Comparative genomics and proteomics in Ensembl Sep 2006

Comparative genomics and proteomics in Ensembl Sep 2006

Overview • Rationale • Species available • Comparative proteomics – Orthologue and paralogue prediction – Protein clustering into families • Comparative genomics – Genome-wide DNA alignments – Synteny block characterisation • Future and perspectives 2 of 56

Compara The Compara database is one single multispecies database • Gene orthology/paralogy prediction • Protein clustering • Whole genome alignments • Synteny regions 3 of 56
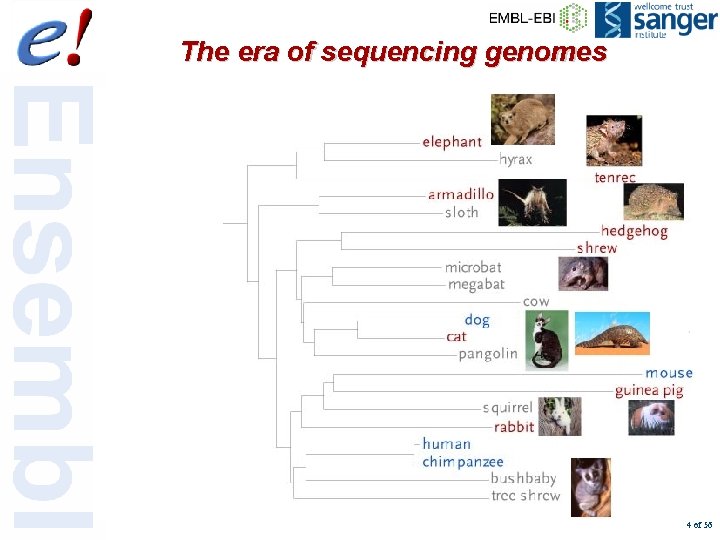
The era of sequencing genomes 23 Red : whole genome assembly available Green : whole genome assembly due within the next year in Ensembl 91 41 92 * 19 species currently in Ensembl + 10 Pre! Ensembl 105 Eutheria 310 Mammalia Amniota Tetrapoda 360 450 550 45 83 65 170 20 Metatheria Aves Vertebrata 74 Amphibia 197 140 Teleostei 70 Chordata 990 ? 25 ? Urochordata 1500? 70? 250 340 Arthropoda H. sapiens (human) * + P. troglodytes (chimpanzee) * M. mulatta (rhesus macaque) * M. musculus (house mouse) * R. norvegicus (Norway rat) * C. familiaris (dog) * F. catus (cat) E. caballus (horse) S. scrofa (pig) B. taurus (cow) * O. aries (sheep) L. africana (elephant) + M. domestica (opossum) * G. gallus (chicken) * X. tropicalis (western clawed frog) * X. laevis (African clawed frog) D. rerio (zebrafish) * O. latipes (Japanese medaka) G. aculeatus (Stickleback) + T. nigroviridis (spotted green pufferfish) * T. rubripes (torafugu) * C. savignyi (sea squirt) + C. intestinalis (transparent sea squirt) * A. aegypti (yellow fever mosquito) + A. gambiae (African malaria mosquito) * D. melanogaster (fruitfly) * A. mellifera (honey bee) * Fungi Nematoda C. elegans (nematode) * S. cerevisiae (baker’s yeast) * Million years 1000 500 400 300 200 100 4 of 56

Comparing different species • From the Ensembl perspective joins species through – orthologous/paralogous genes links – chromosome synteny links – protein family links • From a broader perspective – – – Where are syntenic regions located? How many genes are conserved? Where are orthologous/paralogous genes? Is gene order conserved? Where are potential regulatory regions? What is missing in one species, present only in another? 5 of 56

Orthologue and Paralogue Prediction • Evolutionary studies • Identify potential species-specific proteins/genes • Identify orthologues of (human) genes in model organisms 6 of 56

Gene Evolution Orthologues and Paralogues Reconstruct the Molecular Evolutionary history from the evidence visible within the known extant genes • Divergence • Speciation / Duplication • Change within allelic population • Point Mutations / Selection / Drift • Exon/domain shuffling • Transposition / Translocation • Retroposition (reverse transcription) • Horizontal gene transfer? 7 of 56

Homologue Relationships • Orthologues : any gene pairwise relation where the ancestor node is a speciation event • Paralogues : any gene pairwise relation where the ancestor node is a duplication event 8 of 56
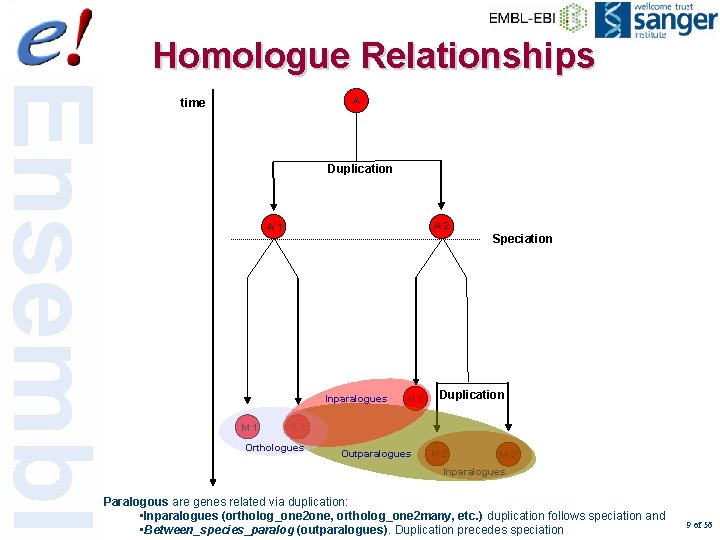
Homologue Relationships A time Duplication Inparalogues A 1 Speciation Inparalogues M 1 A 2 H 2 Duplication H 1 Orthologues Outparalogues M 2’ Inparalogues Orthologous genes have originated from a single ancestor (often have equivalent functions). Paralogous are genes related via duplication: • Inparalogues (ortholog_one 2 one, ortholog_one 2 many, etc. ) duplication follows speciation and • Between_species_paralog (outparalogues). Duplication precedes speciation 9 of 56
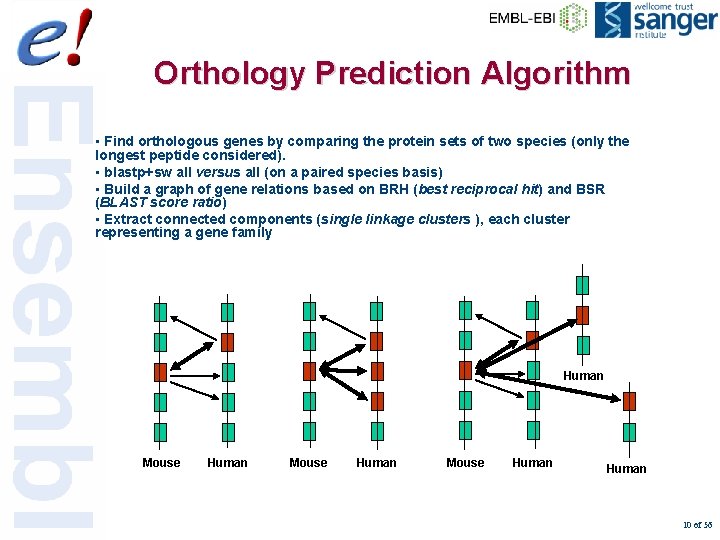
Orthology Prediction Algorithm • Find orthologous genes by comparing the protein sets of two species (only the longest peptide considered). • blastp+sw all versus all (on a paired species basis) • Build a graph of gene relations based on BRH (best reciprocal hit) and BSR (BLAST score ratio) • Extract connected components (single linkage clusters ), each cluster representing a gene family Human Mouse Human 10 of 56
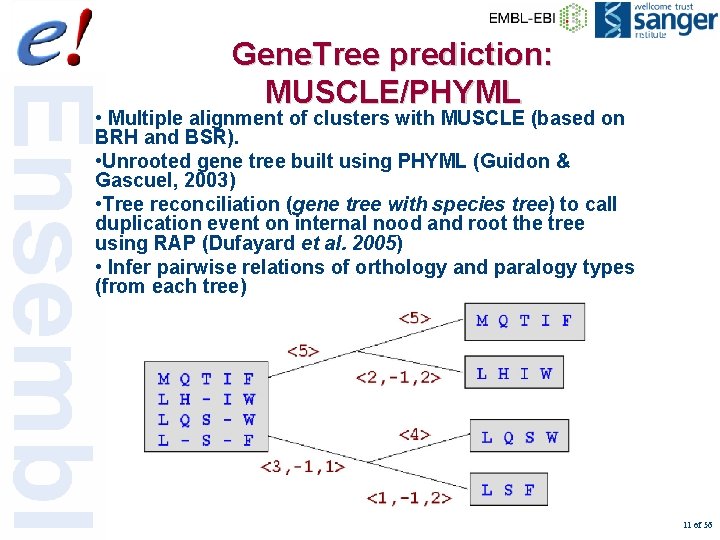
Gene. Tree prediction: MUSCLE/PHYML • Multiple alignment of clusters with MUSCLE (based on BRH and BSR). • Unrooted gene tree built using PHYML (Guidon & Gascuel, 2003) • Tree reconciliation (gene tree with species tree) to call duplication event on internal nood and root the tree using RAP (Dufayard et al. 2005) • Infer pairwise relations of orthology and paralogy types (from each tree) 11 of 56

Molecular Phylogenetics • Protein sequences in different species, both: • Provide information about the history of • evolution Reconstruct evolution • We are after an alignment that equally reflects all species: • Modeling the branching processes by comparing gene and species trees (tree reconciliation) 12 of 56
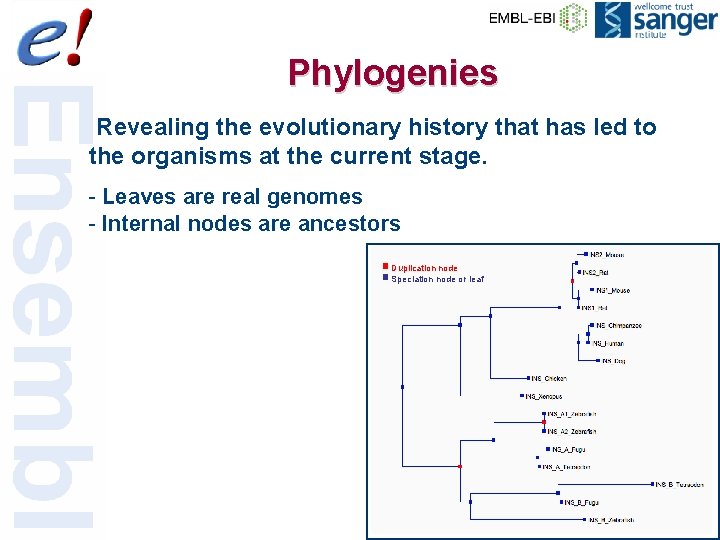
Phylogenies Revealing the evolutionary history that has led to the organisms at the current stage. - Leaves are real genomes - Internal nodes are ancestors Duplication node Speciation node or leaf 13 of 56

Orthologue and Paralogue types • • ortholog_one 2 one ortholog_one 2 many ortholog_many 2 many apparent_ortholog_one 2 one • within_species_paralog • between_species_paralog 14 of 56
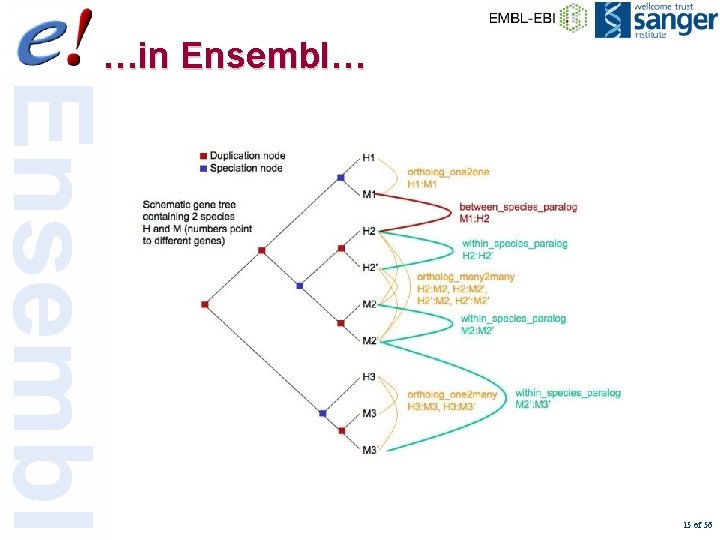
…in Ensembl… 15 of 56
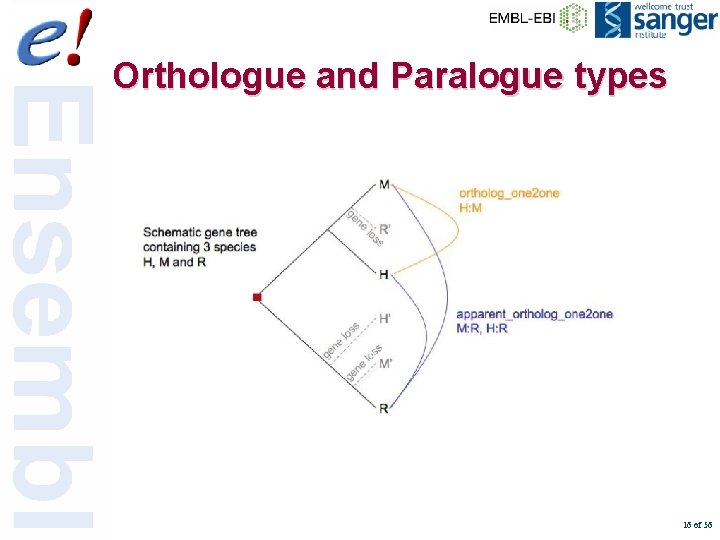
Orthologue and Paralogue types 16 of 56
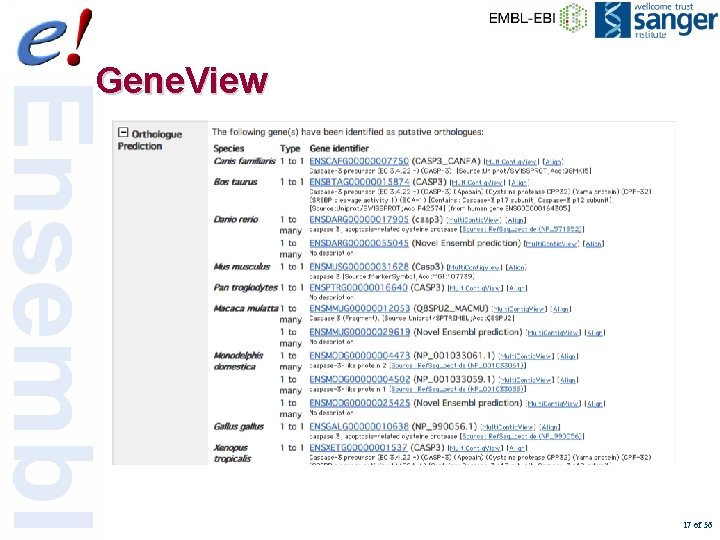
Gene. View 17 of 56
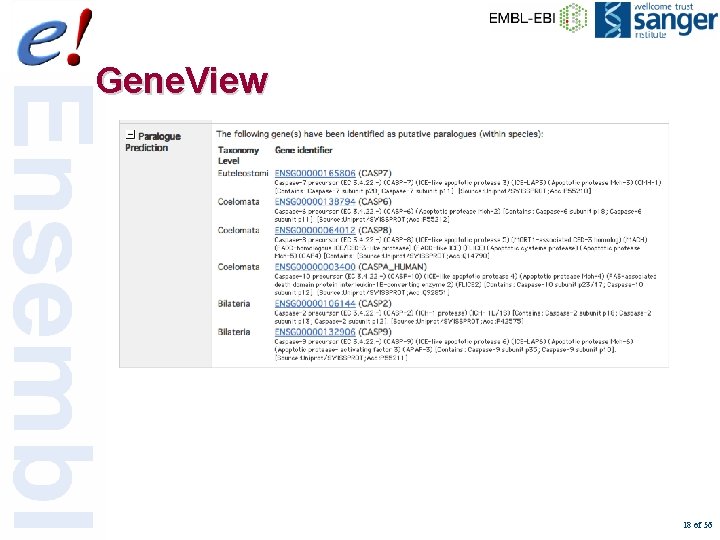
Gene. View 18 of 56
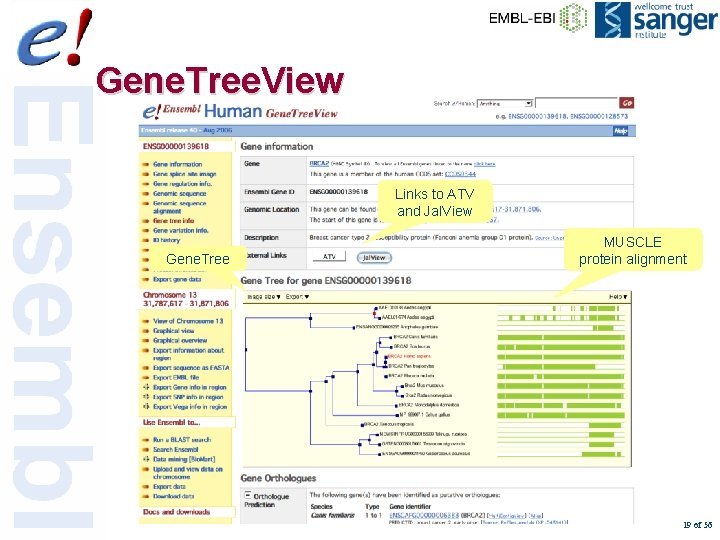
Gene. Tree. View Links to ATV and Jal. View Gene. Tree MUSCLE protein alignment 19 of 56
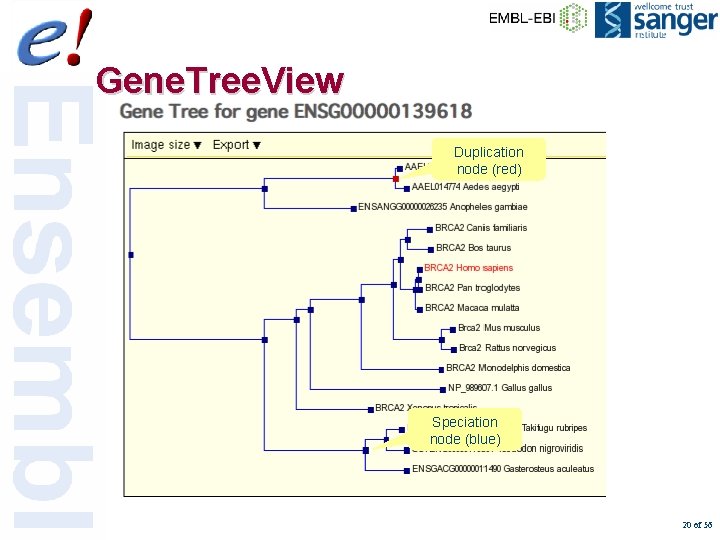
Gene. Tree. View Duplication node (red) Speciation node (blue) 20 of 56
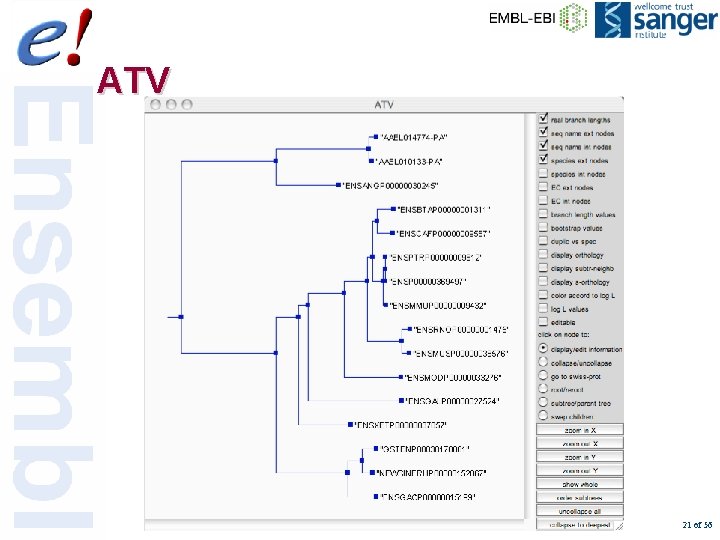
ATV 21 of 56

Protein clustering into families • Cluster proteins from different organisms that may share the same function • Obtain some kind of description for ‘novel’ genes/proteins • Locate family members over the whole genome • Identify possible orthologues and paralogues in other species 22 of 56

Protein Dataset • Nearly a million proteins clustered: – All Ensembl proteins from all species in Ensembl • 513, 256 predicted proteins – All metazoan (animal) proteins in Uni. Prot • 55, 892 Uni. Prot/Swiss-Prot • 469, 725 Uni. Prot/Tr. EMBL • Blastp all versus all, then clustering with MCL 23 of 56

Clustering Strategy • BLASTP all-versus-all comparison • Markov clustering • For each cluster: – Calculation of multiple sequence alignments with Clustal. W – Assignment of a consensus description 24 of 56
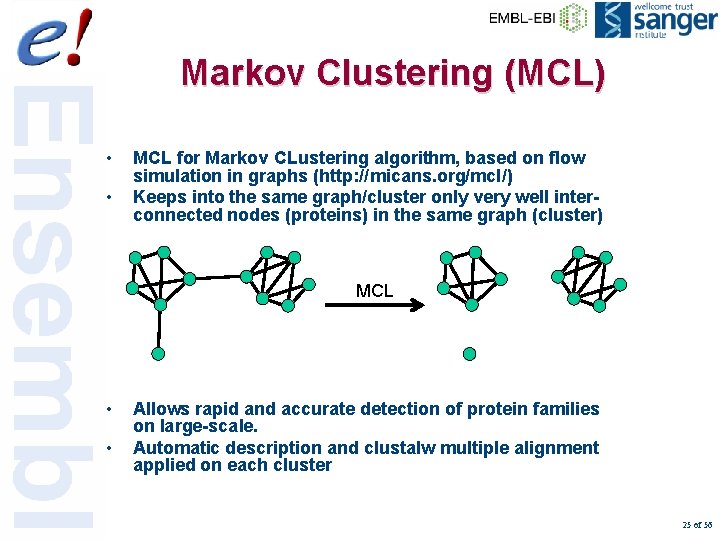
Markov Clustering (MCL) • • MCL for Markov CLustering algorithm, based on flow simulation in graphs (http: //micans. org/mcl/) Keeps into the same graph/cluster only very well interconnected nodes (proteins) in the same graph (cluster) MCL • • Allows rapid and accurate detection of protein families on large-scale. Automatic description and clustalw multiple alignment applied on each cluster 25 of 56
Prot. View Link to Family. View 26 of 56
Family. View Jal. View multiple alignments Ensembl family members within human Ensembl family members in other species 27 of 56

For each cluster • We store – Description and score – Multiple alignment • Future extensions – Improving descriptions – Multiple alignment assessment – Build phylogeny on each cluster • Using the multiple alignment • Using d. S values (mainly inside mammals) • Extend paralogous prediction 28 of 56

Aligning complete genomes 29 of 56

Whole Genome Alignments • Understand what evolution has done on the species compared, after speciation – What is missing in one species, present only in another? – Differences between closely related species may help understanding speciation • Define syntenic regions, those long regions of DNA sequences were order and orientation is highly conserved • Conserved non-coding regions – Guides to putative regulatory regions 30 of 56
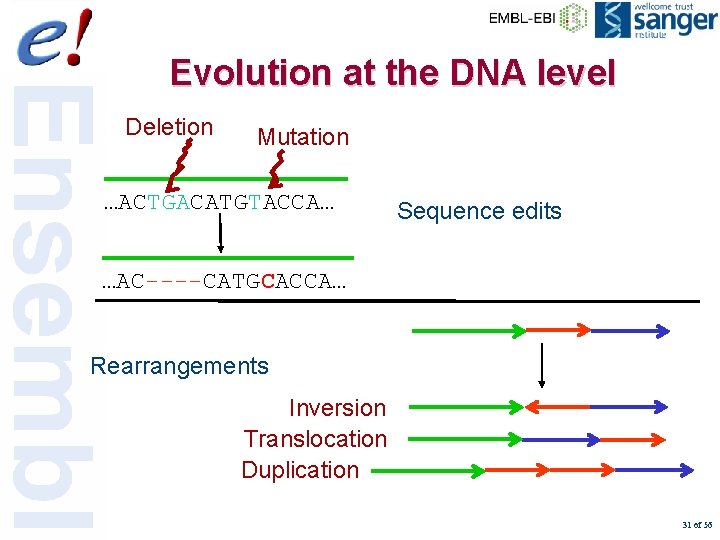
Evolution at the DNA level Deletion Mutation …ACTGACATGTACCA… Sequence edits …AC----CATGCACCA… Rearrangements Inversion Translocation Duplication 31 of 56

Basic Idea • Functional sequences evolve more slowly than non-functional sequences • Comparing genomic sequences from species at different evolutionary distances allows us to identify: – Coding genes – Non-coding regulatory sequences 32 of 56

Aligning large genomic sequences • Independent from protein/gene predictions • Should find all highly similar regions between two sequences • Should allow for segments without similarity, rearrangements etc. – Computes run only by few dedicated groups • Issues – – – Heavy process Scalability, as more and more genomes are sequenced Time constraint Computes run only by few dedicated groups As the «true» alignment is not known, then difficult to measure the alignment accuracy and apply the right method 33 of 56

Using a local aligner • Local alignment – Find all highly similar regions over 2 sequences • Find the orthologous as well as all the paralogous sequences – Separated by segments without alignment – Can handle rearranged sequences – Need post- filtering to limit too much overlapping alignments 34 of 56
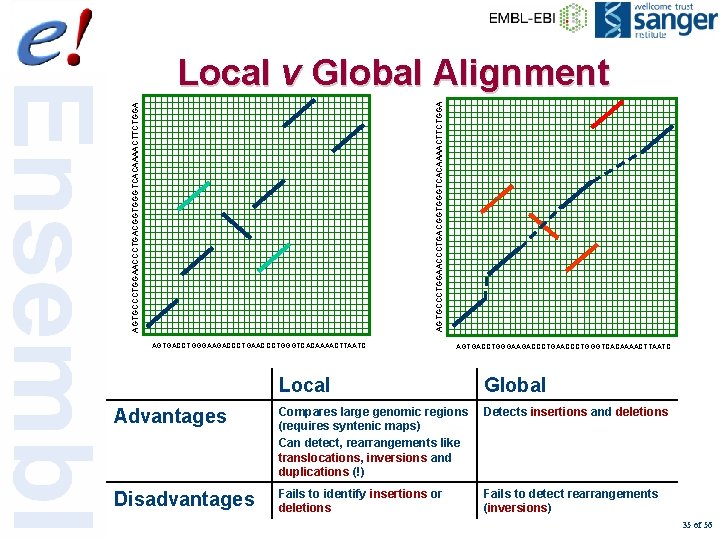
AGTGCCCTGGAACCCTGACGGTGGGTCACAAAACTTCTGGA Local v Global Alignment AGTGACCTGGGAAGACCCTGAACCCTGGGTCACAAAACTTAATC Local Global Advantages Compares large genomic regions (requires syntenic maps) Can detect, rearrangements like translocations, inversions and duplications (!) Detects insertions and deletions Disadvantages Fails to identify insertions or deletions Fails to detect rearrangements (inversions) 35 of 56

Glocal Alignment Problem GTGCCCTGGAACCCTGACGGTGGGTCACAAAACTTCTGGAG Find least cost transformation of one sequence into another using new operations • Sequence edits (indels, mutations) • Inversions • Translocations • Duplications • A combination of these AGTGACCTGGGAAGACCCTGAACCCTGGGTCACAAAACT 36 of 56 Glocal aligner (Brudno et al. , 2003)

BLASTZ-net, t. BLAT and MLAGAN • BLASTZ-net (comparison on nucleotide level) is used for species that are evolutionary close, e. g. human - mouse • Translated BLAT (comparison on amino acid level) is used for evolutionary more distant species, e. g. human - zebrafish • MLAGAN global alignment used for multispecies alignments 37 of 56

all versus all approach using BLASTZ (collaboration with UCSC) • Can handle large sequences • Used 2 -weighted spaced seeding strategy • Dynamic masking • Makes distinction between repeat and non-repeat sequences (soft masking) • Try aligning inside repeats • One iterative step with lower threshold to expand alignments 38 of 56

Blastz strategy • 10 Mb Human fragments (3000) • 30 Mb Mouse fragments (100) • Lineage-specific repeats removed • 48 hours on 1024 CPUs • Generates 9 Gb of output • When filtered for Best hit on Human, reduced to 2. 5 Gb • 10 Mb Human fragments (3000) • 30 Mb Mouse fragments (100) 39 of 56

Blastz human genome coverage • 40% of the human genome is covered by an alignment of mouse sequences By rescoring the alignment over a “tight” matrix that is very stringent and look for high conservation (>70% identity), the coverage goes down to 6% 40 of 56

DNA/DNA matches web display Contig. View human EPO Conserved sequences 41 of 56
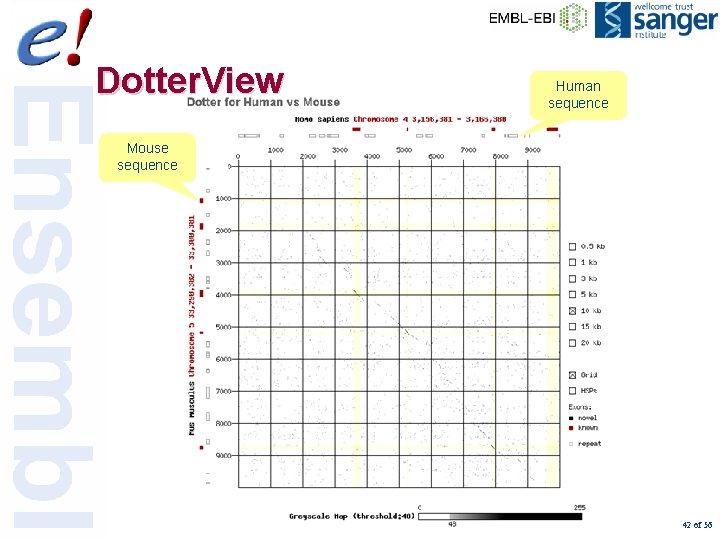
Dotter. View Human sequence Mouse sequence 42 of 56

Multiple alignments • Currently 3 sets: – MLAGAN-primates: – MLAGAN-amniote vertebrates: – MLAGAN-eutherian mammals: 43 of 56
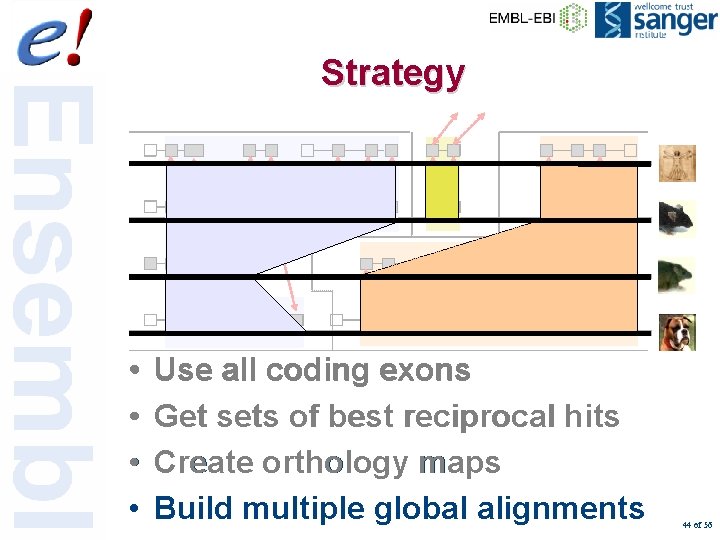
Strategy • • Use all coding exons Get sets of best reciprocal hits Create orthology maps Build multiple global alignments 44 of 56
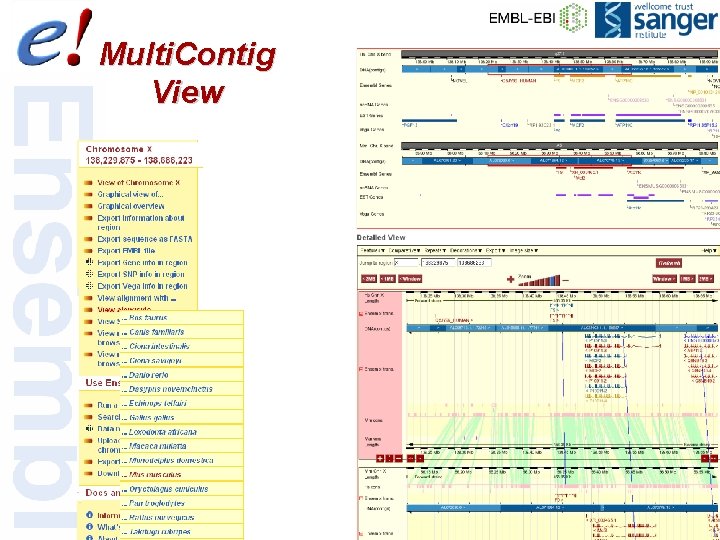
Multi. Contig View 45 of 56

Multiple alignments Contig. View human EPO 46 of 56

Align. Splice. View Export alignments Human Dog Rat Mouse Alignment on basepair level 47 of 56

Multi. Contig. View vs. Align. Slice. View 48 of 56

Align. View 49 of 56
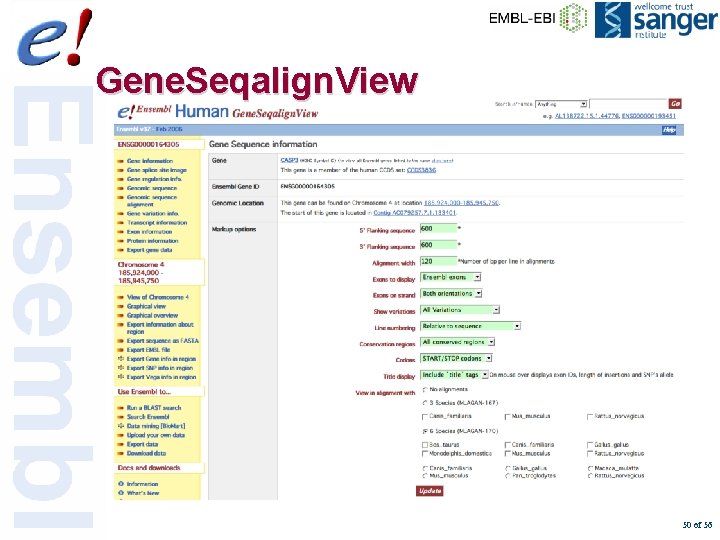
Gene. Seqalign. View 50 of 56

Gene. Seqalign. View 51 of 56

Syntenic Regions • Genome alignments are refined into larger syntenic regions • Alignments are clustered together when the relative distance between them is less than 100 kb and order and orientation are consistent • Any clusters less than 100 kb are discarded 52 of 56
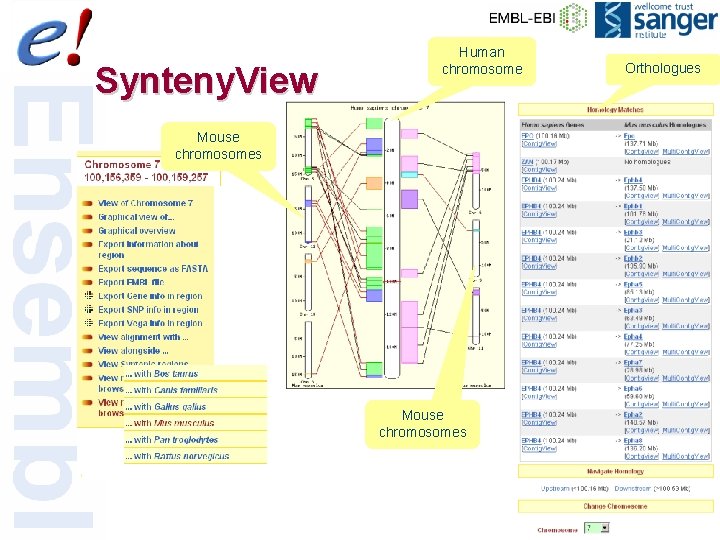
Synteny. View Human chromosome Orthologues Mouse chromosomes 53 of 56
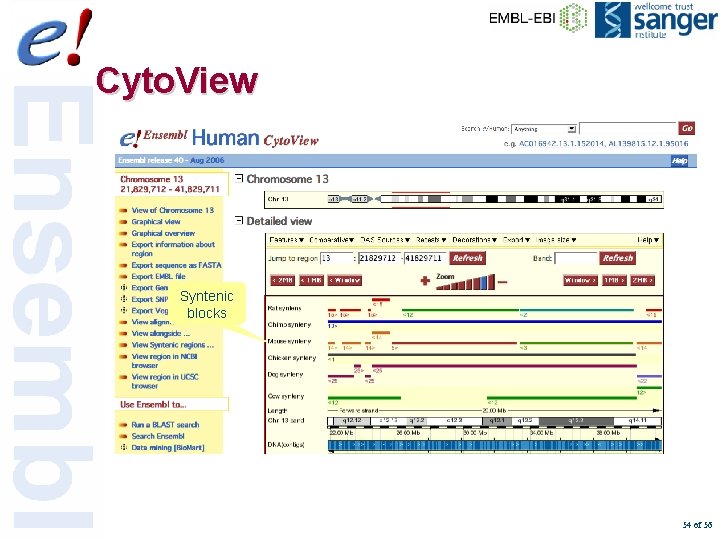
Cyto. View Syntenic blocks 54 of 56

Outlook • Ortho. View • Displaying alignments both from whole genome alignments and on orthologues • Consider all isoforms for each gene • Calculate d. N/d. S 55 of 56

Acknowledgements • • • Abel Ureta-Vidal Benoît Ballester Kathryn Beal Stephen Fitzgerald Javier Herrero Albert Vilella Ensembl team Sep 2006 56 of 56
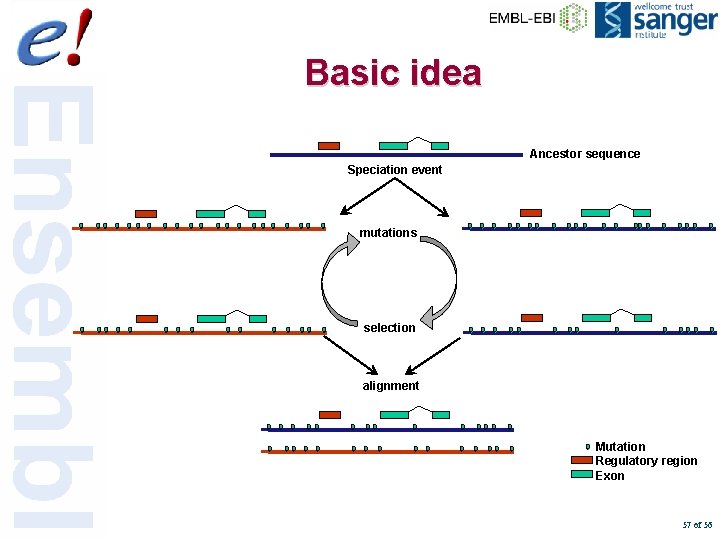
Basic idea Ancestor sequence Speciation event mutations selection alignment Mutation Regulatory region Exon 57 of 56
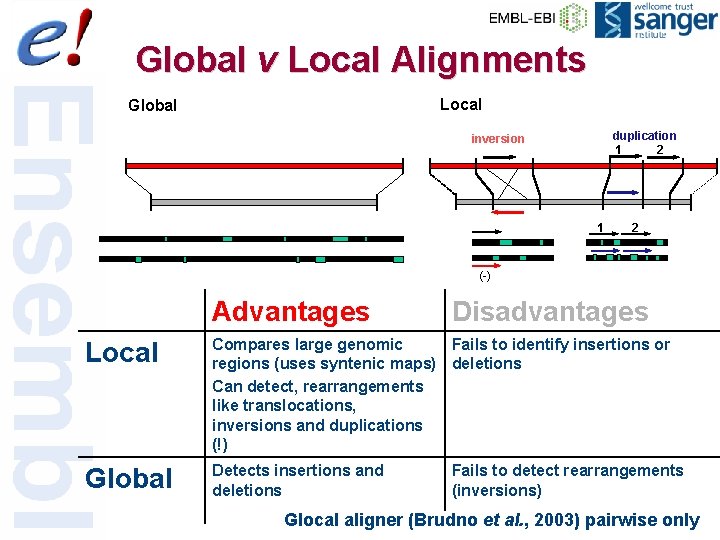
Global v Local Alignments Local Global duplication 1 2 inversion 1 2 (-) Advantages Disadvantages Local Compares large genomic Fails to identify insertions or regions (uses syntenic maps) deletions Can detect, rearrangements like translocations, inversions and duplications (!) Global Detects insertions and deletions Fails to detect rearrangements (inversions) Glocal aligner (Brudno et al. , 2003) pairwise only 58 of 56

Inparalogues vs Outparalogues 59 of 56 Adapted from Sonnhammer & Koonin (2002) TIG 18, 12: 620
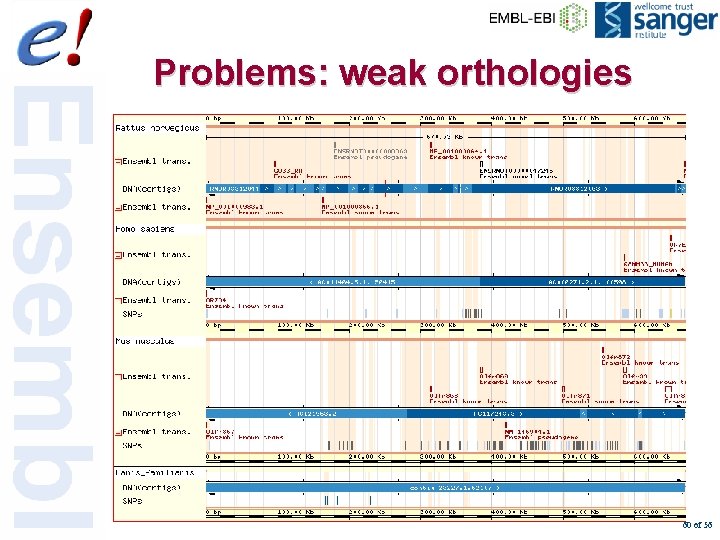
Problems: weak orthologies 60 of 56

Problems: missalignments 61 of 56

Possible solutions • Weak orthologies: • Poor alignments: – report to author – edit alignments, detect wrong edges, redefine blocks – use another aligner 62 of 56
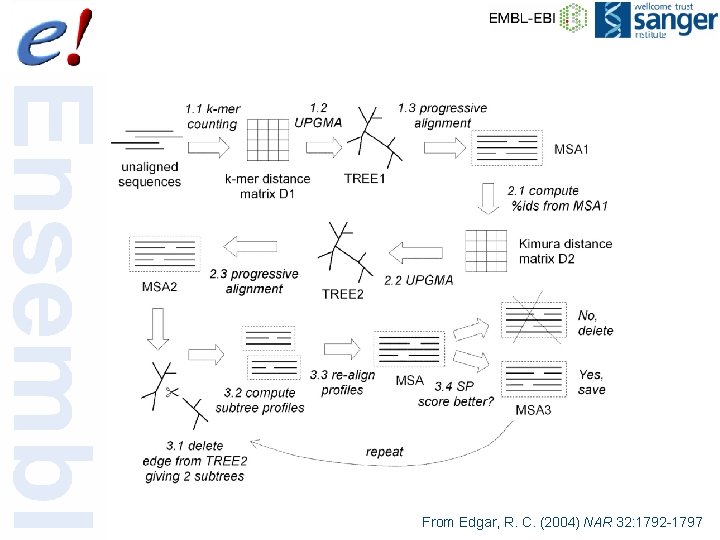
From Edgar, R. C. (2004) NAR 32: 1792 -1797 63 of 56
- Slides: 63