Comparing Cancer Risks between Radiation and Dioxin Exposure

Comparing Cancer Risks between Radiation and Dioxin Exposure Based on Two-Stage Model Tsuyoshi Nakamura Faculty of Environmental Studies, Nagasaki University David G. Hoel Dept. of Biometry and Epidemiology, Medical University of South Carolina 1. Two-Stage Model 2. Historical Aspects 3. Estimation Method 4. Radiation by JANUS 5. Dioxin by Kociba Summary Conclusion 1

Two-Stage Model Three States of Cells Normal, Intermediate and Malignant N I M Four Parameters for Rates m 1 : First Mutation Rate for N I b : Clonal Expansion Rate for I d : Death Rate for I m 2 : Second Mutation Rate for I M 2

History Mathematical tool based on Molecular biology to study Mechanistic processes in Cancer development (Moolgavkar, Venzon, Knudson, 70’s) Special Feature Explicit modeling of Clonal expansion, Differentiation and Mutation of I-cells as a Continuous Stochastic Process Cancer Incidence Data (time, type, covariate) (t, 1, x): endpoint (t, 0, x): censored 3

Problems Unidentifiability All parameters are not identifiable. Reparameterization or Assumption is necessary. Non-Convergence MLE of the identifiable parameters are still often hardly obtainable, because of the peculiar shape of the likelihood surface (Portier et at. 1997). Non-Standard Algorithm Lack of Confidence in Results Lack of Comparison among Studies 4

Survivor function S(t) Probability of No Malignant Cell at t, is obtained by solving a series of differential equations, derived from Stochastic processes on Probability Generating Function (Moolgavkar et al 1990; Kopp et al 1994; Portier et al 1996; ) Stochastic processes M(t)=(x(t), y(t), z(t)) denote the number of N-, I- and M-cells at t, respectively. M(t): Continuous Markov Birth-Death process S(t)= Si, j. Prob{M(t)=(i, j, 0)} 5

Probability Generating Function P(i, j, k|t)=Prob{M(t)=(i, j, k) | M(0)=(1, 0, 0) } G(u, v, w|t)= Si, j, k P(i, j, k|t)uivjwk and Q(i, j, k|t)=Prob{M(t)=(i, j, k) | M(0)=(0, 1, 0) } H(u, v, w|t)= Si, j, k Q(i, j, k|t)uivjwk S(t)=Si, j. P(i, j, 0|t)=G(1, 1, 0|t) Differential Equations It follows that (Portier et al 1996) d. G(t)/dt= m 1 G(t)H(t)-m 1 G (t) d. H(t)/dt= b. H(t)2+d-(b+d+m 2)H(t) G(0)=1, H(0)=1 6

Survivor Function S(t) X 0 = Number of N-cells, Large and Constant u = N I Rate per Cell per unit Time ==> m 1=u. X 0 S(t)=exp{-L(t)}, L(t)=h{t(R+y)/2+log[{R-y+(R+y)e-Rt}/2 R]} L is Cumulative Hazard with new parameters h=m 1/b, y=b-d-m 2 and R 2=(b-d-m 2)2+4 bm 2 Original likelihood y: Net Proliferation Rate r=m 1 m 2=h(R 2 -y 2)/4: Overall Mutation Rate l(h, y, r) based on L(t) is termed Original likelihood. Non-convergence is frequent ! 7
![Conditional likelihood Put d=0 then L(t|d=0)= [ ] m*1, b* and m*2 are employed Conditional likelihood Put d=0 then L(t|d=0)= [ ] m*1, b* and m*2 are employed](http://slidetodoc.com/presentation_image_h2/0765ef221b38d1f6e3aad7d7d0bbbf0c/image-8.jpg)
Conditional likelihood Put d=0 then L(t|d=0)= [ ] m*1, b* and m*2 are employed to emphasize these parameters are valid only when d=0 l(m*1, b*, m*2 ) based on L(t|d=0) is termed Conditional likelihood. Looks Better Shape! 8

Transformation Conditional likelihood converges better! Biological interpretation of parameters is ? It ignores the death of the I-cells. Biological parameters estimated by m*1, b* and m*2 are h=m*1/b*, y=b*-m*2 and r=m*1 m*2 (Nakamura and Hoel 2002) Thus, MLE of h, y and r are obtained from Conditional Likelihood ! Practically y=b*, since m*2 is small 9

Comparison on Experimental Data Conditional vs Original JANUS data for Radiation Risk study On g and Neutron in Mice Argonne National Laboratory (1953 -1970 ) Reliable Pathological information Kociba data for Chronic Toxicity study on TCDD in Rats Dow Chemical (1978) Reliable Pathological information 10
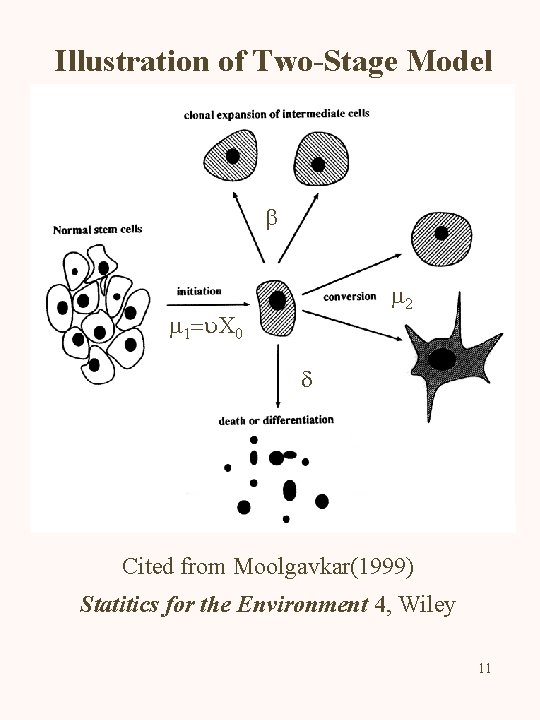
Illustration of Two-Stage Model b m 2 m 1=u. X 0 d Cited from Moolgavkar(1999) Statitics for the Environment 4, Wiley 11
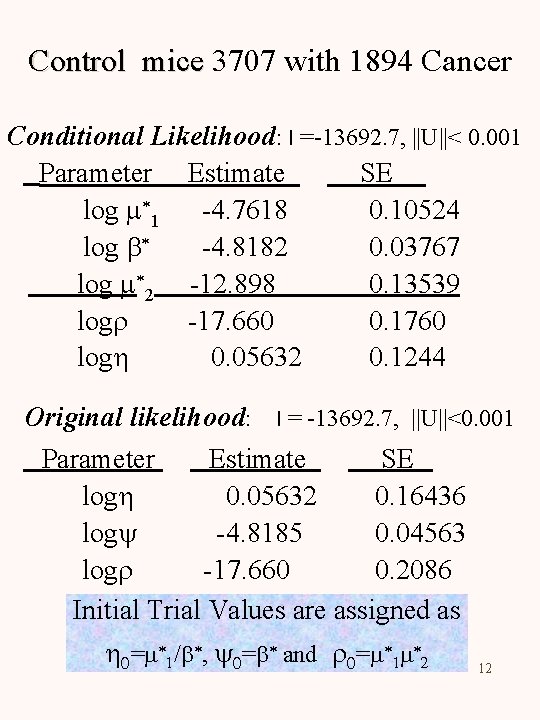
Control mice 3707 with 1894 Cancer Conditional Likelihood: l =-13692. 7, ||U||< 0. 001 Parameter Estimate SE log m*1 -4. 7618 0. 10524 log b* -4. 8182 0. 03767 log m*2 -12. 898 0. 13539 logr -17. 660 0. 1760 logh 0. 05632 0. 1244 Original likelihood: l = -13692. 7, ||U||<0. 001 Parameter Estimate SE logh 0. 05632 0. 16436 logy -4. 8185 0. 04563 logr -17. 660 0. 2086 Initial Trial Values are assigned as h 0=m*1/b*, y 0=b* and r 0=m*1 m*2 12

Regression Model logq=a+b. Dose (Contol + g) 7402 mice with 4133 Cancer Conditional. Likelihood: l=-29446. 65, ||U||=0. 002 Const. a (SE) logm*1 -4. 931 (0. 0817) logb* -4. 851 (0. 0278) logm*2 -12. 43 (0. 1014) logr -17. 37 (0. 1181) Slope b (SE) 0. 00717 (0. 00115) -0. 000345 (0. 000071) -0. 002934 (0. 001126) 0. 00424 (0. 000241) ------------------------------------------------------------------------- Original likelihood: l=-29446. 69, ||U||=0. 2542 Const. a (SE) Slope b (SE) logh -0. 0797(0. 1318) 0. 00749 (0. 00131) logy -4. 852 (0. 0373) 0. 000345 (0. 000077) logr -17. 37 (0. 1536) 0. 00424 (0. 000266) All Estimates are of p<0. 01 13

Effect of Exposure on Mutation and Promotion 1) r=m 1 m 2=u. X 0 m 2 2) X 0 is Constant not affected by Exposure 3) Effect of exposure on u and that on m 2 are the same ( Moolgavkar et al , 1999), 4) logr=a+b. Dose ==> Dose effect on u and that on m 2 is b/2 Dose Effect on Mutation Rate and Net proliferation Rate may be obtained from Conditional likelihood without Additional Assumption! 14

Log Cumulative Hazards Two-Stage (H) vs K-M(V) Dose 0 : Subjects 3707 Cancer 1894 V H 15

Log Cumulative Hazards Two-Stage (H) vs K-M(V) Dose 86 : Subjects 1376 Cancer 960 V H 16

Log Cumulative Hazards Two-Stage (H) vs K-M(V) Dose 756 : Subjects 396 Cancer 190 V H 17

Regression Coefficients for Dioxin 205 rats, 31 cancer, logq=a+blog(1+Dose) Conditional Likelihood: l = -206. 77, ||U||=0. 0004 Const. SE Slope SE logm*1 -3. 780 0. 7075 non logb* -3. 961 0. 1062 0. 0680 0. 01497 logm*2 -20. 82 1. 371 non logr -24. 60 1. 259 non Original Likelihood: l = -207. 012, ||U||= 0. 0012 Const. a SE Slope SE logh 0. 0865 0. 8192 non logy -3. 979 0. 1083 0. 0658 0. 01466 logr -24. 32 1. 216 non Original Likelihood: l =-207. 585, ||U||=5. 7489 Incomplete-convergent case Const. SE Slope SE logh -0. 4724 0. 7395 non logy -3. 773 0. 04589 0. 0631 0. 0200 logr -27. 14 0. 3368 non 18
Log Cumulative Hazards for Dioxin Doses week 100 10 1 0 19

Log Cumulative Hazards for Radiation Doses week 756 400 197. 6 86. 31 43. 15 0 20
- Slides: 20