Cloning Workshop Presented by Alex Klenov Pipette Jockey

Cloning Workshop Presented by Alex Klenov (Pipette. Jockey) Step 1: Design Experiment Step 2: Prepare Fragments Step 3: Assemble Fragments Introduction - Refined to have a high success rate for most applications - Not the fastest, more upfront work with purification - Save time/money by not having to repeat an experiment - Focus on overlap based cloning techniques like Gibson/Hifi/TEDA (vs. restriction enzymes, Golden Gate as exception) Scope - For beginners -> intermediate level Step 4: Transform Cells Caveats/Limitations Step 5: Screen Colonies - Many, many other methods are out there - Some may be better suited for your application - ANY method that gives your desired clone is fair game Do Stuff with Clones

What are the general operations we want to do when cloning? Number of Fragments 1 Fragment 1 Vector 1 Insert (2 Fragments) 1 Vector >1 Insert (>2 fragments) Operation - Make a point mutation/substitution (A few bp within ~50 bp window) - Make a deletion (Any size) - Make a small insertion (Less than ~50 bp) - Transfer DNA from plasmid -> plasmid (“sub-cloning”) Insert synthesized DNA into plasmid Insert genomic DNA into plasmid Clone gene from m. RNA into plasmid - Multi-fragment assemblies Make a substitution (Alternative) Make a deletion (Alternative) Make a small insertion (Alternative) Technique KLD Mutagenesis TEDA assembly Gibson assembly Hifi assembly Golden Gate assembly
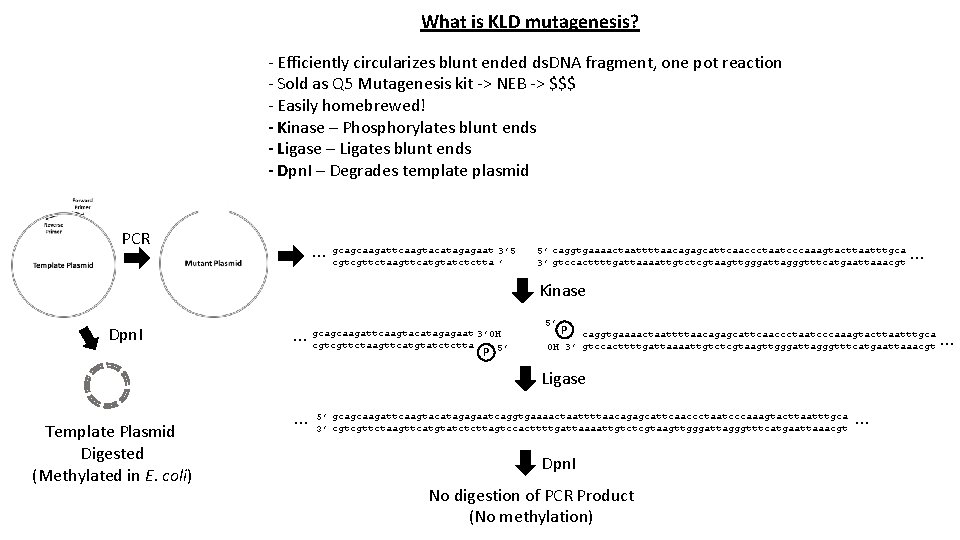
What is KLD mutagenesis? - Efficiently circularizes blunt ended ds. DNA fragment, one pot reaction - Sold as Q 5 Mutagenesis kit -> NEB -> $$$ - Easily homebrewed! - Kinase – Phosphorylates blunt ends - Ligase – Ligates blunt ends - Dpn. I – Degrades template plasmid PCR 3’ 5 … gcagcaagattcaagtacatagagaat cgtcgttctaagttcatgtatctctta ’ 5’ caggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ gtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt … Kinase Dpn. I … gcagcaagattcaagtacatagagaat 3’OH cgtcgttctaagttcatgtatctctta 5’ P caggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca OH 3’ gtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt Ligase Template Plasmid Digested (Methylated in E. coli) … 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt Dpn. I No digestion of PCR Product (No methylation) … …
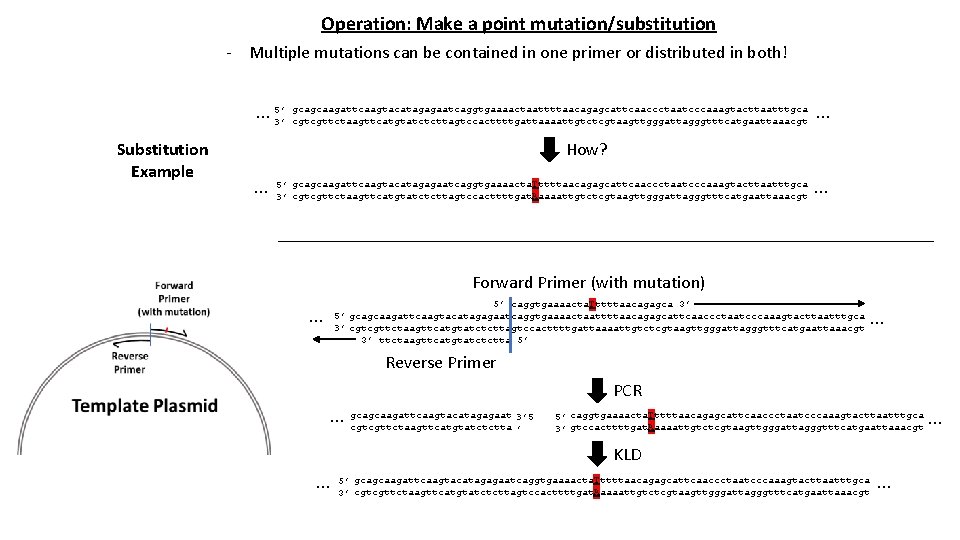
Operation: Make a point mutation/substitution - Multiple mutations can be contained in one primer or distributed in both! … 5’ 3’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca … cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt Substitution Example How? … 5’ gcagcaagattcaagtacatagagaatcaggtgaaaacta. Tttttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgat. Aaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt … Forward Primer (with mutation) … 5’ caggtgaaaacta. Tttttaacagagca 3’ 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt 3’ ttctaagttcatgtatctctta 5’ … Reverse Primer PCR 3’ 5 … gcagcaagattcaagtacatagagaat cgtcgttctaagttcatgtatctctta ’ 5’ caggtgaaaacta. Tttttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ gtccacttttgat. Aaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt KLD … 5’ gcagcaagattcaagtacatagagaatcaggtgaaaacta. Tttttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgat. Aaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt … …
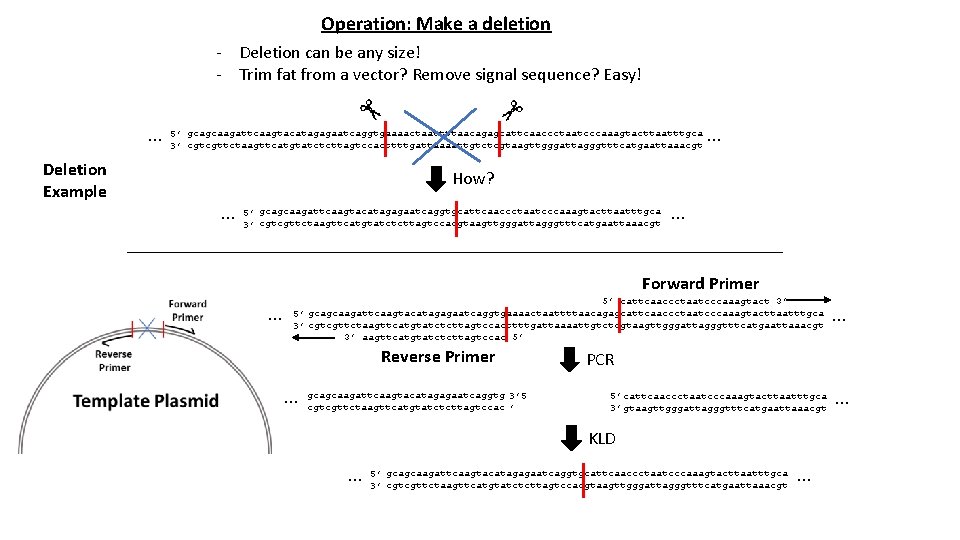
Operation: Make a deletion - Deletion can be any size! - Trim fat from a vector? Remove signal sequence? Easy! … Deletion Example 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt … How? … 5’ gcagcaagattcaagtacatagagaatcaggtgcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacgtaagttgggattagggtttcatgaattaaacgt … Forward Primer … 5’ cattcaaccctaatcccaaagtact 3’ 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt 3’ aagttcatgtatctcttagtccac 5’ Reverse Primer … gcagcaagattcaagtacatagagaatcaggtg 3’ 5 cgtcgttctaagttcatgtatctcttagtccac ’ PCR 5’ cattcaaccctaatcccaaagtacttaatttgca 3’ gtaagttgggattagggtttcatgaattaaacgt KLD … … 5’ gcagcaagattcaagtacatagagaatcaggtgcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacgtaagttgggattagggtttcatgaattaaacgt … …
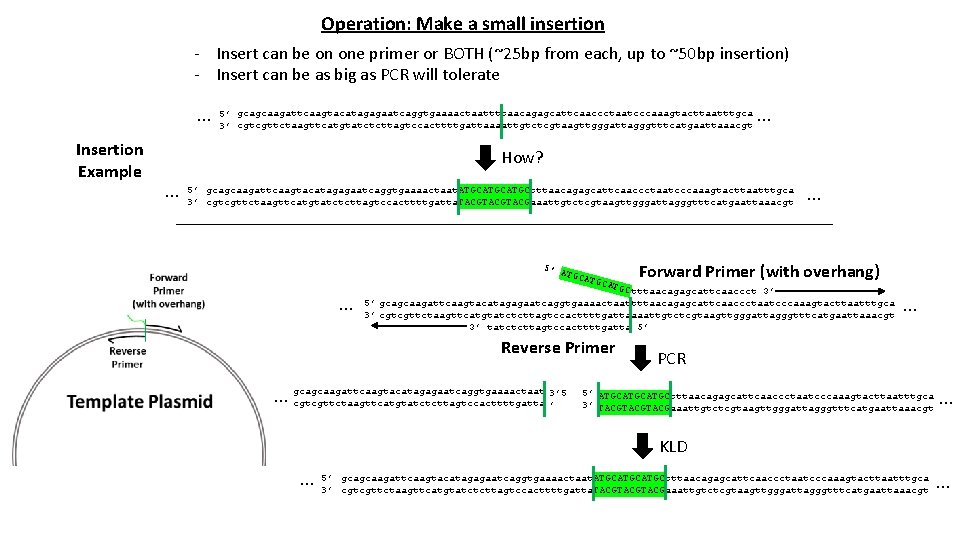
Operation: Make a small insertion - Insert can be on one primer or BOTH (~25 bp from each, up to ~50 bp insertion) - Insert can be as big as PCR will tolerate … Insertion Example 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt … How? … 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaat. ATGCATGCtttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgatta. TACGTACGaaattgtctcgtaagttgggattagggtttcatgaattaaacgt … Forward Primer (with overhang) … 5’ A TGCA TGC tttaacagagcattcaaccct 3’ 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt 3’ tatctcttagtccacttttgatta 5’ Reverse Primer 3’ 5 … gcagcaagattcaagtacatagagaatcaggtgaaaactaat cgtcgttctaagttcatgtatctcttagtccacttttgatta ’ … PCR 5’ ATGCATGCtttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ TACGTACGaaattgtctcgtaagttgggattagggtttcatgaattaaacgt … KLD … 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaat. ATGCATGCtttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgatta. TACGTACGaaattgtctcgtaagttgggattagggtttcatgaattaaacgt …

What are the general operations we want to do when cloning? Number of Fragments 1 Fragment 1 Vector 1 Insert (2 Fragments) 1 Vector >1 Insert (>2 fragments) Operation - Make a point mutation/substitution (A few bp within ~50 bp window) - Make a deletion (Any size) - Make a small insertion (Less than ~50 bp) - Transfer DNA from plasmid -> plasmid (“sub-cloning”) Insert synthesized DNA into plasmid Insert genomic DNA into plasmid Clone gene from m. RNA into plasmid - Multi-fragment assemblies Make a substitution (Alternative) Make a deletion (Alternative) Make a small insertion (Alternative) Technique KLD Mutagenesis TEDA assembly Gibson assembly Hifi assembly Golden Gate assembly

What is TEDA / Gibson / Hifi assembly? - Overlap based cloning techniques - Same principle for all, different blends of enzymes, different efficiencies - Varies widely in cost, easy to homebrew Insert CATCATCATCACAGCAGCGGCatgaaaccgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat TGAGATCCGGCTGCTAACAAA GTAGTAGTAGTGTCGTCGCCGtactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaattttaagta ACTCTAGGCCGACGATTGTTT AGCAGCCATCATCATCACAGCAGCGGC TCGTCGGTAGTAGTAGTGTCGTCGCCG TGAGATCCGGCTGCTAACAAAGCCCGAAAGGAAGCTGAGTTG ACTCTAGGCCGACGATTGTTTCGGGCTTTCCTTCGACTCAAC Destination Vector T 5 Exo Polymerase AGCAGCCATCATCATCACAGCAGCGGC GAAGCTGAGTTG accgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat TGAGATCCGGCTGCTAACAAA TCGTCGG ACTCTAGGCCGACGATTGTTTCGGGCTTTCCTTCGACTCAAC GTAGTAGTAGTGTCGTCGCCGtactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaatttta Polymerase + Ligase (Gibson mix) AGCAGCCATCATCATCACAGCAGCGGC atgaaaccgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat TGAGATCCGGCTGCTAACAAAGCCCGAAAGGAAGCTGAGTTG TCGTCGGTAGTAGTAGTGTCGTCGCCG tactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaattttaagta ACTCTAGGCCGACGATTGTTTCGGGCTTTCCTTCGACTCAAC Noice! Ligase
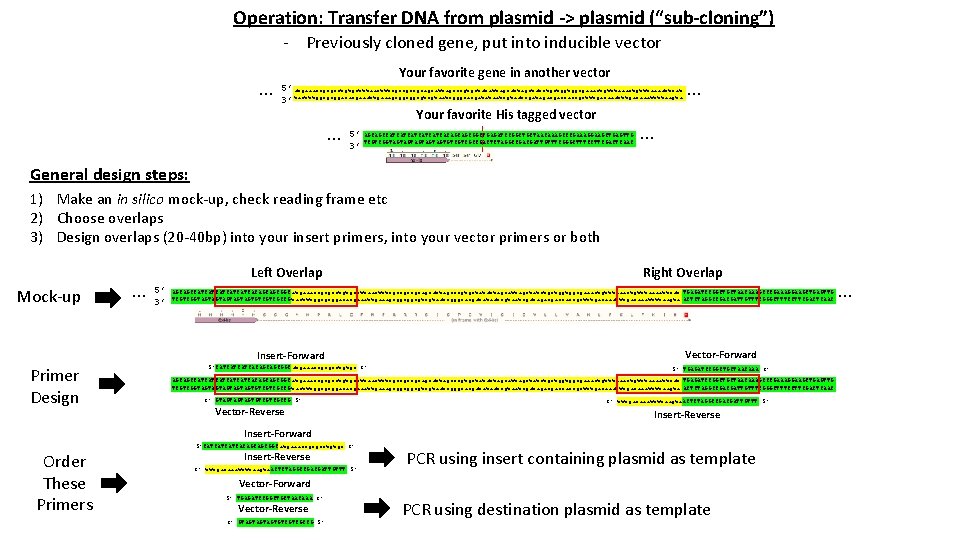
Operation: Transfer DNA from plasmid -> plasmid (“sub-cloning”) - Previously cloned gene, put into inducible vector … Your favorite gene in another vector 5’ atgaaaccgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat 3’ tactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaattttaagta … Your favorite His tagged vector 5’ AGCAGCCATCATCATCACAGCAGCGGCTGAGATCCGGCTGCTAACAAAGCCCGAAAGGAAGCTGAGTTG 3’ TCGTCGGTAGTAGTAGTGTCGTCGCCGACTCTAGGCCGACGATTGTTTCGGGCTTTCCTTCGACTCAAC … … General design steps: 1) Make an in silico mock-up, check reading frame etc 2) Choose overlaps 3) Design overlaps (20 -40 bp) into your insert primers, into your vector primers or both Mock-up … Left Overlap 5’ 3’ AGCAGCCATCATCATCACAGCAGCGGC atgaaaccgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat TGAGATCCGGCTGCTAACAAAGCCCGAAAGGAAGCTGAGTTG TCGTCGGTAGTAGTAGTGTCGTCGCCG tactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaattttaagta ACTCTAGGCCGACGATTGTTTCGGGCTTTCCTTCGACTCAAC Insert-Forward Primer Design Right Overlap 5’CATCATCATCACAGCAGCGGCatgaaaccgcgcctgtgc 3’ Vector-Forward 5’ TGAGATCCGGCTGCTAACAAA 3’ AGCAGCCATCATCATCACAGCAGCGGC atgaaaccgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat TGAGATCCGGCTGCTAACAAAGCCCGAAAGGAAGCTGAGTTG TCGTCGGTAGTAGTAGTGTCGTCGCCG tactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaattttaagta ACTCTAGGCCGACGATTGTTTCGGGCTTTCCTTCGACTCAAC 3’ GTAGTAGTAGTGTCGTCGCCG 5’ Vector-Reverse 3’ tttgacaaattttaagta. ACTCTAGGCCGACGATTGTTT 5’ Insert-Reverse Insert-Forward 5’CATCATCATCACAGCAGCGGCatgaaaccgcgcctgtgc 3’ Order These Primers Insert-Reverse 3’ tttgacaaattttaagta. ACTCTAGGCCGACGATTGTTT 5’ PCR using insert containing plasmid as template Vector-Forward 5’ TGAGATCCGGCTGCTAACAAA 3’ Vector-Reverse 3’ GTAGTAGTAGTGTCGTCGCCG 5’ PCR using destination plasmid as template …
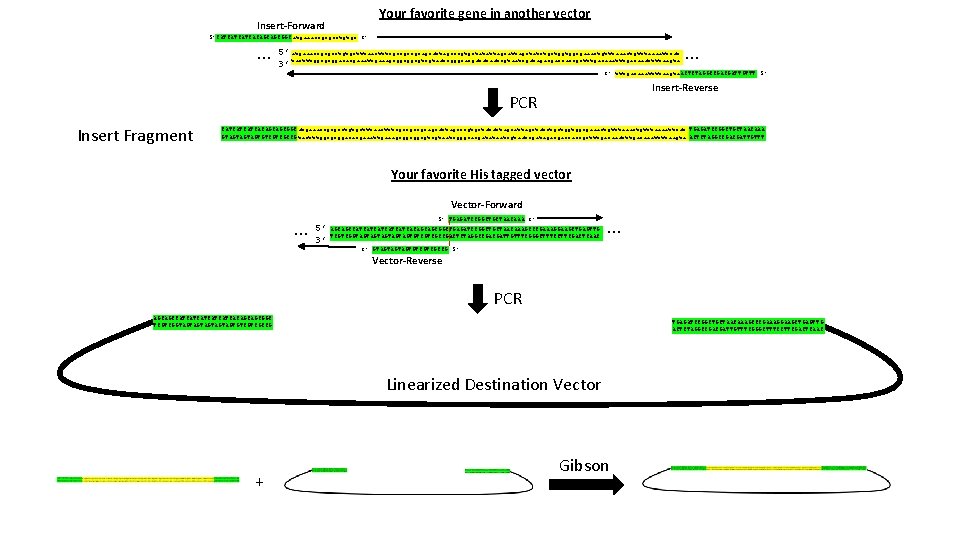
Your favorite gene in another vector Insert-Forward 5’CATCATCATCACAGCAGCGGCatgaaaccgcgcctgtgc 3’ … 5’ atgaaaccgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat 3’ tactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaattttaagta … 3’ tttgacaaattttaagta. ACTCTAGGCCGACGATTGTTT 5’ Insert-Reverse PCR Insert Fragment CATCATCATCACAGCAGCGGCatgaaaccgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat TGAGATCCGGCTGCTAACAAA GTAGTAGTAGTGTCGTCGCCGtactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaattttaagta ACTCTAGGCCGACGATTGTTT Your favorite His tagged vector Vector-Forward … 5’ TGAGATCCGGCTGCTAACAAA 3’ 5’ AGCAGCCATCATCATCACAGCAGCGGCTGAGATCCGGCTGCTAACAAAGCCCGAAAGGAAGCTGAGTTG 3’ TCGTCGGTAGTAGTAGTGTCGTCGCCGACTCTAGGCCGACGATTGTTTCGGGCTTTCCTTCGACTCAAC … 3’ GTAGTAGTAGTGTCGTCGCCG 5’ Vector-Reverse PCR AGCAGCCATCATCATCACAGCAGCGGC TCGTCGGTAGTAGTAGTGTCGTCGCCG TGAGATCCGGCTGCTAACAAAGCCCGAAAGGAAGCTGAGTTG ACTCTAGGCCGACGATTGTTTCGGGCTTTCCTTCGACTCAAC Linearized Destination Vector + Gibson
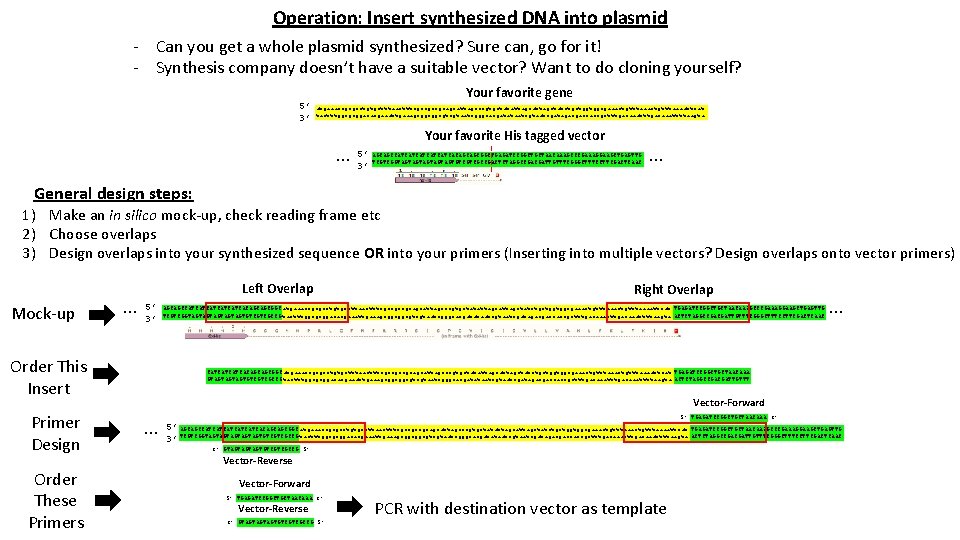
Operation: Insert synthesized DNA into plasmid - Can you get a whole plasmid synthesized? Sure can, go for it! - Synthesis company doesn’t have a suitable vector? Want to do cloning yourself? Your favorite gene 5’ 3’ atgaaaccgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat tactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaattttaagta … Your favorite His tagged vector 5’ AGCAGCCATCATCATCACAGCAGCGGCTGAGATCCGGCTGCTAACAAAGCCCGAAAGGAAGCTGAGTTG 3’ TCGTCGGTAGTAGTAGTGTCGTCGCCGACTCTAGGCCGACGATTGTTTCGGGCTTTCCTTCGACTCAAC … General design steps: 1) Make an in silico mock-up, check reading frame etc 2) Choose overlaps 3) Design overlaps into your synthesized sequence OR into your primers (Inserting into multiple vectors? Design overlaps onto vector primers) Mock-up … Left Overlap 5’ 3’ Order This Insert Primer Design Order These Primers Right Overlap AGCAGCCATCATCATCACAGCAGCGGC atgaaaccgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat TGAGATCCGGCTGCTAACAAAGCCCGAAAGGAAGCTGAGTTG TCGTCGGTAGTAGTAGTGTCGTCGCCG tactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaattttaagta ACTCTAGGCCGACGATTGTTTCGGGCTTTCCTTCGACTCAAC … CATCATCATCACAGCAGCGGCatgaaaccgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat TGAGATCCGGCTGCTAACAAA GTAGTAGTAGTGTCGTCGCCGtactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaattttaagta ACTCTAGGCCGACGATTGTTT Vector-Forward … 5’ TGAGATCCGGCTGCTAACAAA 3’ 5’ AGCAGCCATCATCATCACAGCAGCGGC atgaaaccgcgcctgtgctttaactttcgccgccgcagcattagcccgtgctatattagctatctgctggtggcgaaactgtttaaaattcat TGAGATCCGGCTGCTAACAAAGCCCGAAAGGAAGCTGAGTTG 3’ TCGTCGGTAGTAGTAGTGTCGTCGCCG tactttggcgcggacacgaaattgaaagcggcggcgtcgtaatcgggcacgatataatcgatagacgaccaccgctttgacaaattttaagta ACTCTAGGCCGACGATTGTTTCGGGCTTTCCTTCGACTCAAC 3’ GTAGTAGTAGTGTCGTCGCCG 5’ Vector-Reverse Vector-Forward 5’ TGAGATCCGGCTGCTAACAAA 3’ Vector-Reverse 3’ GTAGTAGTAGTGTCGTCGCCG 5’ PCR with destination vector as template
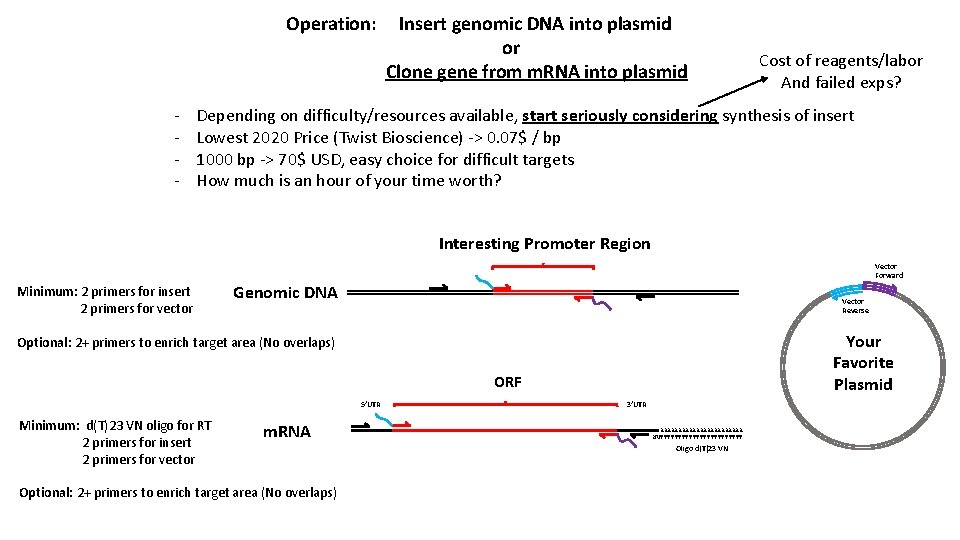
Operation: - Insert genomic DNA into plasmid or Clone gene from m. RNA into plasmid Cost of reagents/labor And failed exps? Depending on difficulty/resources available, start seriously considering synthesis of insert Lowest 2020 Price (Twist Bioscience) -> 0. 07$ / bp 1000 bp -> 70$ USD, easy choice for difficult targets How much is an hour of your time worth? Interesting Promoter Region Minimum: 2 primers for insert 2 primers for vector Vector Forward Genomic DNA Vector Reverse Your Favorite Plasmid Optional: 2+ primers to enrich target area (No overlaps) ORF 5’UTR Minimum: d(T)23 VN oligo for RT 2 primers for insert 2 primers for vector m. RNA Optional: 2+ primers to enrich target area (No overlaps) 3’UTR AAAAAAAAAAAA NVTTTTTTTTTTTT Oligo d(T)23 VN

What are the general operations we want to do when cloning? Number of Fragments 1 Fragment 1 Vector 1 Insert (2 Fragments) 1 Vector >1 Insert (>2 fragments) Operation - Make a point mutation/substitution (A few bp within ~50 bp window) - Make a deletion (Any size) - Make a small insertion (Less than ~50 bp) - Transfer DNA from plasmid -> plasmid (“sub-cloning”) Insert synthesized DNA into plasmid Insert genomic DNA into plasmid Clone gene from m. RNA into plasmid - Multi-fragment assemblies Make a substitution (Alternative) Make a deletion (Alternative) Make a small insertion (Alternative) Technique KLD Mutagenesis TEDA assembly Gibson assembly Hifi assembly Golden Gate assembly
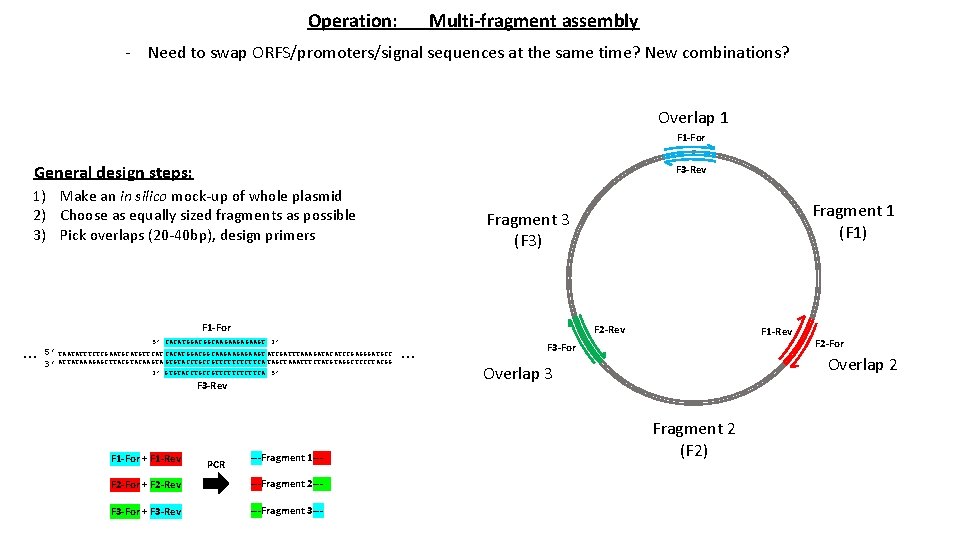
Operation: Multi-fragment assembly - Need to swap ORFS/promoters/signal sequences at the same time? New combinations? Overlap 1 F 1 -For General design steps: F 3 -Rev 1) Make an in silico mock-up of whole plasmid 2) Choose as equally sized fragments as possible 3) Pick overlaps (20 -40 bp), design primers F 1 -For … F 2 -Rev 5’ CACATGGACGGCAAGAAGT 3’ 5’ TAATATTTCTCGAATGCATGTTCAT CACATGGACGGCAAGAAGT ATCGATTTAAAGATACATCCGAGGGATGCC 3’ ATTATAAAGAGCTTACGTACAAGTA GTGTACCTGCCGTTCTTCA TAGCTAAATTTCTATGTAGGCTCCCTACGG 3’ GTGTACCTGCCGTTCTTCA 5’ F 3 -Rev F 1 -For + F 1 -Rev PCR Fragment 1 (F 1) Fragment 3 (F 3) ---Fragment 1 --- F 2 -For + F 2 -Rev ---Fragment 2 --- F 3 -For + F 3 -Rev ---Fragment 3 --- … F 1 -Rev F 3 -For F 2 -For Overlap 2 Overlap 3 Fragment 2 (F 2)
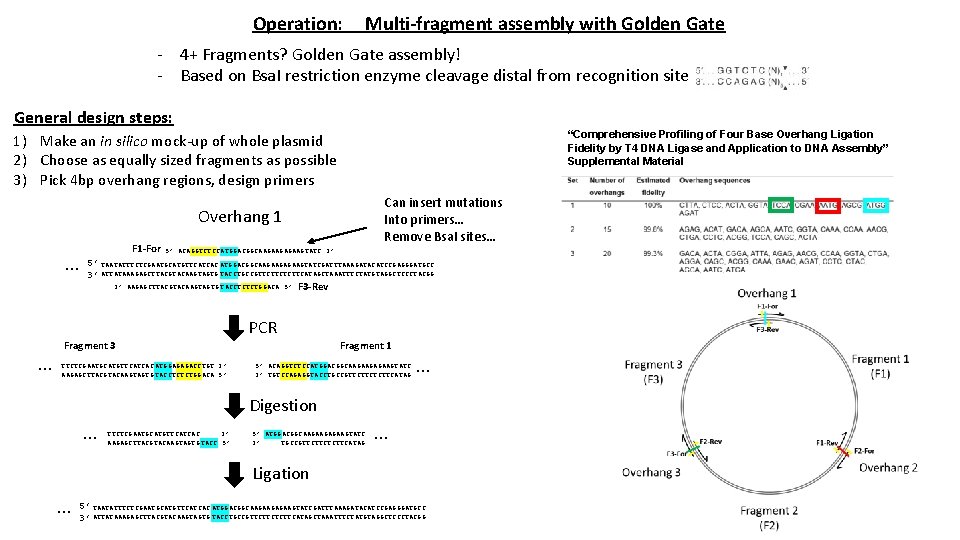
Operation: Multi-fragment assembly with Golden Gate - 4+ Fragments? Golden Gate assembly! - Based on Bsa. I restriction enzyme cleavage distal from recognition site General design steps: “Comprehensive Profiling of Four Base Overhang Ligation Fidelity by T 4 DNA Ligase and Application to DNA Assembly” Supplemental Material 1) Make an in silico mock-up of whole plasmid 2) Choose as equally sized fragments as possible 3) Pick 4 bp overhang regions, design primers Can insert mutations Into primers… Remove Bsa. I sites… Overhang 1 … F 1 -For 5’ ACAGGTCTCCATGGACGGCAAGAAGTATC 3’ 5’ TAATATTTCTCGAATGCATGTTCATCAC ATGGACGGCAAGAAGTATCGATTTAAAGATACATCCGAGGGATGCC 3’ ATTATAAAGAGCTTACGTACAAGTAGTG TACCTGCCGTTCTTCATAGCTAAATTTCTATGTAGGCTCCCTACGG 3’ AAGAGCTTACGTACAAGTAGTG TACCTCTCTGGACA 5’ F 3 -Rev PCR … Fragment 1 Fragment 3 TTCTCGAATGCATGTTCATCAC ATGGAGAGACCTGT 3’ AAGAGCTTACGTACAAGTAGTG TACCTCTCTGGACA 5’ 5’ ACAGGTCTCCATGGACGGCAAGAAGTATC 3’ TGTCCAGAGGTACCTGCCGTTCTTCATAG … Digestion … TTCTCGAATGCATGTTCATCAC 3’ AAGAGCTTACGTACAAGTAGTG TACC 5’ 5’ ATGGACGGCAAGAAGTATC 3’ TGCCGTTCTTCATAG … Ligation … 5’ TAATATTTCTCGAATGCATGTTCATCAC ATGGACGGCAAGAAGTATCGATTTAAAGATACATCCGAGGGATGCC 3’ ATTATAAAGAGCTTACGTACAAGTAGTG TACCTGCCGTTCTTCATAGCTAAATTTCTATGTAGGCTCCCTACGG
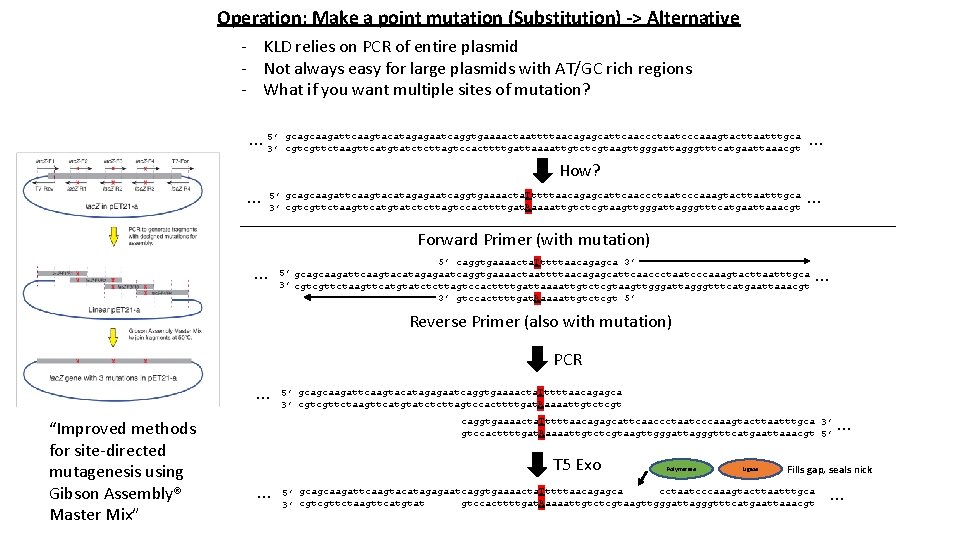
Operation: Make a point mutation (Substitution) -> Alternative - KLD relies on PCR of entire plasmid - Not always easy for large plasmids with AT/GC rich regions - What if you want multiple sites of mutation? … 5’ 3’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca … cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt How? … 5’ gcagcaagattcaagtacatagagaatcaggtgaaaacta. Tttttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgat. Aaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt … Forward Primer (with mutation) … 5’ caggtgaaaacta. Tttttaacagagca 3’ 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt 3’ gtccacttttgat. Aaaaattgtctcgt 5’ … Reverse Primer (also with mutation) PCR … “Improved methods for site-directed mutagenesis using Gibson Assembly® Master Mix” 5’ gcagcaagattcaagtacatagagaatcaggtgaaaacta. Tttttaacagagca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgat. Aaaaattgtctcgt caggtgaaaacta. Tttttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ gtccacttttgat. Aaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt 5’ T 5 Exo … Polymerase Ligase … Fills gap, seals nick cctaatcccaaagtacttaatttgca 5’ gcagcaagattcaagtacatagagaatcaggtgaaaacta. Tttttaacagagca gtccacttttgat. Aaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt 3’ cgtcgttctaagttcatgtat …
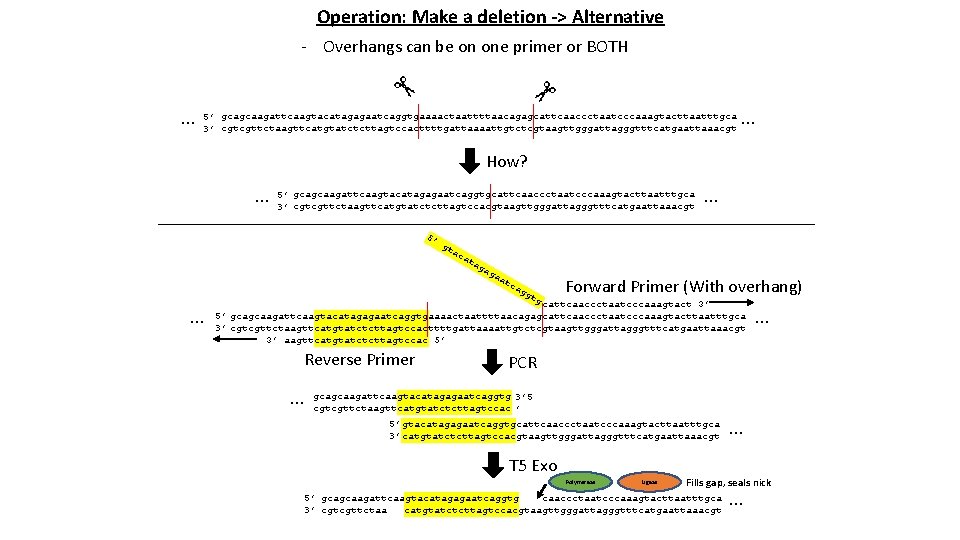
Operation: Make a deletion -> Alternative - Overhangs can be on one primer or BOTH … 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt … How? … 5’ gcagcaagattcaagtacatagagaatcaggtgcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacgtaagttgggattagggtttcatgaattaaacgt 5’ gt ac at ag ag a at c ag … … gt g Forward Primer (With overhang) cattcaaccctaatcccaaagtact 3’ 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt 3’ aagttcatgtatctcttagtccac 5’ Reverse Primer … … PCR gcagcaagattcaagtacatagagaatcaggtg 3’ 5 cgtcgttctaagttcatgtatctcttagtccac ’ 5’ gtacatagagaatcaggtgcattcaaccctaatcccaaagtacttaatttgca 3’ catgtatctcttagtccacgtaagttgggattagggtttcatgaattaaacgt T 5 Exo Polymerase Ligase … Fills gap, seals nick 5’ gcagcaagattcaagtacatagagaatcaggtg caaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaa catgtatctcttagtccacgtaagttgggattagggtttcatgaattaaacgt …
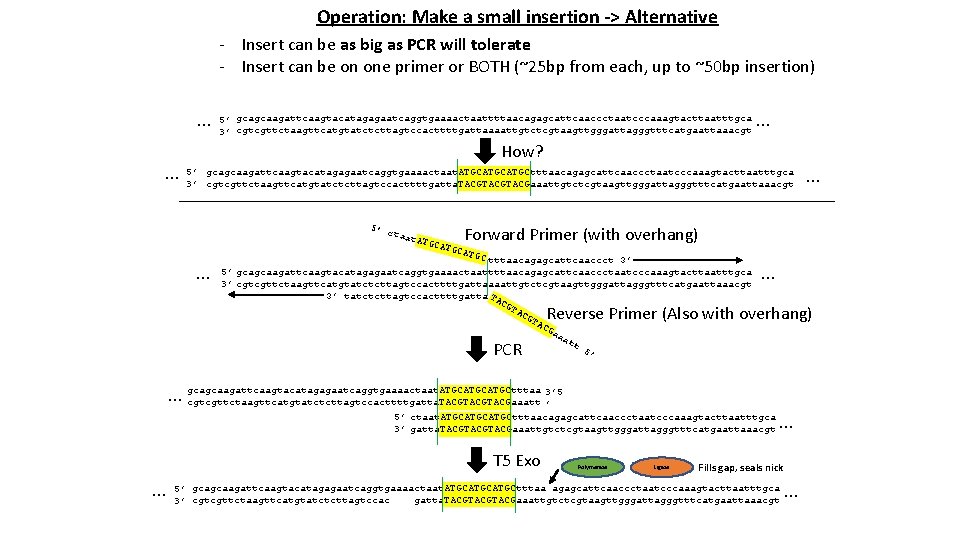
Operation: Make a small insertion -> Alternative - Insert can be as big as PCR will tolerate - Insert can be on one primer or BOTH (~25 bp from each, up to ~50 bp insertion) … … 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt … How? 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaat. ATGCATGCtttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgatta. TACGTACGaaattgtctcgtaagttgggattagggtttcatgaattaaacgt 5’ c taat ATGC … … Forward Primer (with overhang) ATGC tttaacagagcattcaaccct 3’ 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaattttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ cgtcgttctaagttcatgtatctcttagtccacttttgattaaaattgtctcgtaagttgggattagggtttcatgaattaaacgt 3’ tatctcttagtccacttttgatta TA CG aa at t 5’ … Reverse Primer (Also with overhang) PCR 3’ 5 … gcagcaagattcaagtacatagagaatcaggtgaaaactaat. ATGCATGCtttaa cgtcgttctaagttcatgtatctcttagtccacttttgatta. TACGTACGaaatt ’ 5’ ctaat. ATGCATGCtttaacagagcattcaaccctaatcccaaagtacttaatttgca 3’ gatta. TACGTACGaaattgtctcgtaagttgggattagggtttcatgaattaaacgt T 5 Exo … Polymerase Ligase … Fills gap, seals nick 5’ gcagcaagattcaagtacatagagaatcaggtgaaaactaat. ATGCATGCtttaa agagcattcaaccctaatcccaaagtacttaatttgca gatta. TACGTACGaaattgtctcgtaagttgggattagggtttcatgaattaaacgt 3’ cgtcgttctaagttcatgtatctcttagtccac …
- Slides: 18