COMP 5331 Other Clustering Techniques Prepared by Raymond

COMP 5331 Other Clustering Techniques Prepared by Raymond Wong Presented by Raymond Wong raywong@cse

What we learnt n n K-mean Dendrogram

Other Clustering Models n Model-Based Clustering n n Density-Based Clustering n n EM Algorithm DBSCAN Scalable Clustering Method n BIRCH

EM Algorithm n Drawback of the K-means/Dendrogram n n n Each point belongs to a single cluster There is no representation that a point can belong to different clusters with different probabilities Use probability density to associate to each point

EM Algorithm n n Assume that we know there are k clusters Each cluster follows a distribution (e. g. , Gaussian Distribution) n 1 D Gaussian Distribution n n Mean Standard derivation p(x| < , >) =

EM Algorithm n n Since there are k clusters, we have k distributions. Cluster 1 n Gaussian Distribution n Mean 1 n n Cluster 2 n Gaussian Distribution n Mean 2 n n n Standard derivation 1 Standard derivation 2 … Cluster k n Gaussian Distribution n Mean k n Standard derivation k

EM Algorithm n n Expectation-Maximization Algorithm n n Step 1 (Parameter Initialization) n Initialize all i and i One possible implementation: Step 2 (Expectation) p(x|< , >) n For each point x, n n i i jp(x|< j, j>) For each cluster i, n Calculate the probability that x belongs to cluster i Step 3 (Maximization) n For each cluster i, n n p(x Ci) = Calculate the mean i according to the probabilities that all points belong to cluster i Repeat Step 2 and Step 3 until the parameters converge One possible implementation: i = x x . p(x Ci) y p(y Ci)

Other Clustering Models n Model-Based Clustering n n Density-Based Clustering n n EM Algorithm DBSCAN Scalable Clustering Method n BIRCH

DBSCAN n Traditional Clustering n n n Can only represent sphere clusters Cannot handle irregular shaped clusters DBSCAN n Density-Based Spatial Clustering of Applications with Noise

DBSCAN n a b c d e Given a point p and a non-negative real number , n the -neighborhood of point p, denoted by N(p), is the set of points q (including point p itself) such that the distance between p and q is within .
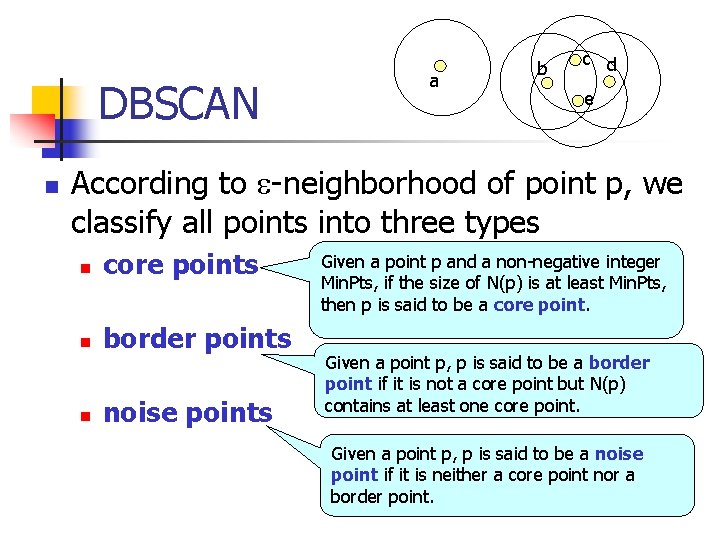
DBSCAN n a b c d e According to -neighborhood of point p, we classify all points into three types n core points n border points n noise points Given a point p and a non-negative integer Min. Pts, if the size of N(p) is at least Min. Pts, then p is said to be a core point. Given a point p, p is said to be a border point if it is not a core point but N(p) contains at least one core point. Given a point p, p is said to be a noise point if it is neither a core point nor a border point.

DBSCAN n n a b c d e Principle 1: Each cluster contains at least one core point. Principle 2: Given any two core points p and q, if N(p) contains q (or N(q) contains p), then p and q are in the same cluster. Principle 3: Consider a border point p to be assigned to one of the clusters formed by Principle 1 and Principle 2. Suppose N(p) contains multiple core points. A border point p is assigned arbitrarily to one of the clusters containing these core points (formed by Principle 1 and Principle 2). Principle 4: All noise points do not belong to any clusters.

Other Clustering Models n Model-Based Clustering n n Density-Based Clustering n n EM Algorithm DBSCAN Scalable Clustering Method n BIRCH

BIRCH n Disadvantage of Previous Algorithms n n n Most previous algorithms cannot handle update Most previous algorithms are not scalable BIRCH n Balanced Iterative Reducing and Clustering Using Hierarchies

BIRCH n Advantages n n Incremental Scalable

BIRCH n Each cluster has the following three terms. n Mean n Radius n Diameter Average distance from member objects to the mean Average pairwise distance within a cluster

BIRCH n n L is to store a list of clusters Idea of the Algorithm n n L {} When there is a new data point x n If L = {} n n n Else n n n Create cluster C containing x only Insert C into L Find the closest cluster C from x Insert x into C If C has diameter D greater than a given threshold, n Split cluster C into two sub-clusters C 1 and C 2 n Remove C from L n Insert C 1 and C 2 into L

BIRCH n If there is no efficient data structure, n n the computation of D is very slow Running time = O(? )
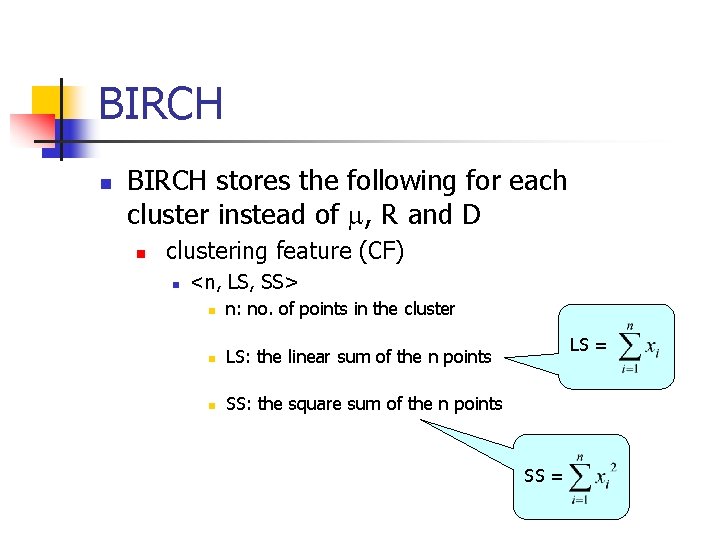
BIRCH n BIRCH stores the following for each cluster instead of , R and D n clustering feature (CF) n <n, LS, SS> n n: no. of points in the cluster n LS: the linear sum of the n points n SS: the square sum of the n points LS = SS =

BIRCH n It is easy to verify that R and D can be derived from CF

BIRCH n n Why Efficient? When a new data point comes, we can update the CF structure efficiently n n Running time = O(? ) clustering feature (CF) n <n, LS, SS> n n: no. of points in the cluster n LS: the linear sum of the n points n SS: the square sum of the n points LS = SS =
- Slides: 21