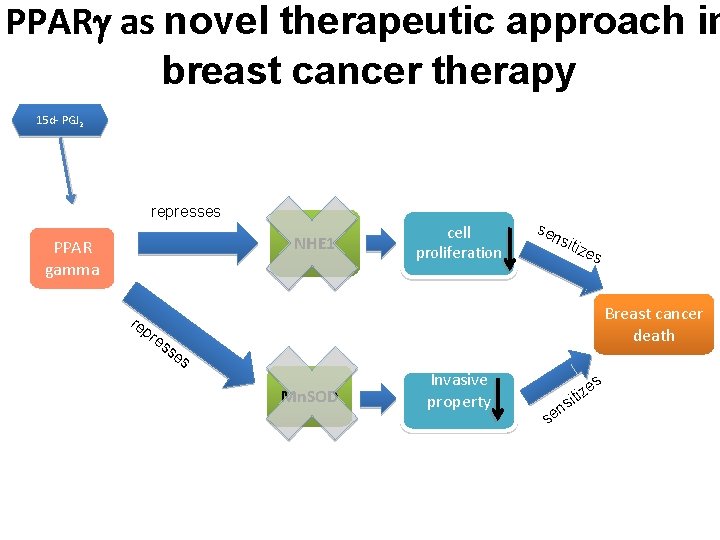
PPAR as novel therapeutic approach in breast cancer therapy 15 d- PGJ 2 represses NHE 1 PPAR gamma re pr es cell proliferation sen siti zes Breast cancer death se s Mn. SOD Invasive property ns e s es z i it

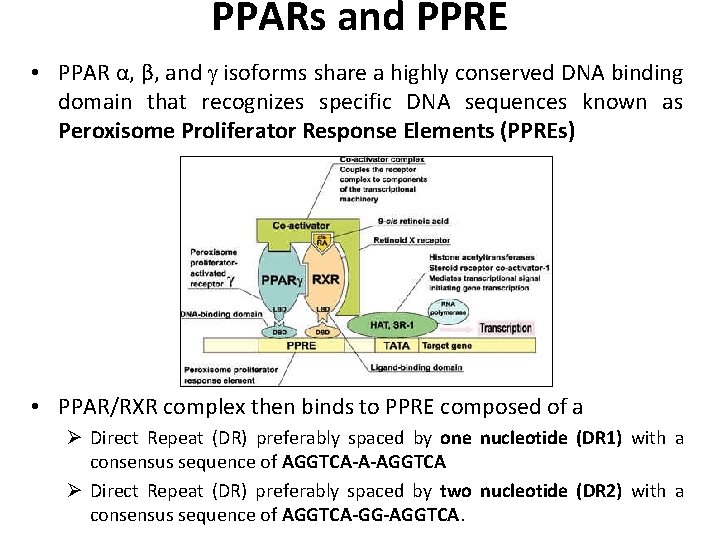
PPARs and PPRE • PPAR α, β, and isoforms share a highly conserved DNA binding domain that recognizes specific DNA sequences known as Peroxisome Proliferator Response Elements (PPREs) • PPAR/RXR complex then binds to PPRE composed of a Ø Direct Repeat (DR) preferably spaced by one nucleotide (DR 1) with a consensus sequence of AGGTCA-A-AGGTCA Ø Direct Repeat (DR) preferably spaced by two nucleotide (DR 2) with a consensus sequence of AGGTCA-GG-AGGTCA.

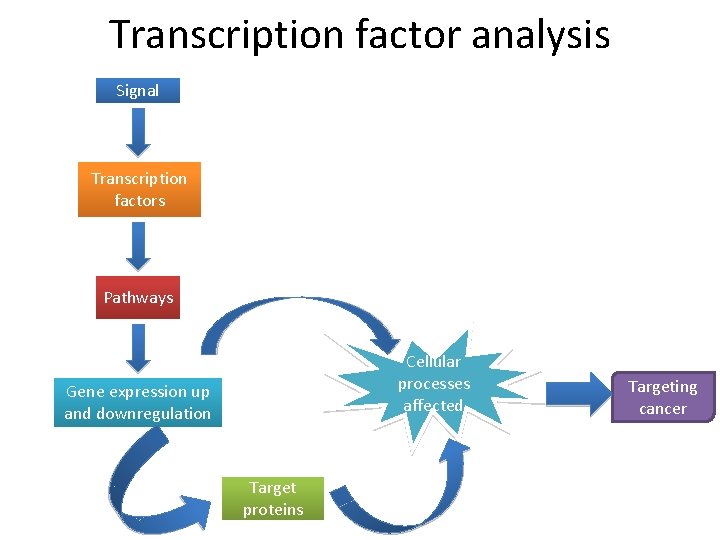
Transcription factor analysis • Transcription factor might be defined as any molecule participating, alone or as part of a complex, in the binding to a gene’s enhancer response element or promoter, with the ultimate outcome being the up- or downregulation of expression of that gene. • Transcription factors participate in stress pathways in cancer by causing the up- or down-regulation of specific genes.

Transcription factor analysis Signal Transcription factors Pathways Cellular processes affected Gene expression up and downregulation Target proteins Targeting cancer