Combinatorial Optimization Problems in Computational Biology Ion Mandoiu

Combinatorial Optimization Problems in Computational Biology Ion Mandoiu CSE Department
![What Is Computational Biology? • [G. Lancia] “Study of mathematical and computational problems of What Is Computational Biology? • [G. Lancia] “Study of mathematical and computational problems of](http://slidetodoc.com/presentation_image_h2/752866a8e2a6d22878bd453473fa7906/image-2.jpg)
What Is Computational Biology? • [G. Lancia] “Study of mathematical and computational problems of modeling biological processes in the cell, removing experimental errors from genomic data, interpreting the data and providing theories about their biological relations” • Multidisciplinary field at the intersection of computer science, biology, discrete mathematics, statistics, optimization, chemistry, physics, …

5 Steps to Solving CB Problems 1. 2. Understand biological problem Represent biological data as mathematical objects (strings, sets, graphs, permutations, …), map biological relations into mathematical relations, and formulate the biological question as optimization or feasibility problem Study computational complexity: Polynomial? NP-hard? Develop efficient algorithms 3. 4. – – 5. If in P, find fast and memory efficient exact algorithms If NP-hard, find practical exact algorithms and/or algorithms with provable approximation guarantees Validate algorithms on biological data

Outline • Shortest Superstring • Sequencing by Hybridization • PCR Primer Selection
Shotgun Sequencing

Shortest Superstring • Given: set of strings s 1, s 2, …, sn • Find: shortest string s containing each si as a substring • Example: Set of strings: 000, 001, 010, 011, 100, 101, 110, 111 Superstring: 0001110100 • NP-Hard [Maier&Storer 77]

Greedy Merging Algorithm - S = {s 1, s 2, …, sn} While |S| > 1 do - Find s, t in S with longest overlap S = ( S {s, t} ) U { s overlapped with t to maximum extent} - Output final string • Approximation factor no better than 2: – s 1 = abk, s 2 =bkc, s 3 = bk+1 – Greedy output: abkcbk+1 length = 2 k+3 – Optimum: abk+1 c length = k+3 • Open problem: prove that greedy superstring is always at most twice longer than optimum

Overlap & Prefix of 2 strings • • Overlap of s and t: longest suffix of s that is a prefix of t Prefix of s and t: s after removing overlap(s, t) s = a 1 a 2 a 3 … a|s|-k+1…a|s| t = b 1 … bk … b|t| prefix(s, t) overlap(s, t)
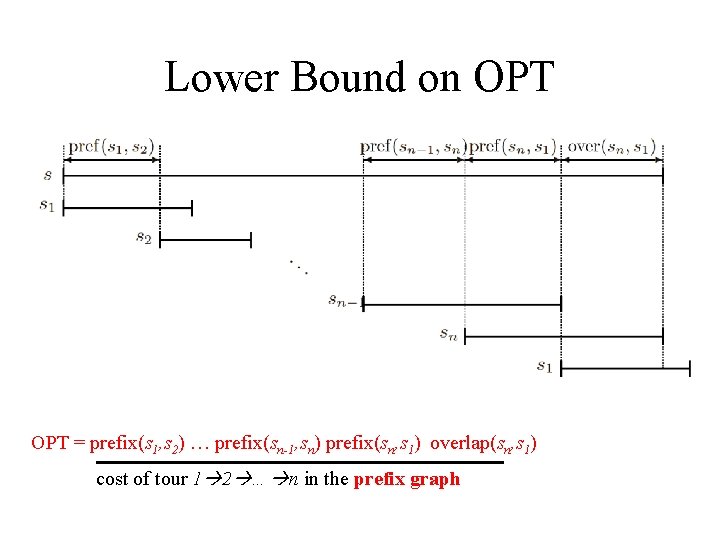
Lower Bound on OPT = prefix(s 1, s 2) … prefix(sn-1, sn) prefix(sn, s 1) overlap(sn, s 1) cost of tour 1 2 … n in the prefix graph

The Cycle Cover Algorithm • Computing TSP in prefix graph is NP-hard • Key idea: lowerbound OPT using min-weight cycle cover • For every cycle c = (i 1 i 2 … il i 1), (c) : = prefix(si 1, si 2) … prefix(sil, si 1) si 1 is a superstring of si 1, …, sil • Cycle cover algorithm:
![The Cycle Cover Algorithm Theorem [Blum, Jiang, Li, Tromp, Yannakakis 94]: Cycle cover algorithm The Cycle Cover Algorithm Theorem [Blum, Jiang, Li, Tromp, Yannakakis 94]: Cycle cover algorithm](http://slidetodoc.com/presentation_image_h2/752866a8e2a6d22878bd453473fa7906/image-11.jpg)
The Cycle Cover Algorithm Theorem [Blum, Jiang, Li, Tromp, Yannakakis 94]: Cycle cover algorithm gives factor 4 approximation. • Length of output is where ri is a “representative” string from cycle ci • wt(C) OPT - If ri no longer than wt(ci) output within factor 2 of optimum! - ri can be much longer than wt(ci) (periodic strings!) - it can be shown that | ri | OPT + 2 wt(C) factor 4
![Improved Algorithm Theorem [Blum, Jiang, Li, Tromp, Yannakakis 94]: The improved algorithm gives factor Improved Algorithm Theorem [Blum, Jiang, Li, Tromp, Yannakakis 94]: The improved algorithm gives factor](http://slidetodoc.com/presentation_image_h2/752866a8e2a6d22878bd453473fa7906/image-12.jpg)
Improved Algorithm Theorem [Blum, Jiang, Li, Tromp, Yannakakis 94]: The improved algorithm gives factor 3 approximation. Proof using that the greedy algorithm gives at least ½ of the optimum compression. Current best approximation factor is 2. 596 [Breslauer, Jiang 97]

Sequencing by Hybridization • Exploits parallel hybridization in DNA arrays • All 4 k probes of a certain length k (k=8 to 10) are synthesized on the array • Target DNA hybridizes at locations containing probes complementary to its k-substrings • Sequencing by Hybridization (SBH) Problem: Reconstruct target DNA given its k-length substrings (spectrum)

Mathematical Formulation of SBH • SBH is a special case of the shortest superstring: solution corresponds to a Hamiltonian path (NP-hard to find) in the “prefix length = 1” graph • [Pevzner 89] SBH is equivalent to finding an Eulerian path (easy to find in linear time) in the following graph: – Vertices are all (k-1)-tuples – Directed edge between two (k-1)-tuples u and v iff there is a k-length string in the spectrum whose first k symbols match u and last k symbols match v • Choose the right mathematical abstraction!
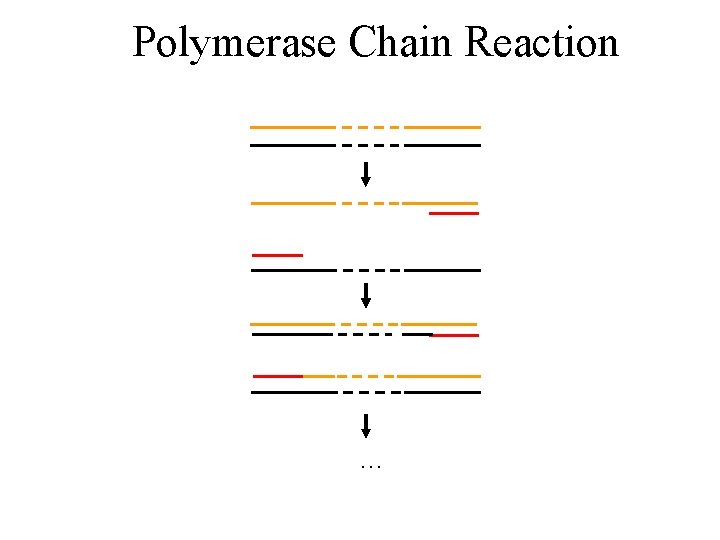
Polymerase Chain Reaction …
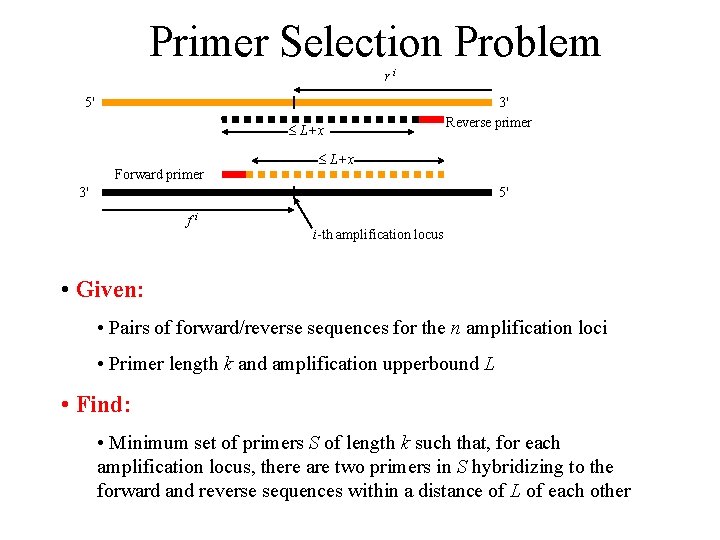
Primer Selection Problem ri 5' L+x Forward primer 3' Reverse primer L+x 3' 5' fi i-th amplification locus • Given: • Pairs of forward/reverse sequences for the n amplification loci • Primer length k and amplification upperbound L • Find: • Minimum set of primers S of length k such that, for each amplification locus, there are two primers in S hybridizing to the forward and reverse sequences within a distance of L of each other
![Previous Work • [Pearson et al. 96] Logarithmic approximation factor using greedy set cover Previous Work • [Pearson et al. 96] Logarithmic approximation factor using greedy set cover](http://slidetodoc.com/presentation_image_h2/752866a8e2a6d22878bd453473fa7906/image-17.jpg)
Previous Work • [Pearson et al. 96] Logarithmic approximation factor using greedy set cover algorithm for a formulation that does not distinguish between forward and reverse primers • Similar formulations used by [Linhart&Shamir’ 02, Souvenir et al. ’ 03] • To enforce bound of L on amplification length must truncate forward and reverse sequences to length L/2 • [Fernandes&Skiena’ 02] model primer selection as a minimum multicolored subgraph problem: • Vertices are candidate primers • Add edge colored by color i between primers u and v if they hybridize to i-th forward and reverse sequences within a distance of L • Find minimum size set of vertices inducing edges of all colors • No non-trivial approximation factor proposed
![Improved Approximations • [Konwar, M, Russell, Shvartsman 04] • Logarithmic approximation factor using “potential Improved Approximations • [Konwar, M, Russell, Shvartsman 04] • Logarithmic approximation factor using “potential](http://slidetodoc.com/presentation_image_h2/752866a8e2a6d22878bd453473fa7906/image-18.jpg)
Improved Approximations • [Konwar, M, Russell, Shvartsman 04] • Logarithmic approximation factor using “potential function” greedy for the bounded amplification length primer selection problem • O(Lln n) approximation factor based on randomized rounding for the minimum multicolored subgraph problem of [Fernandes&Skiena’ 02]
![Improved Approximations • [Konwar, M, Russell, Shvartsman 04] • Logarithmic approximation factor using “potential Improved Approximations • [Konwar, M, Russell, Shvartsman 04] • Logarithmic approximation factor using “potential](http://slidetodoc.com/presentation_image_h2/752866a8e2a6d22878bd453473fa7906/image-19.jpg)
Improved Approximations • [Konwar, M, Russell, Shvartsman 04] • Logarithmic approximation factor using “potential function” greedy for the bounded amplification length primer selection problem • O(Lln n) approximation factor based on randomized rounding for the minimum multicolored subgraph problem of [Fernandes&Skiena’ 02]
- Slides: 19