Combining Different Marker Densities in Genomic Evaluation Paul

Combining Different Marker Densities in Genomic Evaluation Paul Van. Raden 1, Jeff O’Connell 2, George Wiggans 1, Kent Weigel 3 1 Animal Improvement Programs Lab, USDA, Beltsville, MD, USA 2 University of Maryland School of Medicine, Baltimore, MD, USA 3 University of Wisconsin Dept. Dairy Science, Madison, WI, USA Paul. Van. Raden@ars. usda. gov 2010 2007

Topics Ø Filling missing SNPs (imputation) • • • Find haplotypes from genotypes Use lower density to track higher Programs implemented April 2010 Ø Actual mixes of 3 K with 50 K Ø Simulated mixes of 50 K with 500 K Ø Calculating reliabilities Interbull meeting, Riga, Latvia May 2010 (2) Paul Van. Raden 2010

Mixing Different Chips Interbull meeting, Riga, Latvia May 2010 (3) Paul Van. Raden 2010

What is imputation? Ø Ø Ø Genotypes indicate how many copies of each allele were inherited Haplotypes indicate which alleles are on which chromosome Use observed genotypes to impute unknown haplotypes • • Pedigree haplotyping uses relatives Population haplotyping finds matching allele patterns Interbull meeting, Riga, Latvia May 2010 (4) Paul Van. Raden 2010

Why impute haplotypes? Ø Predict unknown SNP from known • • • Measure 3, 000, predict 50, 000 SNP Measure 50, 000, predict 500, 000 Measure each haplotype at highest density only a few times Ø Predict dam from progeny SNP Ø Increase reliabilities for less cost Interbull meeting, Riga, Latvia May 2010 (5) Paul Van. Raden 2010

Haplotyping Program findhap. f 90 Ø Begin with population haplotyping • • • Ø Divide chromosomes into segments, ~250 SNP / segment List haplotypes by genotype match Similar to Fast. Phase, IMPUTE End with pedigree haplotyping • • Detect crossover, fix noninheritance Impute nongenotyped ancestors Interbull meeting, Riga, Latvia May 2010 (6) Paul Van. Raden 2010

Recent Program Revisions Ø Ø Imputation and GEBV reliability are better than in 9 WCGALP paper Changes since January 2010 • • Use known haplotype if second is unknown Use current instead of base frequency Combine parent haplotypes if crossover is detected Begin search with parent or grandparent haplotypes Interbull meeting, Riga, Latvia May 2010 (7) Paul Van. Raden 2010

Most Frequent Haplotypes 5. 16% 4. 37% 4. 36% 3. 67% 3. 66% 3. 65% 3. 51% 3. 42% 3. 24% 3. 22% 02222020020020202000020020200002202222220 02202022000200220222000022002002000002002222000022020022202200200022020220000222200202220 02202022200202202020000202220000202002 0222202022020200220000020222202000002020220002022 022020022202200200022020220000222200202222 02200222202022022020022200000002022220002220 022002220022022002020020220200020202002020 02222020200000022020220020220200020202002020 0220022200202000222000020220000020202220 Most frequent haplotype in first segment of chromosome 15 for Holsteins had 4, 316 copies = 41, 822 * 2 *. 0516 Interbull meeting, Riga, Latvia May 2010 (8) Paul Van. Raden 2010

Example Bull: O-Style USA 137611441, Sire = O-Man Ø Read genotypes, write haplotypes Interbull meeting, Riga, Latvia May 2010 (9) Paul Van. Raden 2010

Find Haplotypes – AB coding Genotypes: Oman BB, AA, AB, AA, AB Ostyle BB, AA, AB, AA, AA, AB Haplotypes: OStyle (pat) B A OStyle (mat) B A A A Interbull meeting, Riga, Latvia May 2010 (10) _ _ A B A A A A _ _ Paul Van. Raden 2010

Find Haplotypes – 0, 1, 2 coding Genotypes: codes 0 = BB, 1 = AB or BA, 2 = AA Oman 0 2 2 1 1 2 2 1 Ostyle 0 2 2 1 1 2 2 1 Haplotypes: codes 0 = B , 1 = unknown, 2 = A OStyle (pat) OStyle (mat) 0 2 2 1 2 2 2 1 0 2 2 2 2 1 Interbull meeting, Riga, Latvia May 2010 (11) Paul Van. Raden 2010
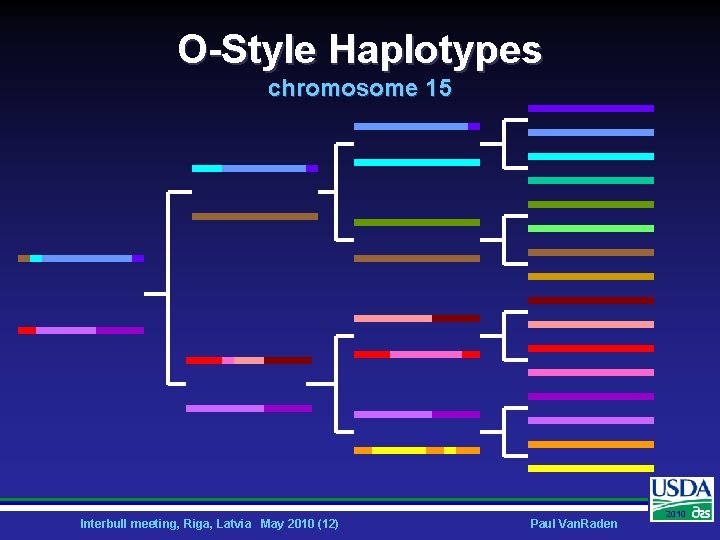
O-Style Haplotypes chromosome 15 Interbull meeting, Riga, Latvia May 2010 (12) Paul Van. Raden 2010

How does imputation work? Ø Identify haplotypes in population using many markers Ø Track haplotypes with fewer markers Ø e. g. , use 5 SNP to track 25 SNP • • 5 SNP: 22020 25 SNP: 20220200200202200 Interbull meeting, Riga, Latvia May 2010 (13) Paul Van. Raden 2010

Imputed Dams Ø If progeny and sire both genotyped • • • First progeny inherits 1 of dam’s 2 haplotypes Second progeny has 50: 50 chance to get same or other haplotype Haplotypes known with 1, 2, 3, etc. progeny are ~50%, 75%, 87%, etc. Interbull meeting, Riga, Latvia May 2010 (14) Paul Van. Raden 2010

Better Communication is Needed Ø Ø Ø “Progeny genotypes should affect dam, but programs are not yet available” Jan 2009 USDA Changes Memo “Programs are available to impute 1300 dams” Oct 2009 USDA report to Council “Encourage USDA to use genotypes, derived by imputation, in genetic evaluation” Oct 2009 Holstein USA Board of Directors (in Holstein Pulse) Interbull meeting, Riga, Latvia May 2010 (15) Paul Van. Raden 2010

Haplotyping Tests – Real Data Ø Half of young animals assigned 3 K • • Ø Proven bulls, cows all had 50 K Dams imputed using 50 K and 3 K Half of ALL animals assigned 3 K • • • Could 3 K reference animals help? 10, 000 proven bulls yet to genotype Should cows with 3 K be predictors? Interbull meeting, Riga, Latvia May 2010 (16) Paul Van. Raden 2010
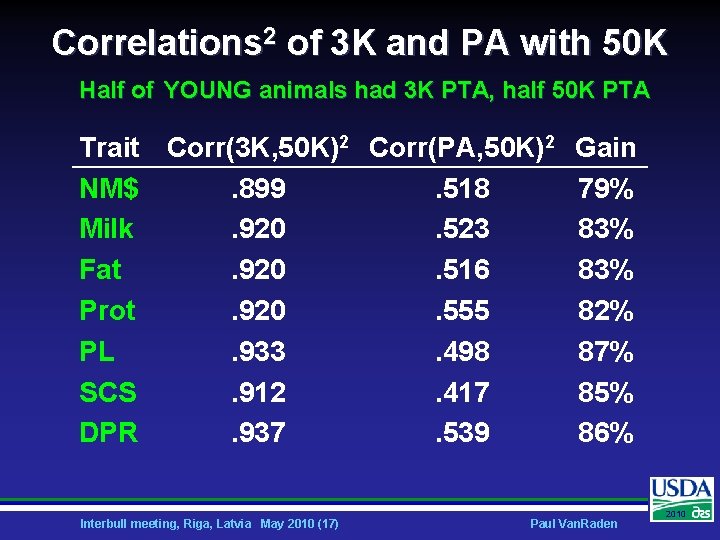
Correlations 2 of 3 K and PA with 50 K Half of YOUNG animals had 3 K PTA, half 50 K PTA Trait Corr(3 K, 50 K)2 Corr(PA, 50 K)2 NM$. 899. 518 Milk. 920. 523 Fat. 920. 516 Prot. 920. 555 PL. 933. 498 SCS. 912. 417 DPR. 937. 539 Interbull meeting, Riga, Latvia May 2010 (17) Gain 79% 83% 82% 87% 85% 86% Paul Van. Raden 2010

Using 3 K as Reference Genotypes Half of ALL animal NM$ were from 3 K, half 50 K REL Gain as compared to all 50 K Breed 50 K prog 3 K prog HO 90% 73% Imputed dams 36% JE 82% 56% 44% BS 84% 72% 55% Interbull meeting, Riga, Latvia May 2010 (18) Paul Van. Raden 2010

Simulated 500 K Genotypes Ø Linkage in base population • Similar to actual linkage reported by: – – • Ø De Roos et al, 2008 Genetics 179: 1503 Villa-Angulo et al, 2009 BMC Genetics 10: 19 Underlying linkage corresponds to D’ Three subsets of mixed 50 K and 500 K: • • • Of 33, 414, only 1, 586 (young) had 500 K Also bulls > 99% REL, total 3, 726 Also bulls > 90% REL, total 7, 398 Interbull meeting, Riga, Latvia May 2010 (19) Paul Van. Raden 2010
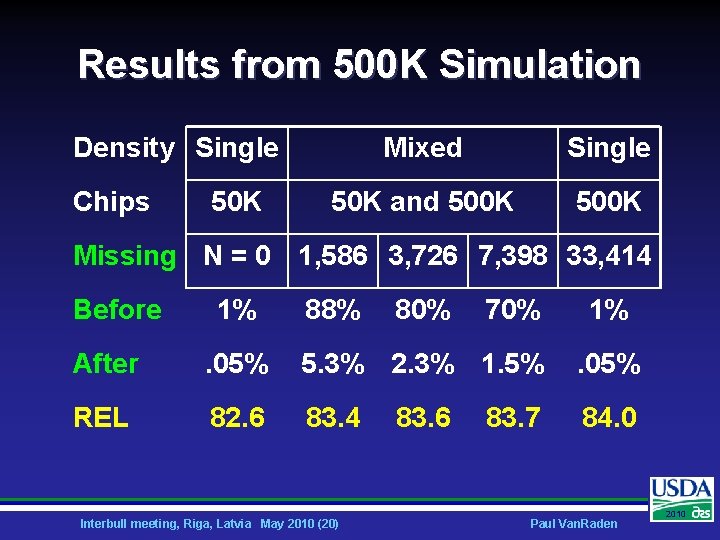
Results from 500 K Simulation Density Single Chips 50 K Mixed Single 50 K and 500 K Missing N = 0 1, 586 3, 726 7, 398 33, 414 Before 1% 88% 80% 70% 1% After . 05% 5. 3% 2. 3% 1. 5% . 05% REL 82. 6 83. 4 84. 0 Interbull meeting, Riga, Latvia May 2010 (20) 83. 6 83. 7 Paul Van. Raden 2010
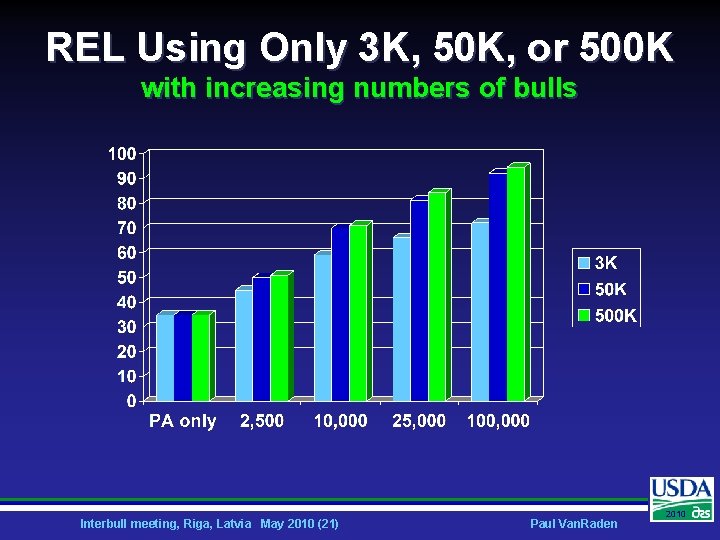
REL Using Only 3 K, 50 K, or 500 K with increasing numbers of bulls Interbull meeting, Riga, Latvia May 2010 (21) Paul Van. Raden 2010

Conclusions Ø Genomic evaluations can mix different chip densities to save $ (or € or ¥) • Ø Ø New programs implemented in April 2010 Only a few thousand of highest density genotypes needed, and other animals imputed More animals can be genotyped to increase selection differential and size of reference population Interbull meeting, Riga, Latvia May 2010 (22) Paul Van. Raden 2010

Acknowledgments Ø Ø Ø Curt Van Tassell of BFGL selected the 3, 209 low density SNP Bob Schnabel of U. Missouri fixed map locations for several SNP Mel Tooker assisted with computation Interbull meeting, Riga, Latvia May 2010 (23) Paul Van. Raden 2010
- Slides: 23