Color distribution can accelerate network alignment Md Mahmudul

Color distribution can accelerate network alignment Md Mahmudul Hasan and Tamer Kahveci University of Florida
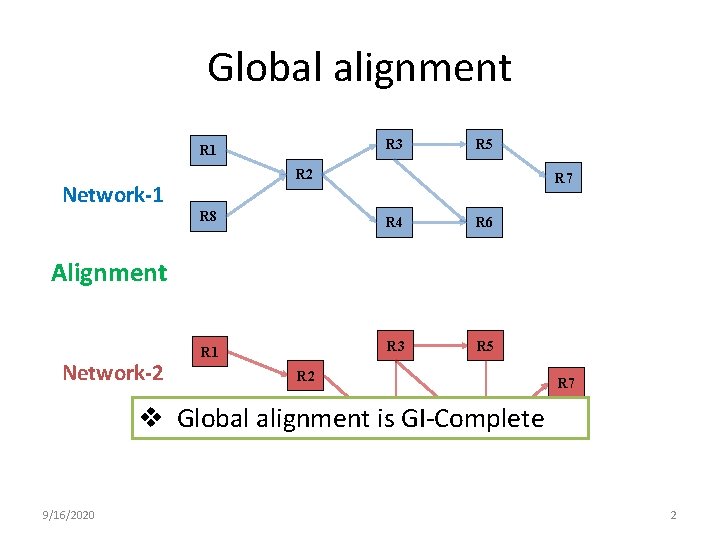
Global alignment R 3 R 1 R 5 R 2 Network-1 R 7 R 8 R 4 R 6 R 1 R 3 R 5 Alignment Network-2 R 7 v Global alignment is GI-Complete R 4 R 6 9/16/2020 2
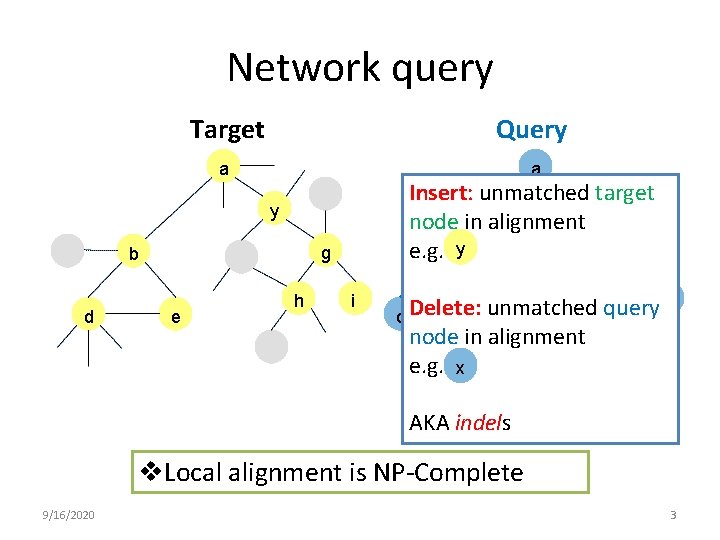
Network query Target Query a a Insert: unmatched target x node in alignment e. g. g by y g b d e h i Delete: unmatched query d e node in alignment e. g. x AKA indels v. Local alignment is NP-Complete 9/16/2020 3

Existing works in a nutshell • Heuristic alignment – Iso. RANK (Singh et al. , 2007) – Sub. MAP (Ay and Kahveci, 2010) – GRAAL (Kuchaiev et al. , 2010) –… • Approximate alignment – QPath (Shlomi et al. , 2006) – QNet (Dost et al. , 2008) – TOPAC (Gulsoy et al. 2012) –… 9/16/2020 4
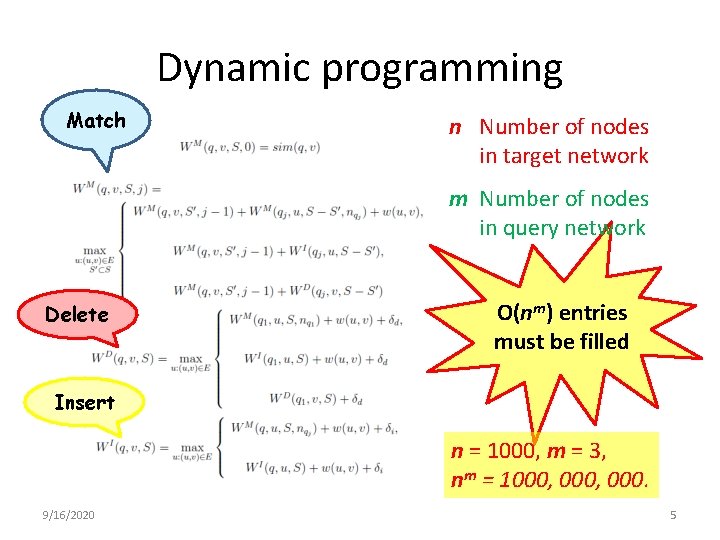
Dynamic programming Match n Number of nodes in target network m Number of nodes in query network Delete O(nm) entries must be filled Insert n = 1000, m = 3, nm = 1000, 000. 9/16/2020 5
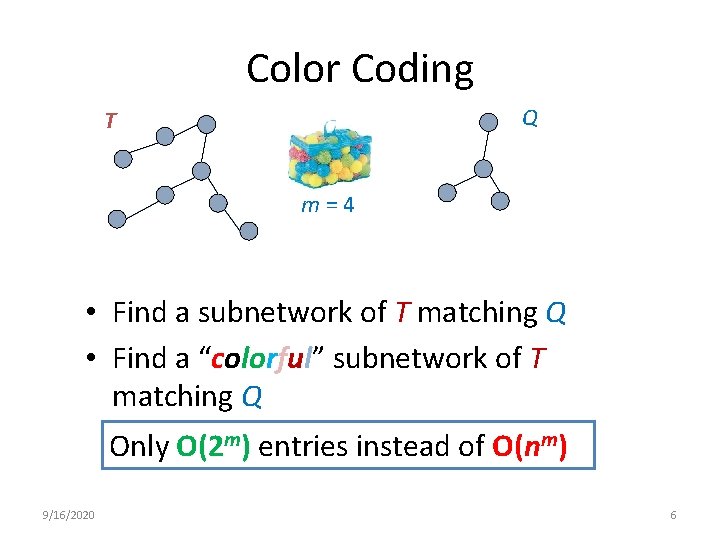
Color Coding Q T m = 4 • Find a subnetwork of T matching Q • Find a “colorful” subnetwork of T matching Q Only O(2 m) entries instead of O(nm) 9/16/2020 6

Color coding details • 9/16/2020 7
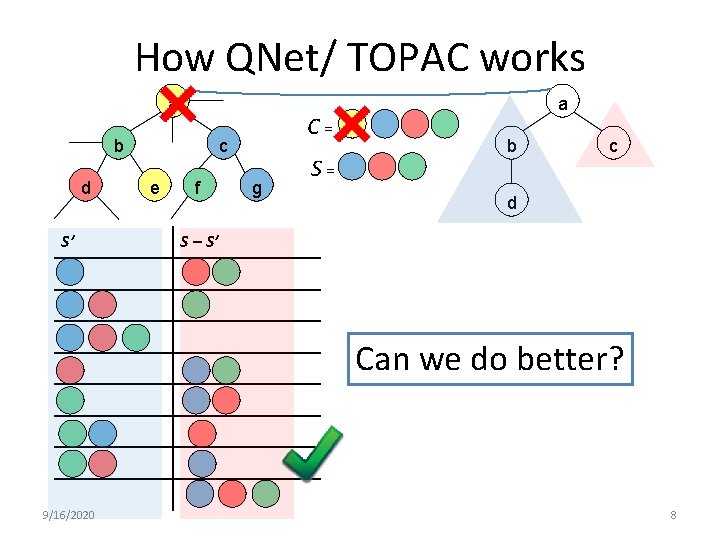
How QNet/ TOPAC works a b d S’ C = c e f g S = a b c d S – S’ Can we do better? 9/16/2020 8
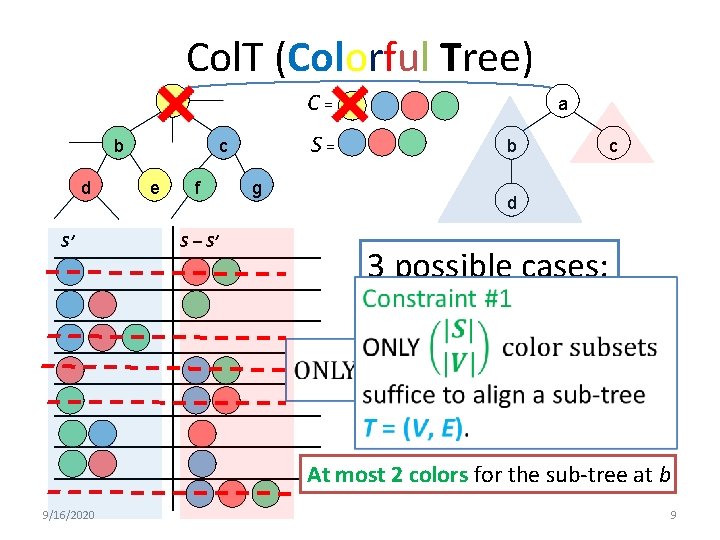
Col. T (Colorful Tree) C = a b d S’ S = c e f g a b c d S – S’ 3 possible cases: match: 2 colors insert: 2 colors delete: < 2 colors At most 2 colors for the sub-tree at b 9/16/2020 9
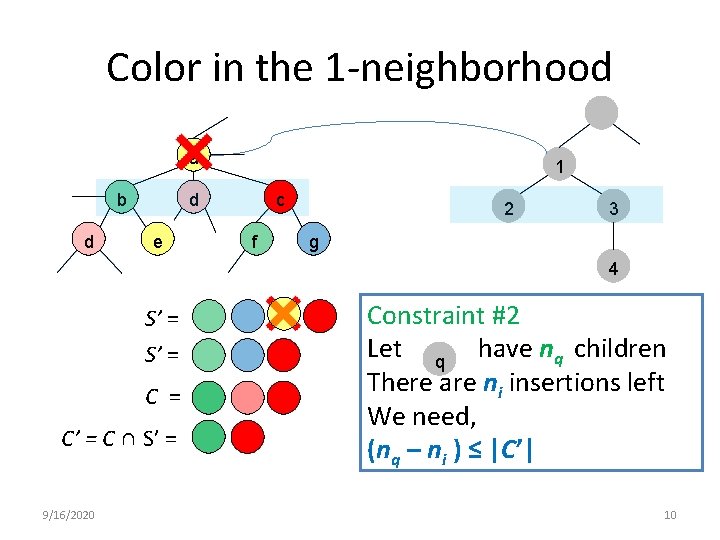
Color in the 1 -neighborhood a b d 1 d e c f 2 3 g 4 S’ = C = C’ = C ∩ S’ = 9/16/2020 Constraint #2 Insufficient color in 1 -neighborhood Let have n of for the alignment. a q q children There are ni insertions left Do not align with in this 1 a We need, coloring. (nq – ni ) ≤ |C’| 10
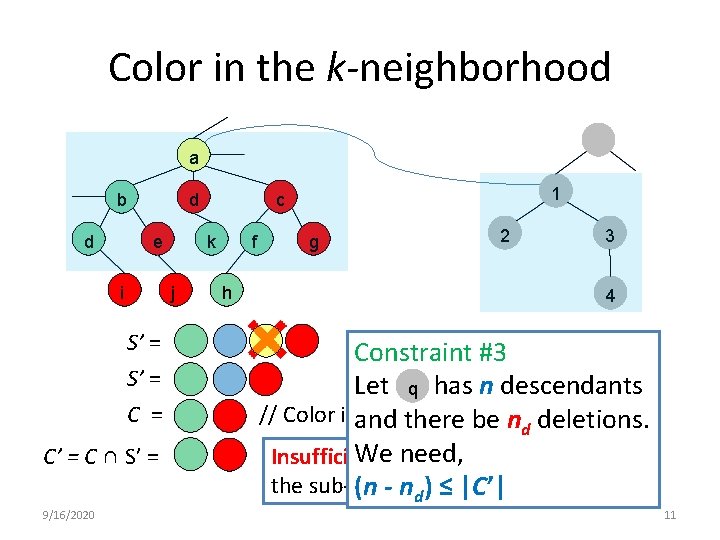
Color in the k-neighborhood a b d d e i S’ = C = C’ = C ∩ S’ = 9/16/2020 k j 1 c f h g 2 3 4 Constraint #3 Let has n descendants q // Color in 2 -neighborhood of a and there be nd deletions. We need, Insufficient color to align the sub-tree rooted at (n - nd) ≤ |C’|1 11

Experiments • Datasets – Synthetic networks • Erdős-Rényi model • Barabási-Albert model • Watts-Strogatz model (small world) – Gene regulatory networks • 297 networks, 46 organisms, 21 signaling pathways – Protein-protein interaction(PPI) network (fly) • 7, 481 proteins, 26, 201 interactions 9/16/2020 12
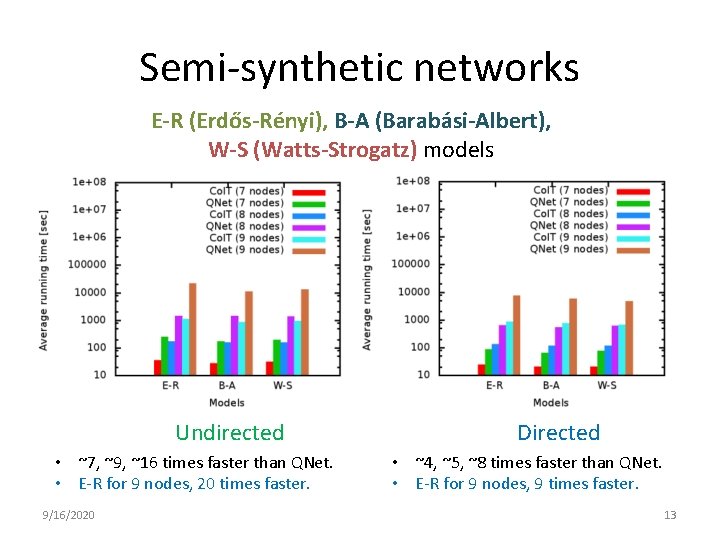
Semi-synthetic networks E-R (Erdős-Rényi), B-A (Barabási-Albert), W-S (Watts-Strogatz) models Undirected • ~7, ~9, ~16 times faster than QNet. • E-R for 9 nodes, 20 times faster. 9/16/2020 Directed • ~4, ~5, ~8 times faster than QNet. • E-R for 9 nodes, 9 times faster. 13
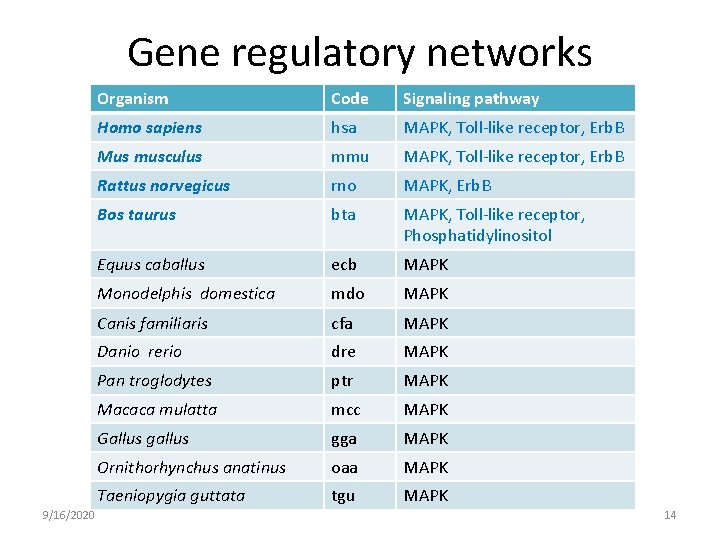
Gene regulatory networks 9/16/2020 Organism Code Signaling pathway Homo sapiens hsa MAPK, Toll-like receptor, Erb. B Mus musculus mmu MAPK, Toll-like receptor, Erb. B Rattus norvegicus rno MAPK, Erb. B Bos taurus bta MAPK, Toll-like receptor, Phosphatidylinositol Equus caballus ecb MAPK Monodelphis domestica mdo MAPK Canis familiaris cfa MAPK Danio rerio dre MAPK Pan troglodytes ptr MAPK Macaca mulatta mcc MAPK Gallus gga MAPK Ornithorhynchus anatinus oaa MAPK Taeniopygia guttata tgu MAPK 14
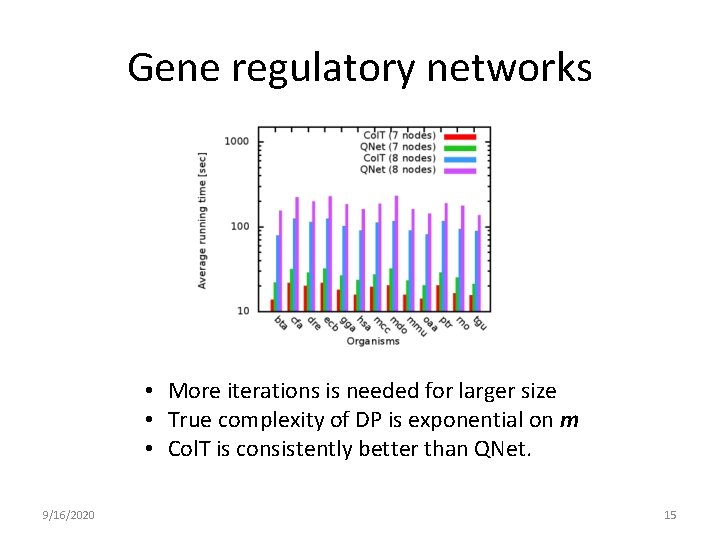
Gene regulatory networks • More iterations is needed for larger size • True complexity of DP is exponential on m • Col. T is consistently better than QNet. 9/16/2020 15
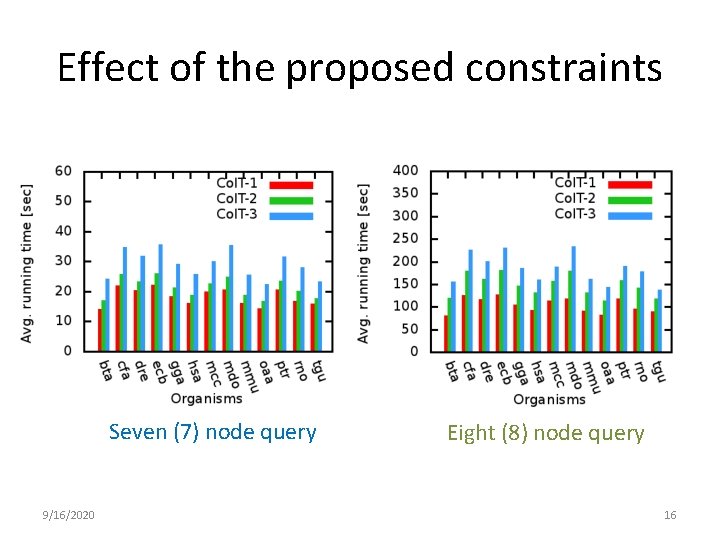
Effect of the proposed constraints Seven (7) node query 9/16/2020 Eight (8) node query 16
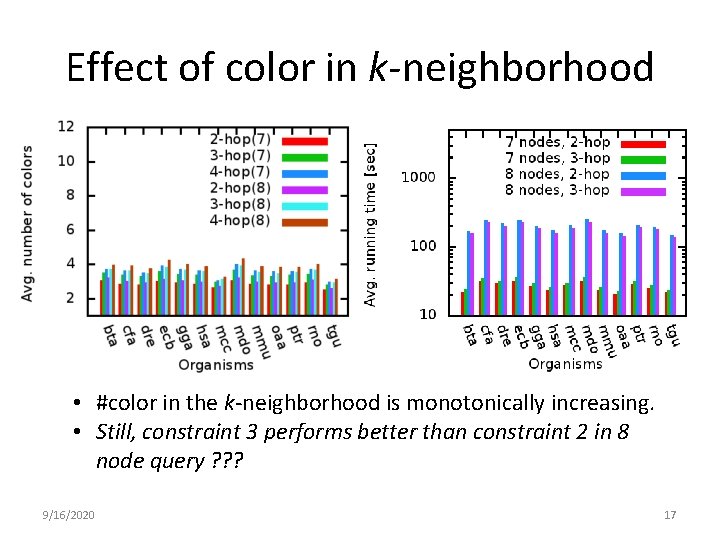
Effect of color in k-neighborhood • #color in the k-neighborhood is monotonically increasing. • Still, constraint 3 performs better than constraint 2 in 8 node query ? ? ? 9/16/2020 17
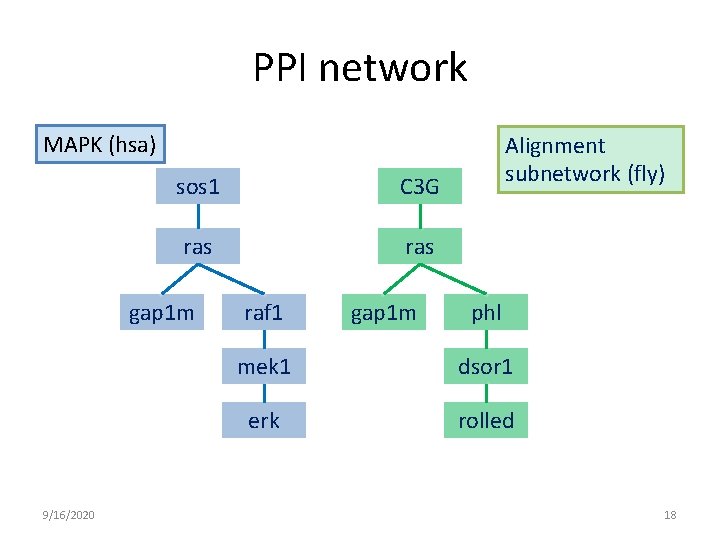
PPI network MAPK (hsa) sos 1 C 3 G ras gap 1 m 9/16/2020 raf 1 gap 1 m Alignment subnetwork (fly) phl mek 1 dsor 1 erk rolled 18

Conclusions • Col. T utilizes the color distribution in the network in color-coding. • Substantially improves over QNet (also used in methods like TOPAC). • Finds functionally similar networks in different organisms FASTER. 9/16/2020 19

Acknowledgements NSF IIS-0845439 Tamer Kahveci 9/16/2020 20

Thank You 9/16/2020 21
- Slides: 21