Composite Segmentation 1 Objectives We have two segmentation

Composite Segmentation 1

Objectives • We have two segmentation methods. – Previous work has identified that these generate similar qualitative states but are different in segment length and transition frequency. • We would like to summarise these in the Integrated analysis paper to provide statistics on the genome. • We would like to provide a summary “Encode view” of the genome for users. 2

Previous • Bake-off – Calculate Precision-Recall type stats for relevant data of biological interest – Look at relationship to hold out data e. g. methylation, RNA – Biologist look and feel test 3
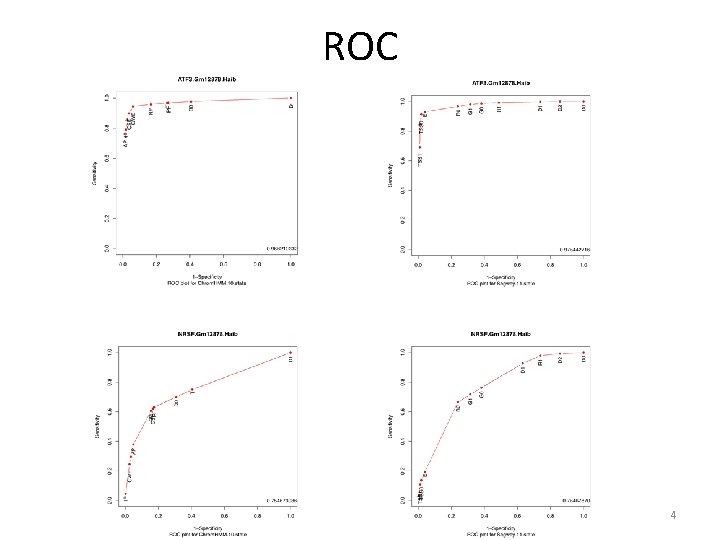
ROC 4
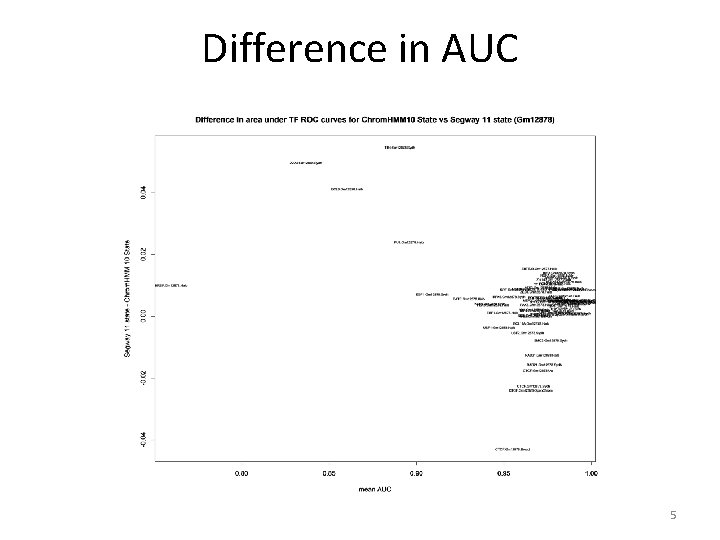
Difference in AUC 5
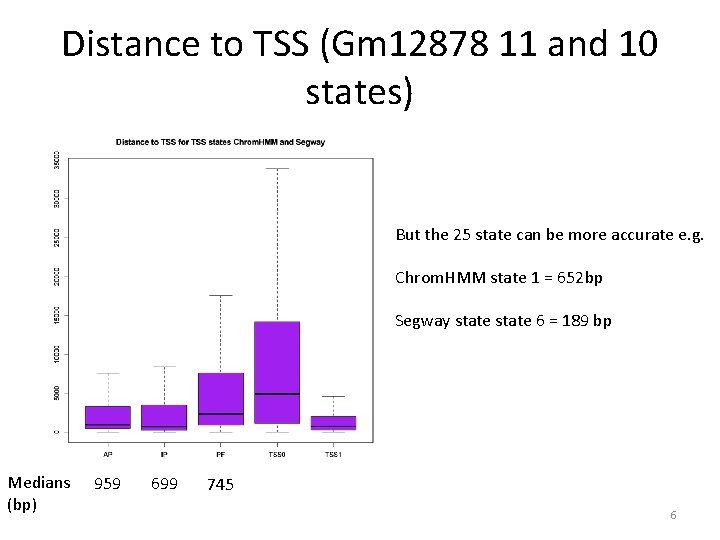
Distance to TSS (Gm 12878 11 and 10 states) But the 25 state can be more accurate e. g. Chrom. HMM state 1 = 652 bp Segway state 6 = 189 bp Medians (bp) 959 699 745 6

User/Browser views • Purely on a vote count there was preference for 10/11 states but no choice between Chrom. HMM/Segway • A consistent comment is that Segway is choppy but is better defined around interesting known regions. People like the continuity of Chrom. HMM, but say it lacks some resolution at high zooms for sites of interest (enhancers, other elements, but also TSS). • There are things missed by both or by one or the other. • There is at least one criticism that says neither are good enough because they don’t tag enhancers well enough. • There is support to use both as they have different advanteages 7

Decision • We use both and report statistics on a pick and choose basis. We would need to explain well to the reader what we are doing and how we get our figures. • This has some support and removes the need for a bake-off. 8
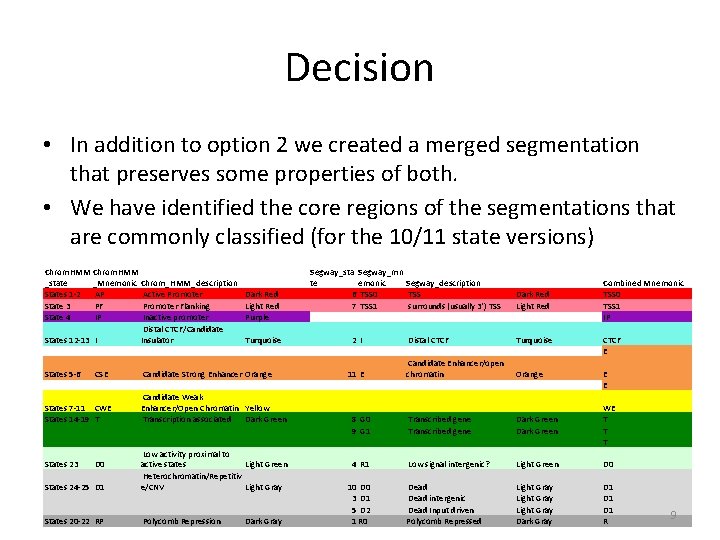
Decision • In addition to option 2 we created a merged segmentation that preserves some properties of both. • We have identified the core regions of the segmentations that are commonly classified (for the 10/11 state versions) Chrom. HMM _state _Mnemonic Chrom_HMM_description States 1 -2 AP Active Promoter State 3 PF Promoter Flanking State 4 IP Inactive promoter Distal CTCF/Candidate States 12 -13 I Insulator States 5 -6 CSE States 7 -11 CWE States 14 -19 T Dark Red Light Red Purple Turquoise Candidate Strong Enhancer Orange Candidate Weak Enhancer/Open Chromatin Yellow Transcription associated Dark Green States 24 -25 D 1 Low activity proximal to active states Light Green Heterochromatin/Repetitiv e/CNV Light Gray States 20 -22 RP Polycomb Repression States 23 D 0 Dark Gray Segway_sta Segway_mn te emonic Segway_description 6 TSS 0 TSS 7 TSS 1 surrounds (usually 3') TSS 2 I Dark Red Light Red Combined Mnemonic TSS 0 TSS 1 IP Distal CTCF Turquoise CTCF E Candidate Enhancer/open chromatin Orange E E 8 G 0 9 G 1 Transcribed gene Dark Green WE T T T 4 R 1 Low signal intergenic? Light Green D 0 Dead intergenic Dead Input driven Polycomb Repressed Light Gray Dark Gray D 1 D 1 R 11 E 10 D 0 3 D 1 5 D 2 1 R 0 9

Approach • Establish equivalence matrix of states and use this to include bases where equivalences occur in the merged segmentation. 10
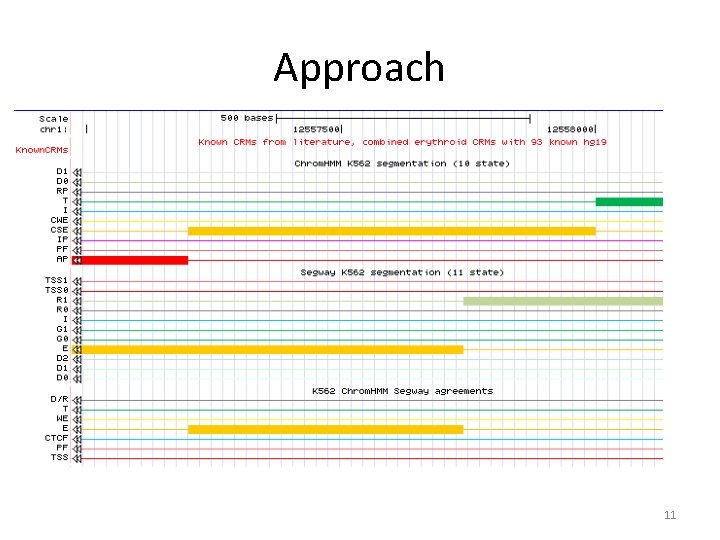
Approach 11

Initial Merge (v 1) Cell line Bases in Core Combined Segmentation Gm 12878 H 1 hesc Helas 3 Hepg 2 Huvec K 562 1, 746, 359, 416 (61. 7%) 1, 375, 544, 206 (48. 6%) 1, 090, 294, 321 (38. 5%) 1, 703, 449, 033 (60. 2%) 1, 585, 992, 714 (56. 1%) 1, 941, 411, 363 (68. 6%) Comments: • Coverage is not good enough, and variable between cell lines. • Manual inspection and confusion matrix indicates there additional equivalences we want to capture 12
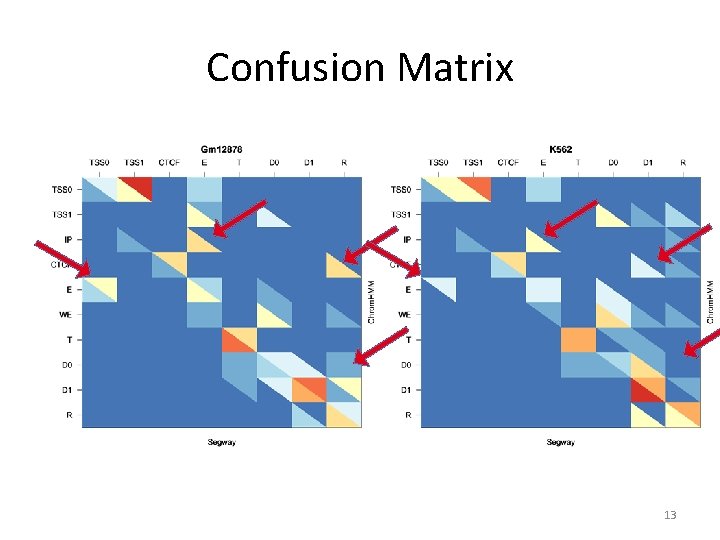
Confusion Matrix 13
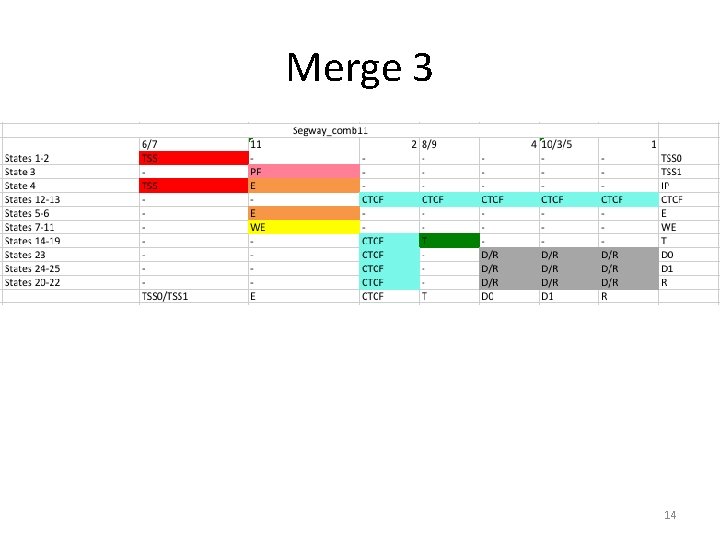
Merge 3 14

Merge v 3 Cell line Bases in Segmentation Gm 12878 H 1 hesc Helas 3 Hepg 2 Huvec K 562 2, 532, 210, 925 (89. 5%) 2, 505, 876, 348 (88. 6%) 2, 309, 076, 343 (81. 6%) 2, 509, 784, 276 (87. 9%) 2, 281, 643, 167 (80. 6%) 2, 543, 419, 766 (89. 9%) • Thisis the version that was circulated to the AWG list, and the following analysis refers to. 15
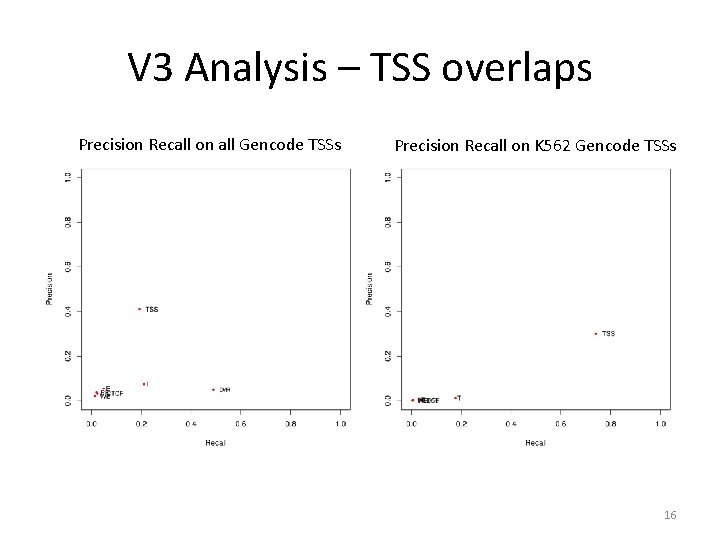
V 3 Analysis – TSS overlaps Precision Recall on all Gencode TSSs Precision Recall on K 562 Gencode TSSs 16
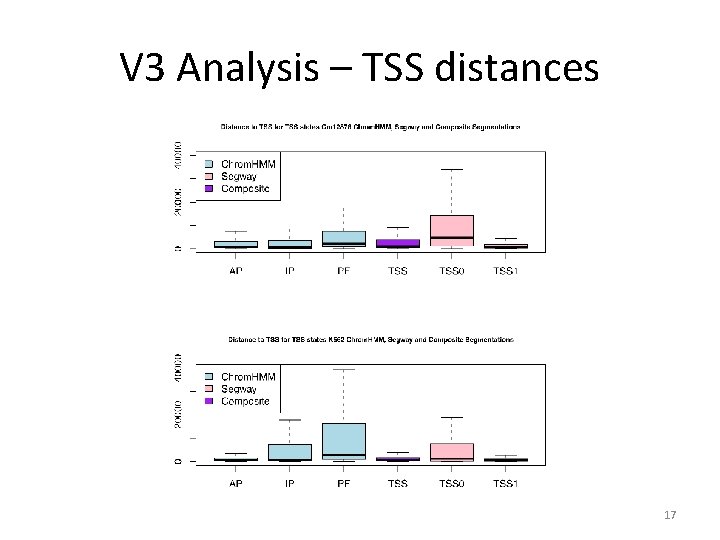
V 3 Analysis – TSS distances 17
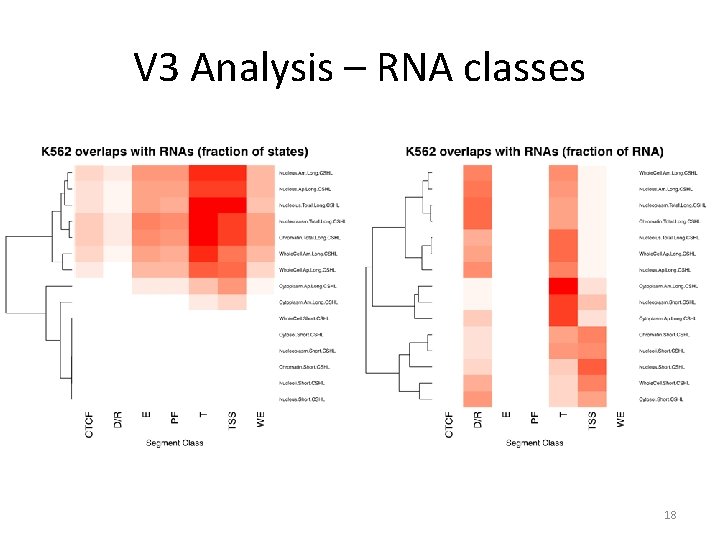
V 3 Analysis – RNA classes 18
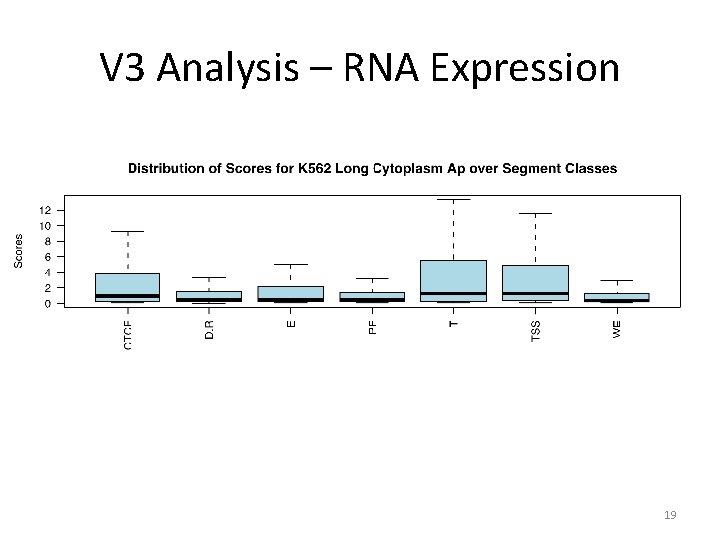
V 3 Analysis – RNA Expression 19
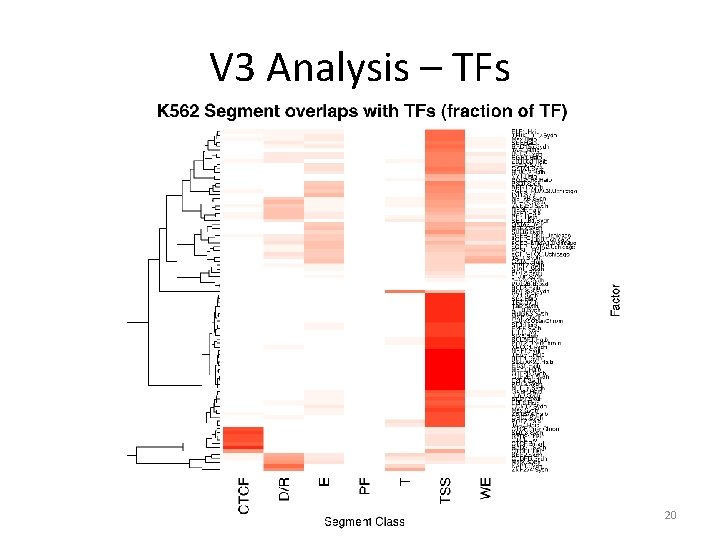
V 3 Analysis – TFs 20

Biologist look and feel test. • http: //encodewiki. ucsc. edu/Encode. DCC/index. php/Segmenta tion_bake_off#Composite_Segmentation 21
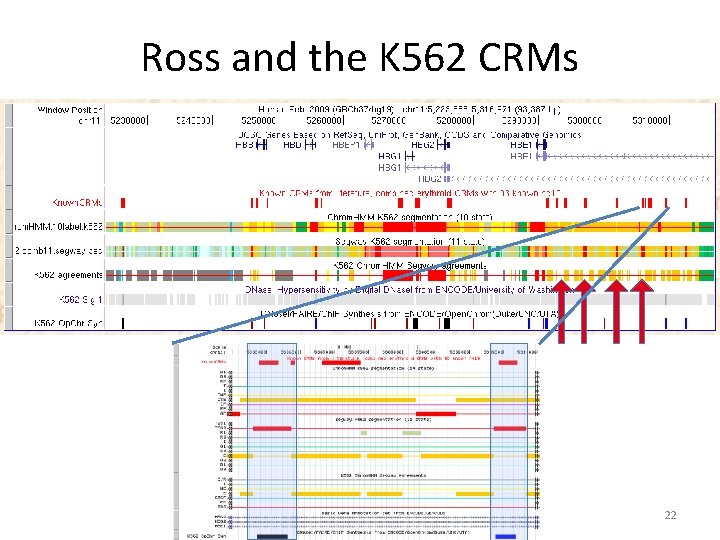
Ross and the K 562 CRMs 22
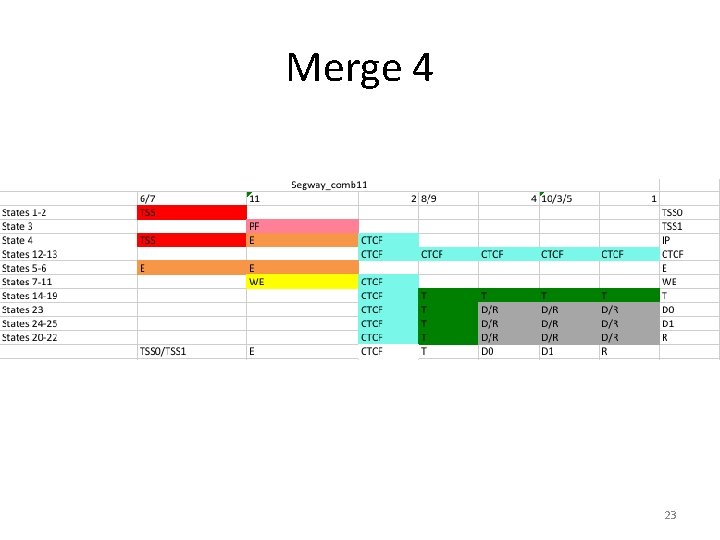
Merge 4 23
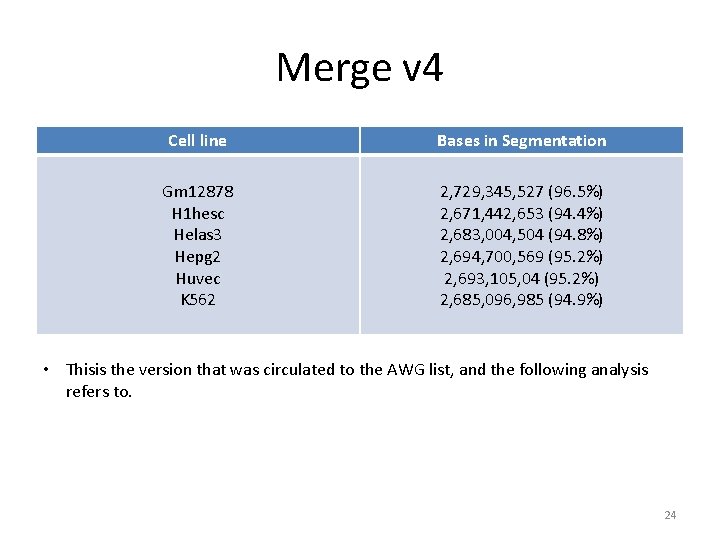
Merge v 4 Cell line Bases in Segmentation Gm 12878 H 1 hesc Helas 3 Hepg 2 Huvec K 562 2, 729, 345, 527 (96. 5%) 2, 671, 442, 653 (94. 4%) 2, 683, 004, 504 (94. 8%) 2, 694, 700, 569 (95. 2%) 2, 693, 105, 04 (95. 2%) 2, 685, 096, 985 (94. 9%) • Thisis the version that was circulated to the AWG list, and the following analysis refers to. 24
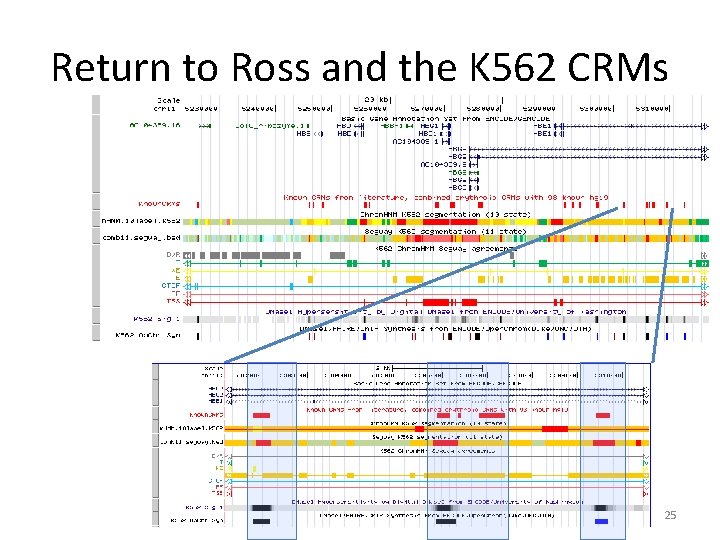
Return to Ross and the K 562 CRMs 25

Names • Core – suggests intersection • Merge – describes the process • Composite – implying a more complex process, perhaps? 26
- Slides: 26