Clustering Gene Expression Data EMBnet DNA Microarrays Workshop

Clustering Gene Expression Data EMBnet: DNA Microarrays Workshop Mar. 4 – Mar. 8, 2002 , UNIL & EPFL, Lausanne Gaddy Getz, Weizmann Institute, Israel • Gene Expression Data • Clustering of Genes and Conditions • Methods – Agglomerative Hierarchical: Average Linkage – Centroids: K-Means – Physically motivated: Super-Paramagnetic Clustering • Coupled Two-Way Clustering Mar 2002 (GG) 1
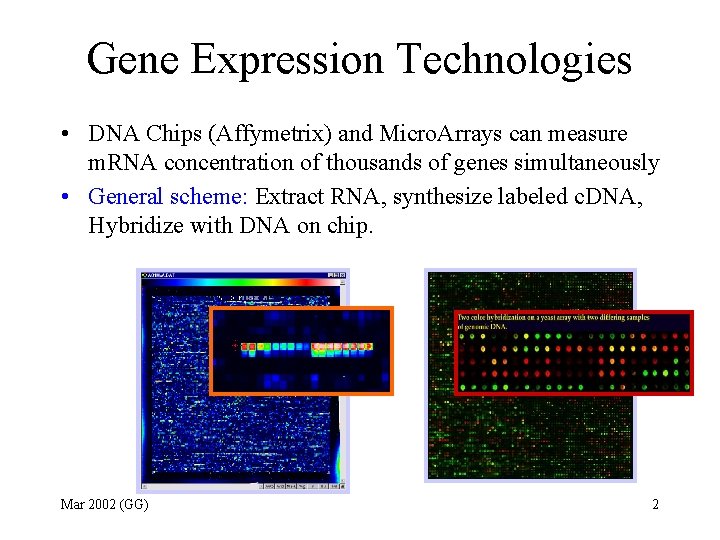
Gene Expression Technologies • DNA Chips (Affymetrix) and Micro. Arrays can measure m. RNA concentration of thousands of genes simultaneously • General scheme: Extract RNA, synthesize labeled c. DNA, Hybridize with DNA on chip. Mar 2002 (GG) 2

Single Experiment • After hybridization – Scan the Chip and obtain an image file – Image Analysis (find spots, measure signal and noise) Tools: Scan. Alyze, Affymetrix, … • Output File – Affymetrix chips: For each gene a reading proportional to the concentrations and a present/absent call. (Average Difference, Absent Call) – c. DNA Micro. Arrays: competing hybridization of target and control. For each gene the log ratio of target and control. (CH 1 I-CH 1 B, CH 2 I-CH 2 B) Mar 2002 (GG) 3

Preprocessing: From one experiment to many • Chip and Channel Normalization – Aim: bring readings of all experiments to be on the same scale – Cause: different RNA amounts, labeling efficiency and image acquisition parameters – Method: Multiply readings of each array/channel by a scaling factor such that: • The sum of the scaled readings will be the same for all arrays • Find scaling factor by a linear fit of the highly expressed genes – Note: In multi-channel experiments normalize each channel separately. Mar 2002 (GG) 4
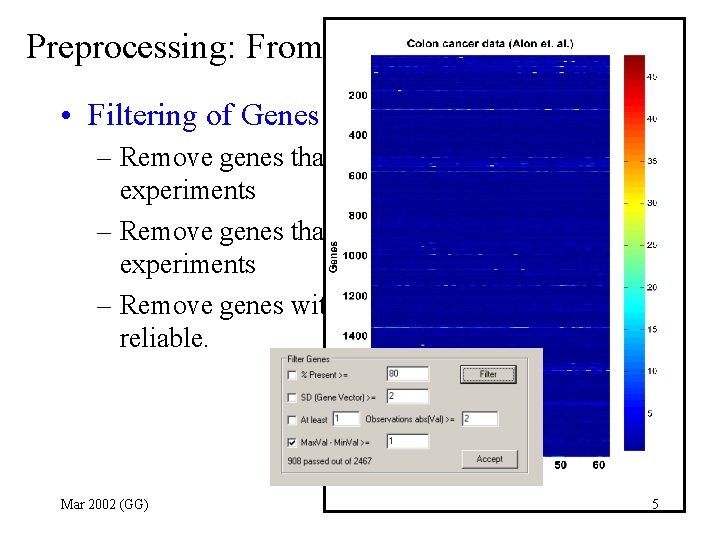
Preprocessing: From one experiment to many • Filtering of Genes – Remove genes that are absent in most experiments – Remove genes that are constant in all experiments – Remove genes with low readings which are not reliable. Mar 2002 (GG) 5
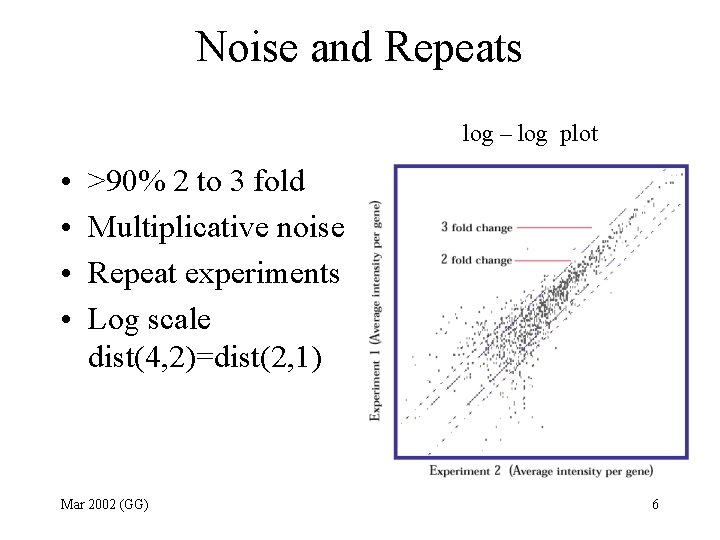
Noise and Repeats log – log plot • • >90% 2 to 3 fold Multiplicative noise Repeat experiments Log scale dist(4, 2)=dist(2, 1) Mar 2002 (GG) 6

We can. Supervised ask many Methods questions? (use predefined labels) • Which genes are expressed differently in two known types of conditions? • What is the minimal set of genes needed to distinguish one type of conditions from the others? • Which genes behave similarly in the experiments? • How many different types of conditions are there? Unsupervised Methods (use only the data) Mar 2002 (GG) 7

Unsupervised Analysis • Goal A: Find groups of genes that have correlated expression profiles. These genes are believed to belong to the same biological process and/or are co-regulated. • Goal B: Divide conditions to groups with similar gene expression profiles. Example: divide drugs according to their effect on gene expression. Clustering Methods Mar 2002 (GG) 8
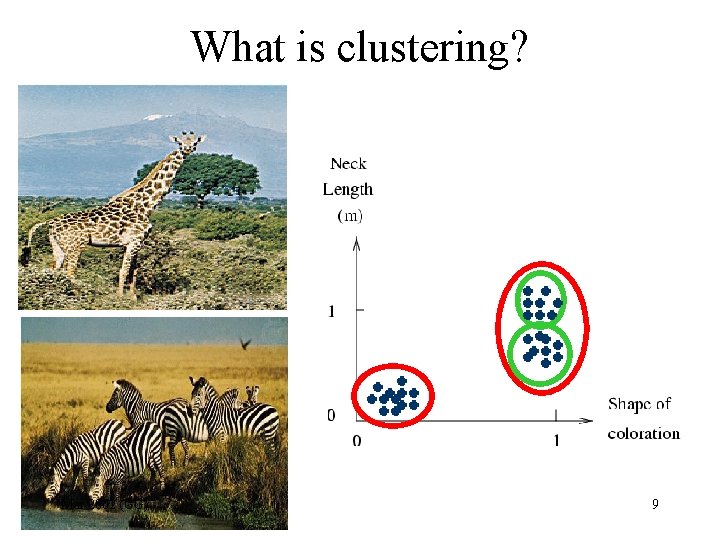
What is clustering? Mar 2002 (GG) 9
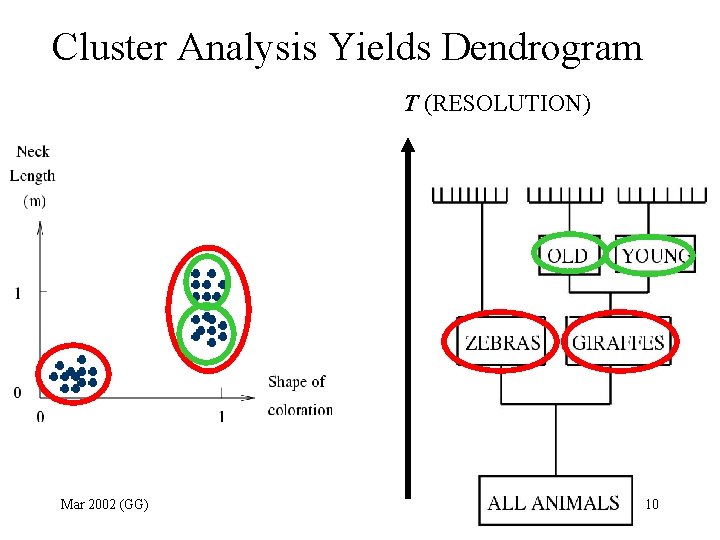
Cluster Analysis Yields Dendrogram T (RESOLUTION) Mar 2002 (GG) 10

What is clustering? More Mathematically • Input: N data points, Xi, i=1, 2, …, N in a D dimensional space. • Goal: Find “natural” groups or clusters. Data point of same cluster - “more similar” • Tasks: – Determine number of clusters – Generate a dendrogram – Identify significant “stable” clusters Mar 2002 (GG) 11

Clustering is ill-posed • Problem specific definitions • Similarity: which points should be considered close? – Correlation coefficient – Euclidean distance • Resolution: specify/hierarchical results • Shape of clusters: general, spherical. Mar 2002 (GG) 12
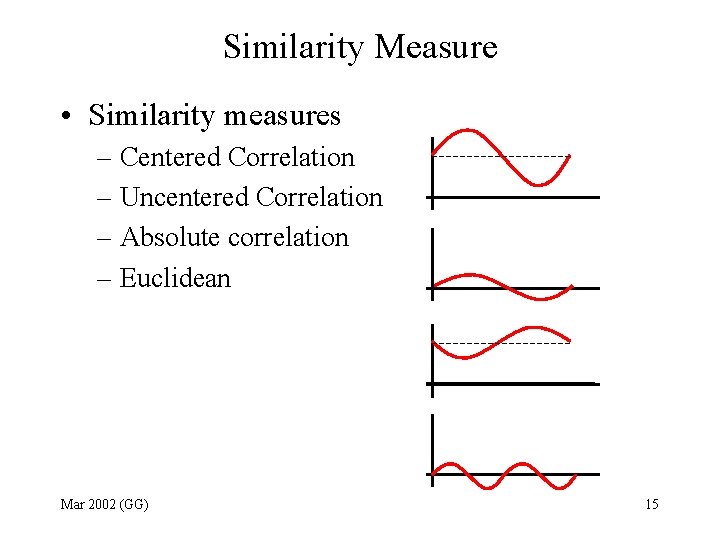
Similarity Measure • Similarity measures – Centered Correlation – Uncentered Correlation – Absolute correlation – Euclidean Mar 2002 (GG) 15
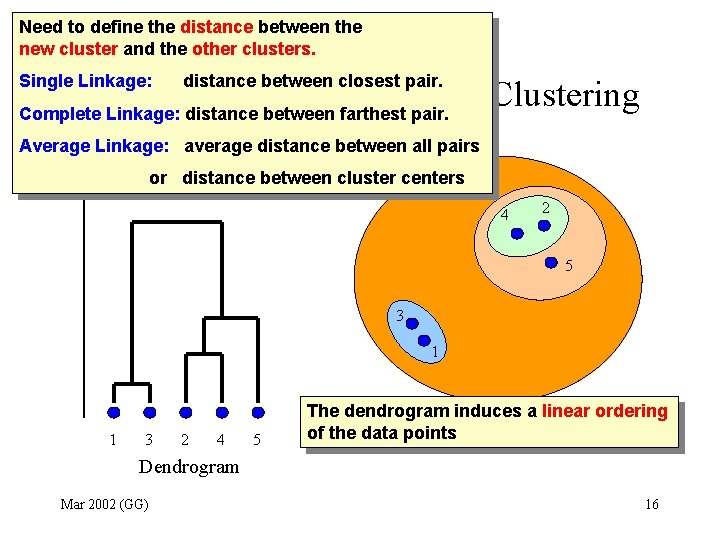
Need to define the distance between the new cluster and the other clusters. Single Linkage: distance between closest pair. Agglomerative Hierarchical Clustering Complete Linkage: distance between farthest pair. Average Linkage: average Distance between joined clustersdistance between all pairs or distance between cluster centers 4 2 5 3 1 1 3 2 4 5 The dendrogram induces a linear ordering of the data points Dendrogram Mar 2002 (GG) 16

Agglomerative Hierarchical Clustering • Results depend on distance update method – Single Linkage: elongated clusters – Complete Linkage: sphere-like clusters • Greedy iterative process • NOT robust against noise • No inherent measure to choose the clusters Mar 2002 (GG) 17
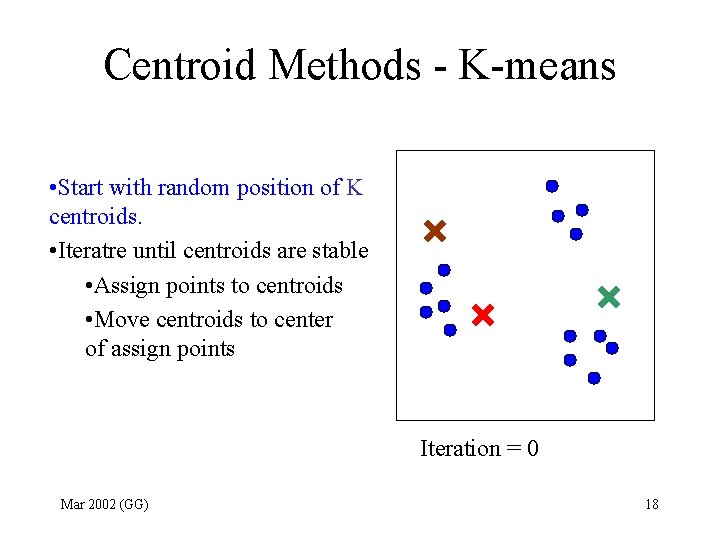
Centroid Methods - K-means • Start with random position of K centroids. • Iteratre until centroids are stable • Assign points to centroids • Move centroids to center of assign points Iteration = 0 Mar 2002 (GG) 18
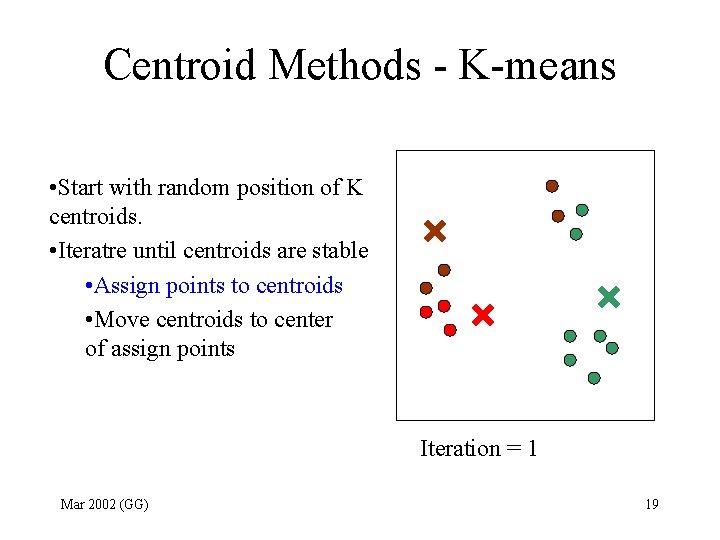
Centroid Methods - K-means • Start with random position of K centroids. • Iteratre until centroids are stable • Assign points to centroids • Move centroids to center of assign points Iteration = 1 Mar 2002 (GG) 19
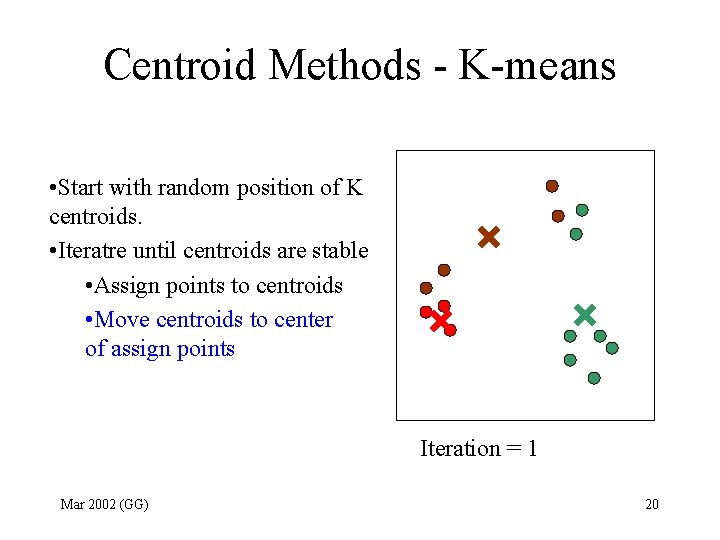
Centroid Methods - K-means • Start with random position of K centroids. • Iteratre until centroids are stable • Assign points to centroids • Move centroids to center of assign points Iteration = 1 Mar 2002 (GG) 20
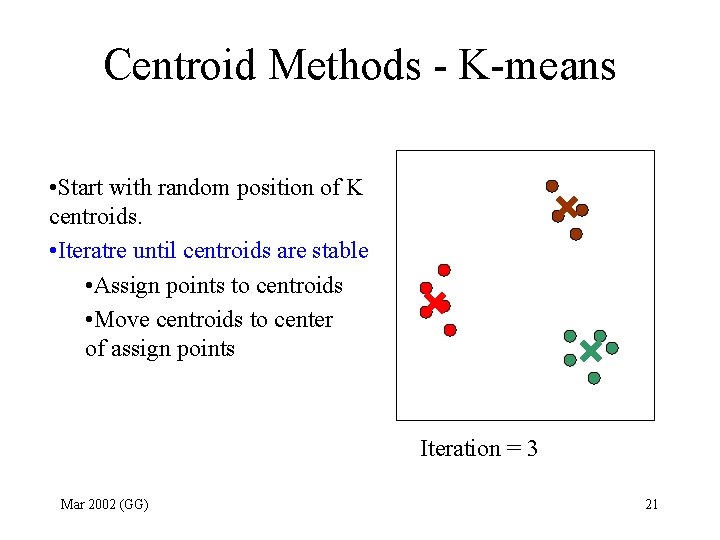
Centroid Methods - K-means • Start with random position of K centroids. • Iteratre until centroids are stable • Assign points to centroids • Move centroids to center of assign points Iteration = 3 Mar 2002 (GG) 21

Centroid Methods - K-means • Result depends on initial centroids’ position • Fast algorithm: compute distances from data points to centroids • No way to choose K. • Example: 3 clusters / K=2, 3, 4 • Breaks long clusters Mar 2002 (GG) 22

Super-Paramagnetic Clustering (SPC) M. Blatt, S. Weisman and E. Domany (1996) Neural Computation • The idea behind SPC is based on the physical properties dilute magnets. • Calculating correlation between magnet orientations at different temperatures (T). T=Low Mar 2002 (GG) 23

Super-Paramagnetic Clustering (SPC) M. Blatt, S. Weisman and E. Domany (1996) Neural Computation • The idea behind SPC is based on the physical properties dilute magnets. • Calculating correlation between magnet orientations at different temperatures (T). T=High Mar 2002 (GG) 24

Super-Paramagnetic Clustering (SPC) M. Blatt, S. Weisman and E. Domany (1996) Neural Computation • The idea behind SPC is based on the physical properties dilute magnets. • Calculating correlation between magnet orientations at different temperatures (T). T=Intermediate Mar 2002 (GG) 25
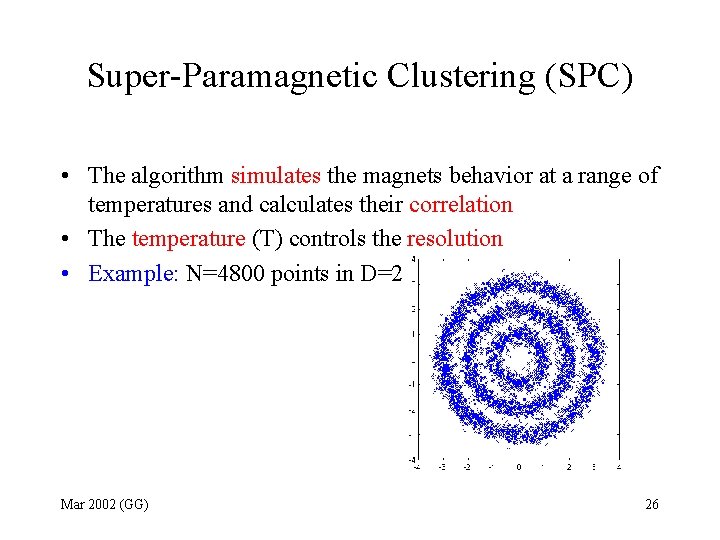
Super-Paramagnetic Clustering (SPC) • The algorithm simulates the magnets behavior at a range of temperatures and calculates their correlation • The temperature (T) controls the resolution • Example: N=4800 points in D=2 Mar 2002 (GG) 26
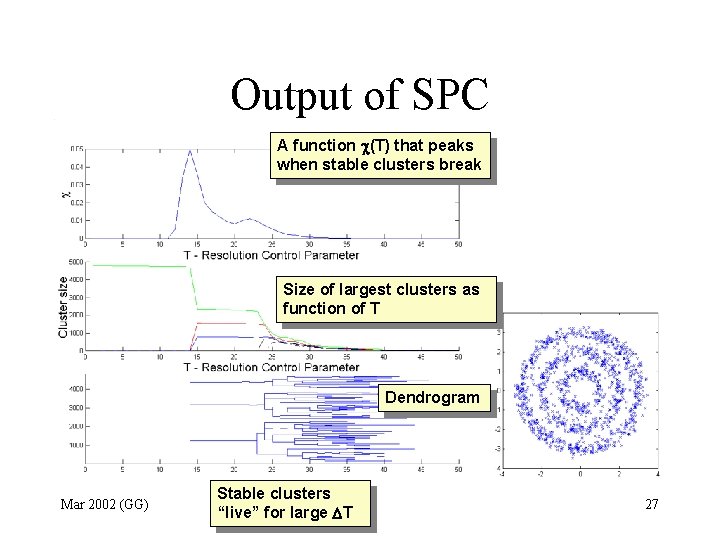
Output of SPC A function (T) that peaks when stable clusters break Size of largest clusters as function of T Dendrogram Mar 2002 (GG) Stable clusters “live” for large T 27

Choosing a value for T Mar 2002 (GG) 28

Advantages of SPC • Scans all resolutions (T) • Robust against noise and initialization calculates collective correlations. • Identifies “natural” ( ) and stable clusters ( T) • No need to pre-specify number of clusters • Clusters can be any shape Mar 2002 (GG) 29

Many clustering methods applied to expression data • Agglomerative Hierarchical – Average Linkage (Eisen et. al. , PNAS 1998) • Centroid (representative) – K-Means (Golub et. al. , Science 1999) – Self Organized Maps (Tamayo et. al. , PNAS 1999) • Physically motivated – Deterministic Annealing (Alon et. al. , PNAS 1999) – Super-Paramagnetic Clustering (Getz et. al. , Physica A 2000) Mar 2002 (GG) 30

Available Tools • Software packages: – M. Eisen’s programs for clustering and display of results (Cluster, Tree. View) • Predefined set of normalizations and filtering • Agglomerative, K-means, 1 D SOM • Web sites: – Coupled Two-Way Clustering (CTWC) website http: //ctwc. weizmann. ac. il both CTWC and SPC – http: //ep. ebi. ac. uk/EP/EPCLUST/ • General mathematical tools – MATLAB • Agglomerative, public m-files. – Statistical programs (SPSS, SAS, S-plus) Mar 2002 (GG) 31
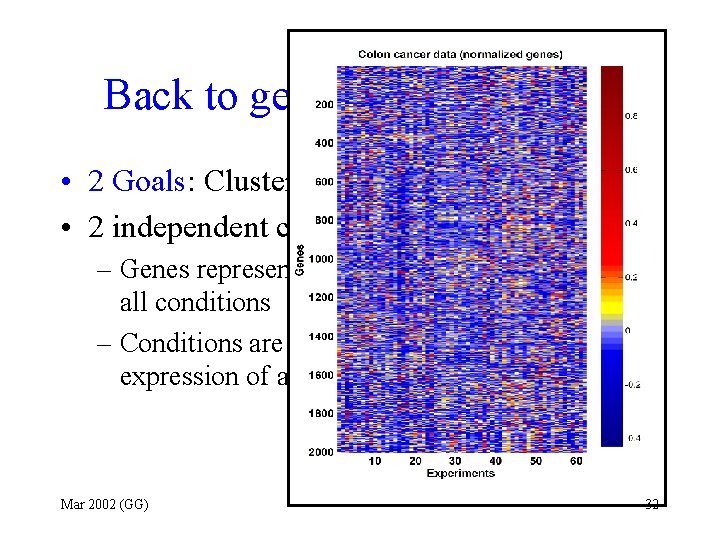
Back to gene expression data • 2 Goals: Cluster Genes and Conditions • 2 independent clustering: – Genes represented as vectors of expression in all conditions – Conditions are represented as vectors of expression of all genes Mar 2002 (GG) 32

First clustering - Experiments 1. Identify tissue classes (tumor/normal) Mar 2002 (GG) 33
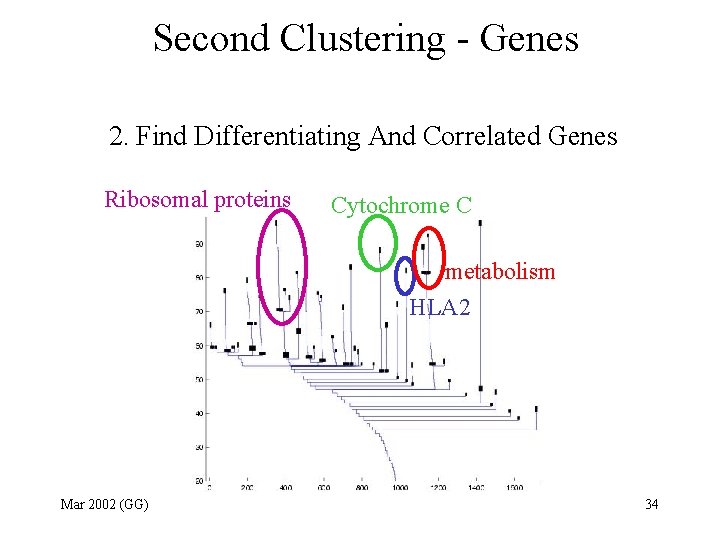
Second Clustering - Genes 2. Find Differentiating And Correlated Genes Ribosomal proteins Cytochrome C metabolism HLA 2 Mar 2002 (GG) 34
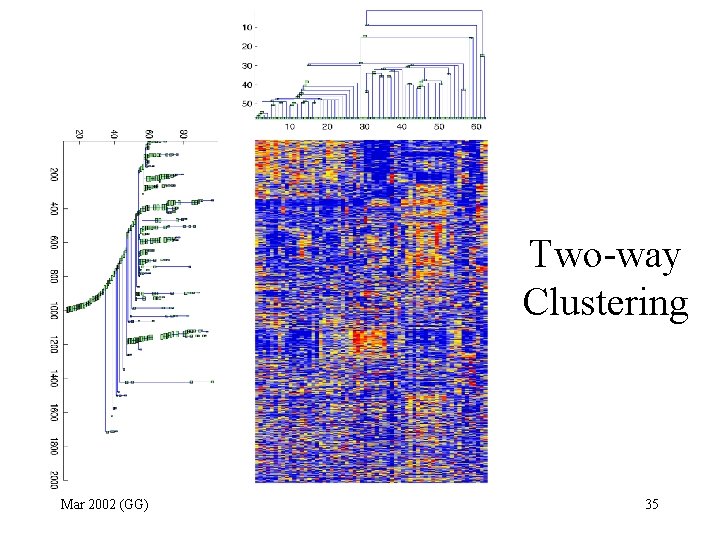
Two-way Clustering Mar 2002 (GG) 35

Coupled Two-Way Clustering (CTWC) G. Getz, E. Levine and E. Domany (2000) PNAS • Motivation: Only a small subset of genes play a role in a particular biological process; the other genes introduce noise, which may mask the signal of the important players. Only a subset of the samples exhibit the expression patterns of interest. • New Goal: Use subsets of genes to study subsets of samples (and vice versa) • A non-trivial task – exponential number of subsets. • CTWC is a heuristic to solve this problem. Mar 2002 (GG) 36

Booing Cheering Mar 2002 (GG) 37
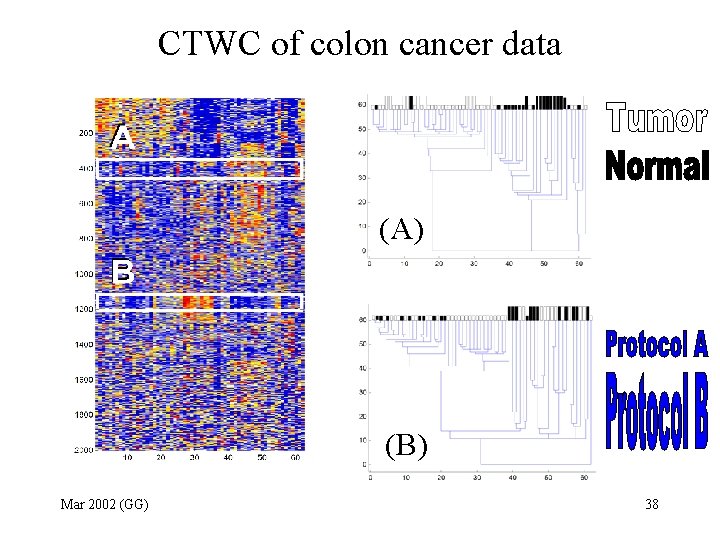
CTWC of colon cancer data (A) (B) Mar 2002 (GG) 38
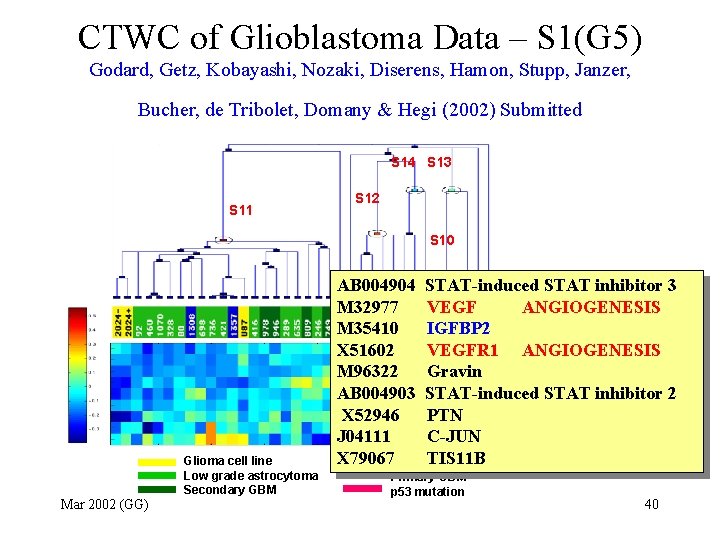
CTWC of Glioblastoma Data – S 1(G 5) Godard, Getz, Kobayashi, Nozaki, Diserens, Hamon, Stupp, Janzer, Bucher, de Tribolet, Domany & Hegi (2002) Submitted S 14 S 13 S 11 S 12 S 10 Glioma cell line Low grade astrocytoma Secondary GBM Mar 2002 (GG) AB 004904 M 32977 M 35410 X 51602 M 96322 AB 004903 X 52946 J 04111 X 79067 STAT-induced STAT inhibitor 3 VEGF ANGIOGENESIS IGFBP 2 VEGFR 1 ANGIOGENESIS Gravin STAT-induced STAT inhibitor 2 PTN C-JUN TIS 11 B Primary GBM p 53 mutation 40

Biological Work • Literature search for the genes • Genomics: search for common regulatory signal upstream of the genes • Proteomics: infer functions. • Design next experiment – get more data to validate result. • Find what is in common with sets of experiments/conditions. Mar 2002 (GG) 41

Summary • Clustering methods are used to – find genes from the same biological process – group the experiments to similar conditions • Different clustering methods can give different results. The physically motivated ones are more robust. • Focusing on subsets of the genes and conditions can uncover structure that is masked when using all genes and conditions http: //ctwc. weizmann. ac. il Mar 2002 (GG) 42
- Slides: 39