Cloning Overview 42919 Monday Night Meeting Ben Maldonato

Cloning Overview 4/29/19 Monday Night Meeting Ben Maldonato 1

Helpful Tools Oligonucleotide (Primer) Analysis Tools: • https: //www. eurofinsgenomics. eu/en/ecom/tools/oligo-analysis/ • https: //www. genscript. com/tools/pcr-primers-designer New England Biolabs (NEB) Tool Suite: • https: //www. neb. com/tools-and-resources/interactive-tools 2

Other Protocol Walkthroughs • https: //www. addgene. org/protocols/pcr-cloning/ • https: //www. addgene. org/protocols/ 3

Outline • Background and Intro • Plasmids and Why We Care • Base Pairing and Reverse Complement • Restriction Enzymes • Sticky vs Blunt Ends • Polymerase Chain Reaction (PCR) • Primer Design and Considerations • Restriction Digestion • Ligation and Transformation • Sequencing • Walkthrough with Data! 4
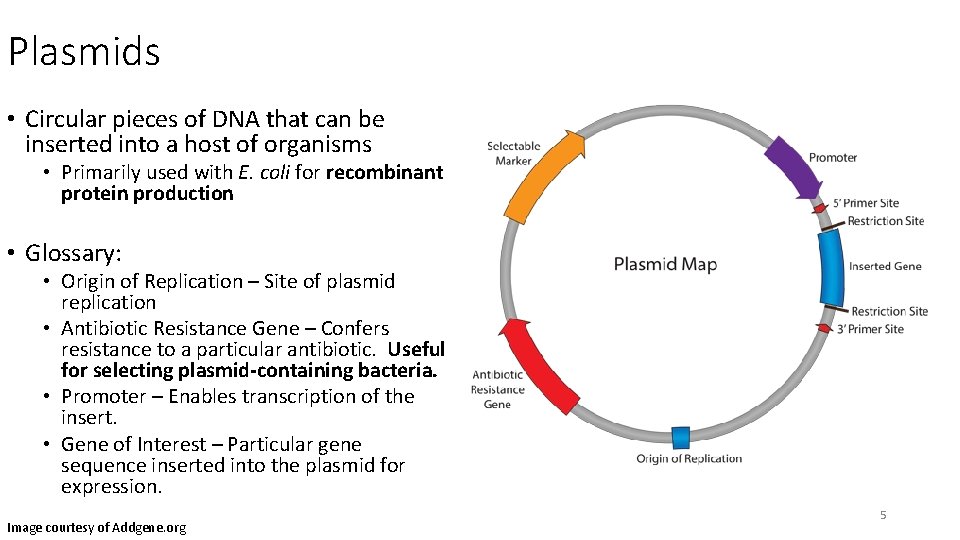
Plasmids • Circular pieces of DNA that can be inserted into a host of organisms • Primarily used with E. coli for recombinant protein production • Glossary: • Origin of Replication – Site of plasmid replication • Antibiotic Resistance Gene – Confers resistance to a particular antibiotic. Useful for selecting plasmid-containing bacteria. • Promoter – Enables transcription of the insert. • Gene of Interest – Particular gene sequence inserted into the plasmid for expression. Image courtesy of Addgene. org 5

DNA Structure and Reverse Complement • Adenine (A) and thymine (T) form a pair with 2 hydrogen bonds • Guanine (G) and cytosine (C) form a pair with 3 hydrogen bonds • Using the top sequence “ATGCGCT…” as our template strand: • The reverse sequence is the top strand read from 3’ to 5’ • The complement sequence is the bottom strand read from 3’ to 5’ • The reverse complement is the bottom strand read from 5’ to 3’ • This is what we would synthesize if we were designing a reverse primer as it binds correctly to the 6 template strand. Image courtesy of Wikipedia. org
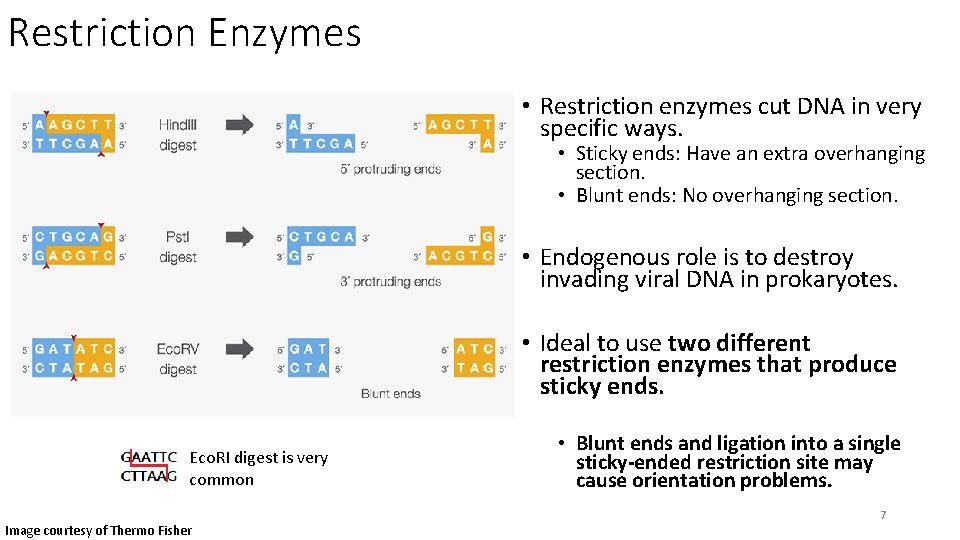
Restriction Enzymes • Restriction enzymes cut DNA in very specific ways. • Sticky ends: Have an extra overhanging section. • Blunt ends: No overhanging section. • Endogenous role is to destroy invading viral DNA in prokaryotes. • Ideal to use two different restriction enzymes that produce sticky ends. Eco. RI digest is very common Image courtesy of Thermo Fisher • Blunt ends and ligation into a single sticky-ended restriction site may cause orientation problems. 7
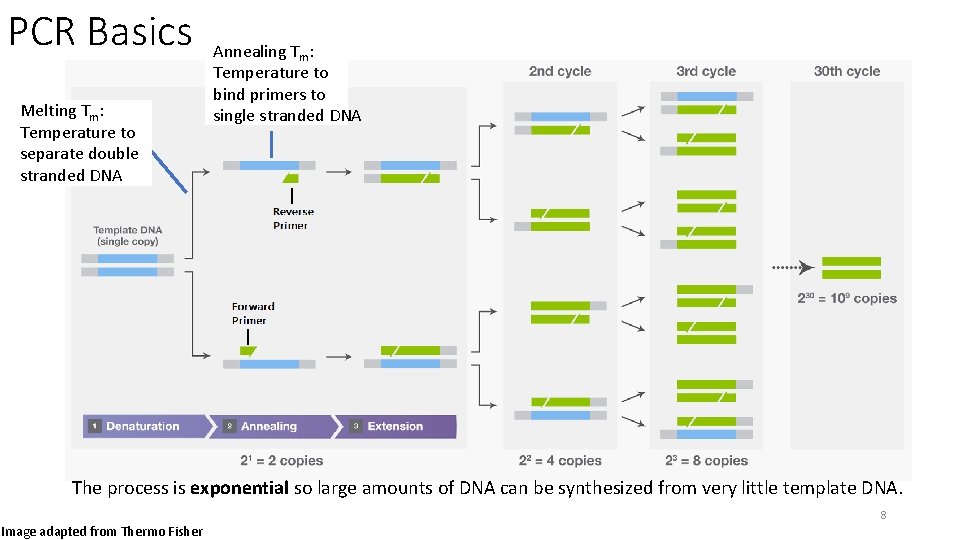
PCR Basics Melting Tm: Temperature to separate double stranded DNA Annealing Tm: Temperature to bind primers to single stranded DNA The process is exponential so large amounts of DNA can be synthesized from very little template DNA. 8 Image adapted from Thermo Fisher
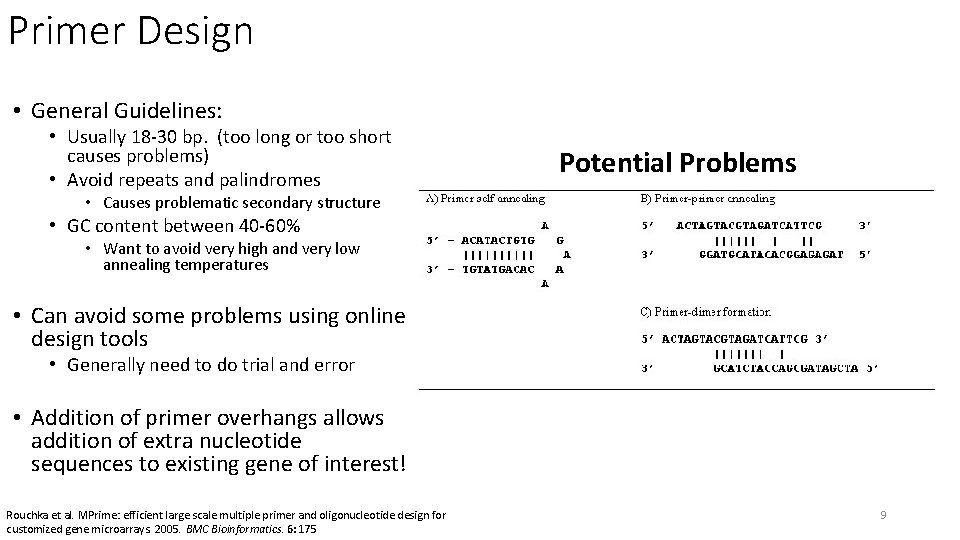
Primer Design • General Guidelines: • Usually 18 -30 bp. (too long or too short causes problems) • Avoid repeats and palindromes Potential Problems • Causes problematic secondary structure • GC content between 40 -60% • Want to avoid very high and very low annealing temperatures • Can avoid some problems using online design tools • Generally need to do trial and error • Addition of primer overhangs allows addition of extra nucleotide sequences to existing gene of interest! Rouchka et al. MPrime: efficient large scale multiple primer and oligonucleotide design for customized gene microarrays. 2005. BMC Bioinformatics. 6: 175 9
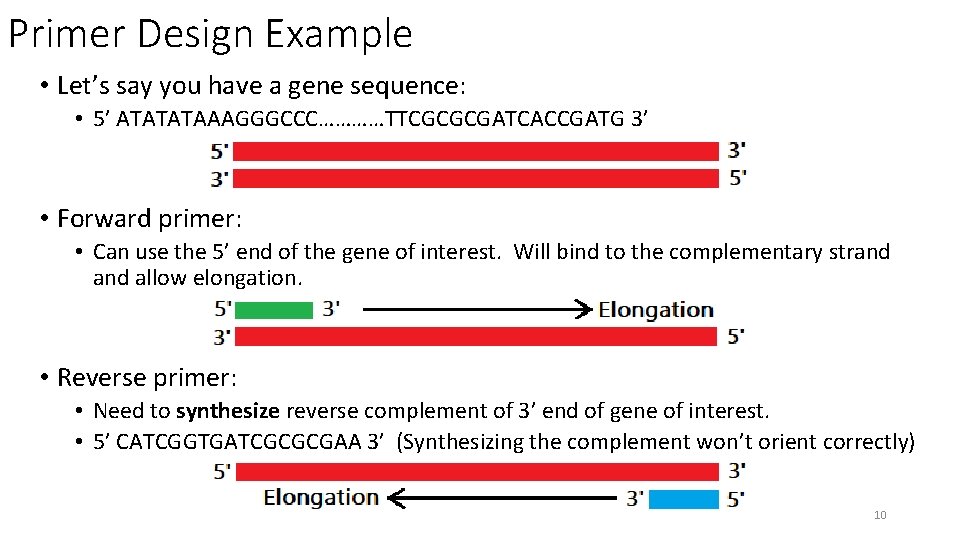
Primer Design Example • Let’s say you have a gene sequence: • 5’ ATATATAAAGGGCCC…………TTCGCGCGATCACCGATG 3’ • Forward primer: • Can use the 5’ end of the gene of interest. Will bind to the complementary strand allow elongation. • Reverse primer: • Need to synthesize reverse complement of 3’ end of gene of interest. • 5’ CATCGGTGATCGCGCGAA 3’ (Synthesizing the complement won’t orient correctly) 10
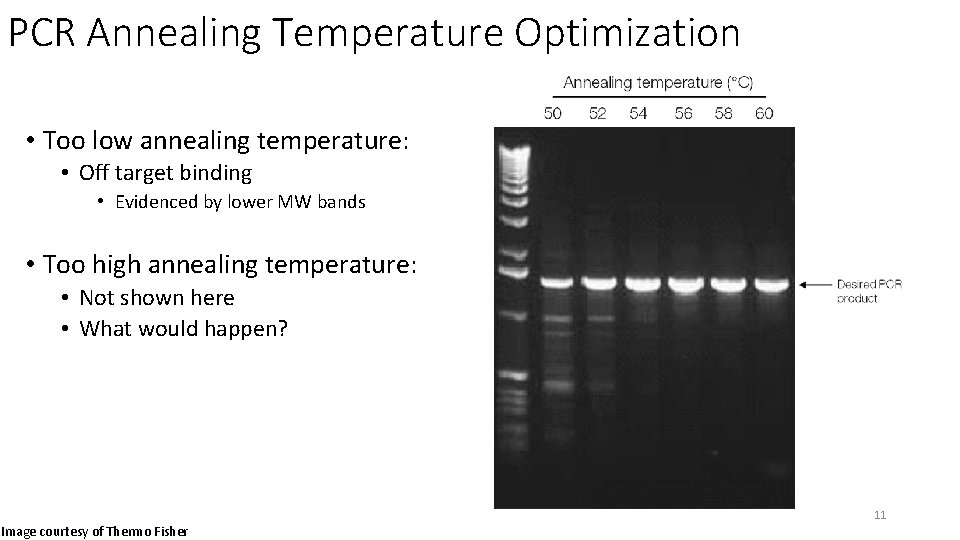
PCR Annealing Temperature Optimization • Too low annealing temperature: • Off target binding • Evidenced by lower MW bands • Too high annealing temperature: • Not shown here • What would happen? • No PCR products observed usually • Too warm for the primers to efficiently bind to the single stranded DNA before the next PCR cycle starts. 11 Image courtesy of Thermo Fisher
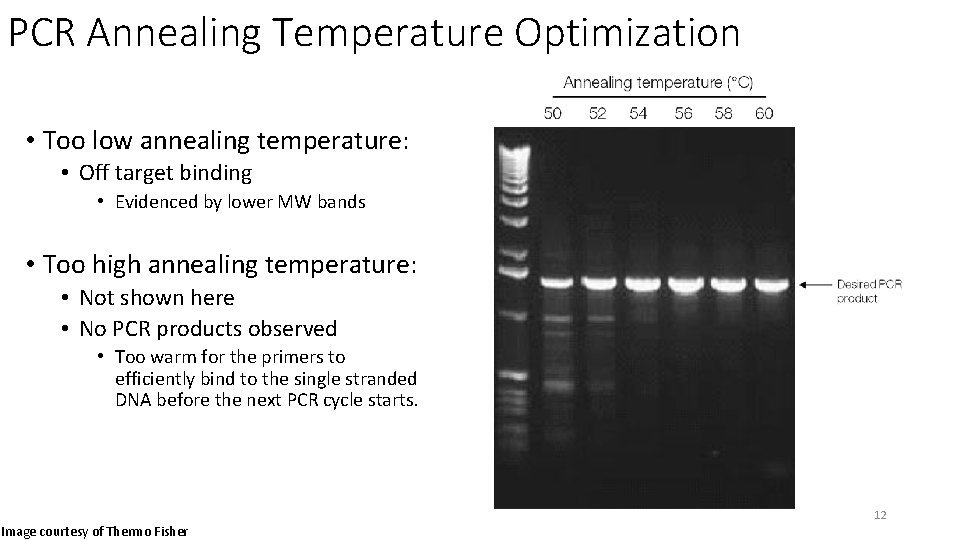
PCR Annealing Temperature Optimization • Too low annealing temperature: • Off target binding • Evidenced by lower MW bands • Too high annealing temperature: • Not shown here • No PCR products observed • Too warm for the primers to efficiently bind to the single stranded DNA before the next PCR cycle starts. 12 Image courtesy of Thermo Fisher
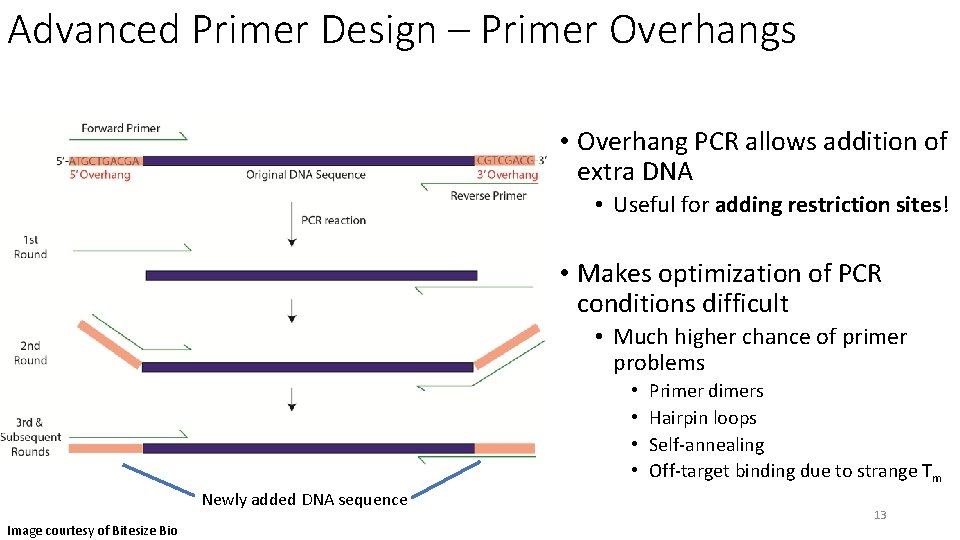
Advanced Primer Design – Primer Overhangs • Overhang PCR allows addition of extra DNA • Useful for adding restriction sites! • Makes optimization of PCR conditions difficult • Much higher chance of primer problems • • Newly added DNA sequence Image courtesy of Bitesize Bio Primer dimers Hairpin loops Self-annealing Off-target binding due to strange Tm 13
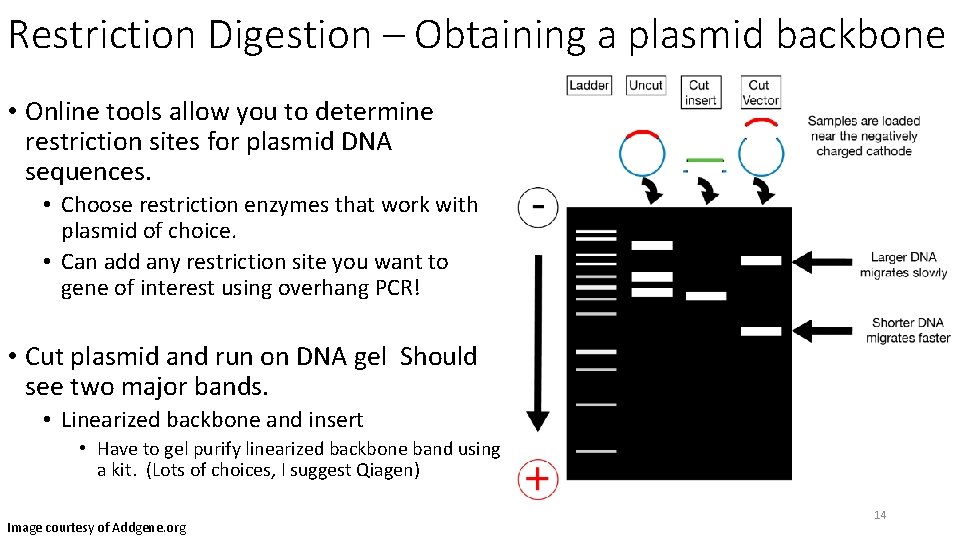
Restriction Digestion – Obtaining a plasmid backbone • Online tools allow you to determine restriction sites for plasmid DNA sequences. • Choose restriction enzymes that work with plasmid of choice. • Can add any restriction site you want to gene of interest using overhang PCR! • Cut plasmid and run on DNA gel Should see two major bands. • Linearized backbone and insert • Have to gel purify linearized backbone band using a kit. (Lots of choices, I suggest Qiagen) Image courtesy of Addgene. org 14

Ligation – Formation of a New Plasmid • Combine digested insert and linearized backbone in presence of DNA ligase and ATP. • Shoot for a molar ratio of 1: 3 backbone: insert • May need to alter if ligation isn’t working well • Very important that backbone has been purified! • Only want your insert present, not the original one too. • As always, use a kit. Qiagen is good Image courtesy of Addgene. org 15

Transformation – First Chance at Plasmid Verification • Heat shock transformation protocol with E. coli host strain (DH 5α, Stellar, etc) • Plate multiple different amounts of transformed stock • Helps give appropriate colony density without having to guess transformation efficiency • Make sure LB agar plates contain correct antibiotic/selection marker! • Check plates for transformant colonies 12 -24 hours later 16 • Can include a non-transformed negative control and a known plasmid positive control Image courtesy of Thermo Fisher
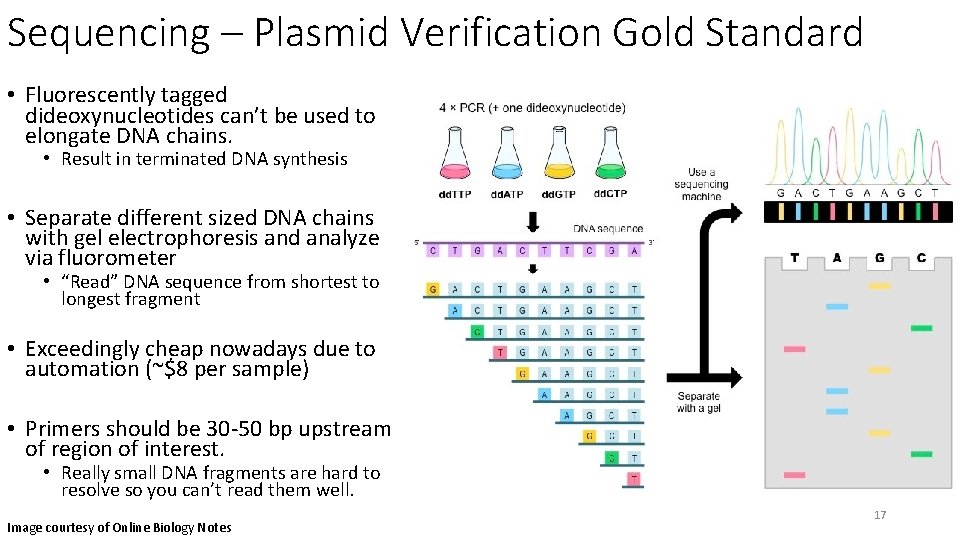
Sequencing – Plasmid Verification Gold Standard • Fluorescently tagged dideoxynucleotides can’t be used to elongate DNA chains. • Result in terminated DNA synthesis • Separate different sized DNA chains with gel electrophoresis and analyze via fluorometer • “Read” DNA sequence from shortest to longest fragment • Exceedingly cheap nowadays due to automation (~$8 per sample) • Primers should be 30 -50 bp upstream of region of interest. • Really small DNA fragments are hard to resolve so you can’t read them well. Image courtesy of Online Biology Notes 17
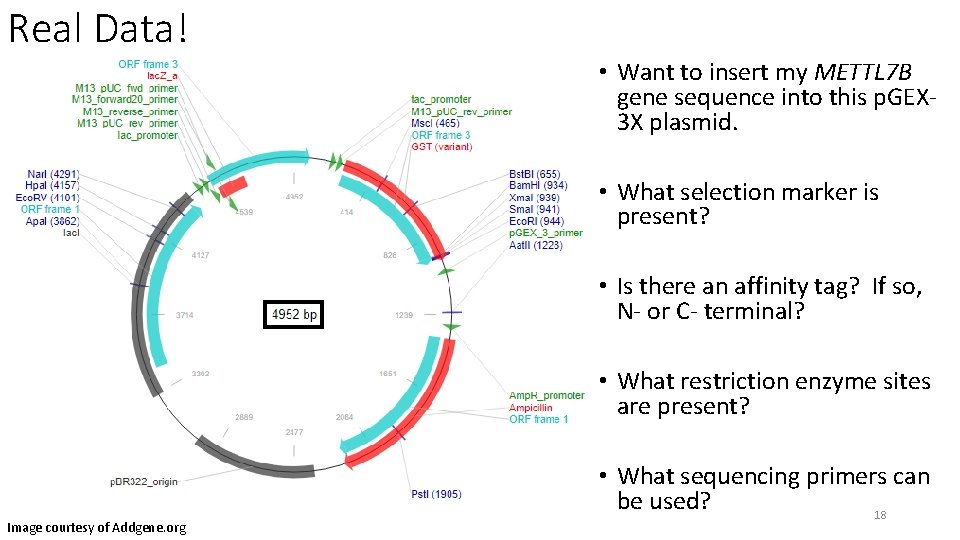
Real Data! • Want to insert my METTL 7 B gene sequence into this p. GEX 3 X plasmid. • What selection marker is present? • Is there an affinity tag? If so, N- or C- terminal? • What restriction enzyme sites are present? Image courtesy of Addgene. org • What sequencing primers can be used? 18
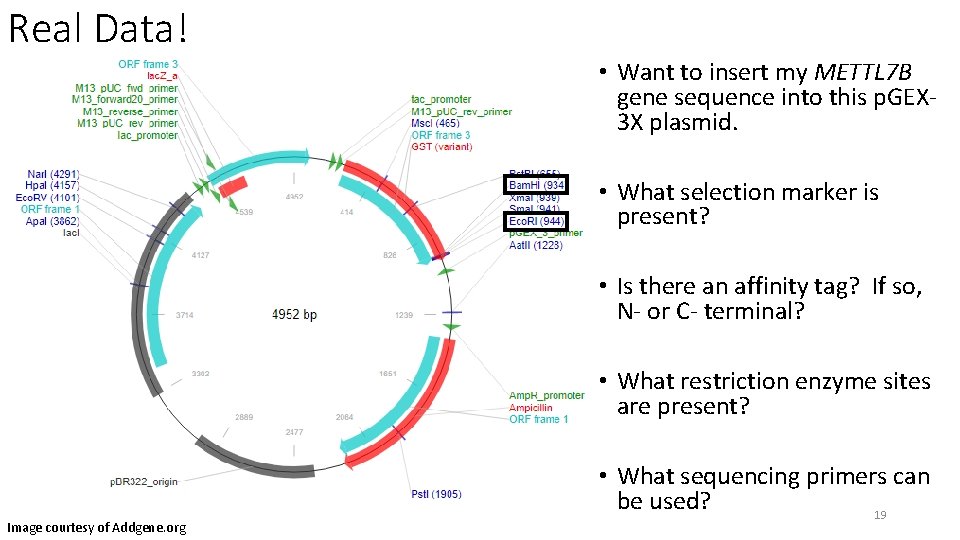
Real Data! • Want to insert my METTL 7 B gene sequence into this p. GEX 3 X plasmid. • What selection marker is present? • Is there an affinity tag? If so, N- or C- terminal? • What restriction enzyme sites are present? Image courtesy of Addgene. org • What sequencing primers can be used? 19
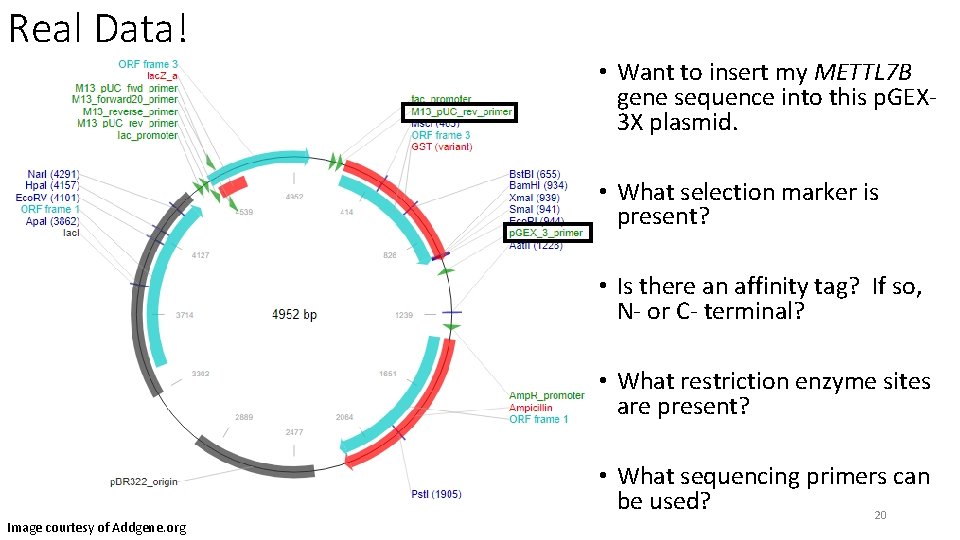
Real Data! • Want to insert my METTL 7 B gene sequence into this p. GEX 3 X plasmid. • What selection marker is present? • Is there an affinity tag? If so, N- or C- terminal? • What restriction enzyme sites are present? Image courtesy of Addgene. org • What sequencing primers can be used? 20

METTL 7 B Plasmid Gene Sequence ATGGATATTTTAGTGCCATTGTTACAGCTTCTTGTGTTGCTGCTGACTCTTCACTTGATGGCGTTGT TAGGTTGTTGGCAACCTTTGTGTAAAAGCTACTTCCCCTACTTAATGGCTGTGTTAACCCCAAAATCGAACC GGAAGATGGAATCCAAAAAGAGTTATTTTCTCAGATTAAGGGGCTTACAGGTGCGTCTGGTAAGG TAGCACTGTTGGAGTTAGGATGTGGGACCGGAGCTAACTTCCAATTTTATCCTCCGGGCTGCCGGGTTAC GTGCCTTGACCCTAATCCGCATTTTGAAAAATTCTTGACTAAATCTATGGCAGAAAATCGTCACTTACAGTA CGAGAGATTCGTTGTGGCTCCGGGTGAAGATATGAGACAACTGGCGGATCAATGGATGTTGTTGT ATGCACCCTTGTGTTATGCTCCGTTCAGTCCCCACGGAAGGTTCTGCAAGAGGTACGGCGGGTTCTGCGC CCAGGGGGAGTGTTATTTTTCTGGGAGCACGTAGCCGAACCATACGGGTCATGGGCATTCATGTGGCAGC AGGTCTTCGAACCCACTTGGAAACACATCGGAGACGGCTGCTGTTTAACGCGGGAAACGTGGAAGGACT TGGAAAATGCACAGTTTAGCGAAATTCAGATGGAAAGACAACCCCGCTGAAATGGTTGCCTGTAGG CCCGCATATTATGGGCAAAGCCGTCAAGACGCGT • Restriction sites available are the upstream Bam. HI (934) and downstream Eco. RI (944) sites • Bam. HI recognition sequence: GGATCC • Eco. RI recognition sequence: GAATTC • Will use overhang PCR to introduce restriction sites as well as extra stop codons after the gene sequence 21
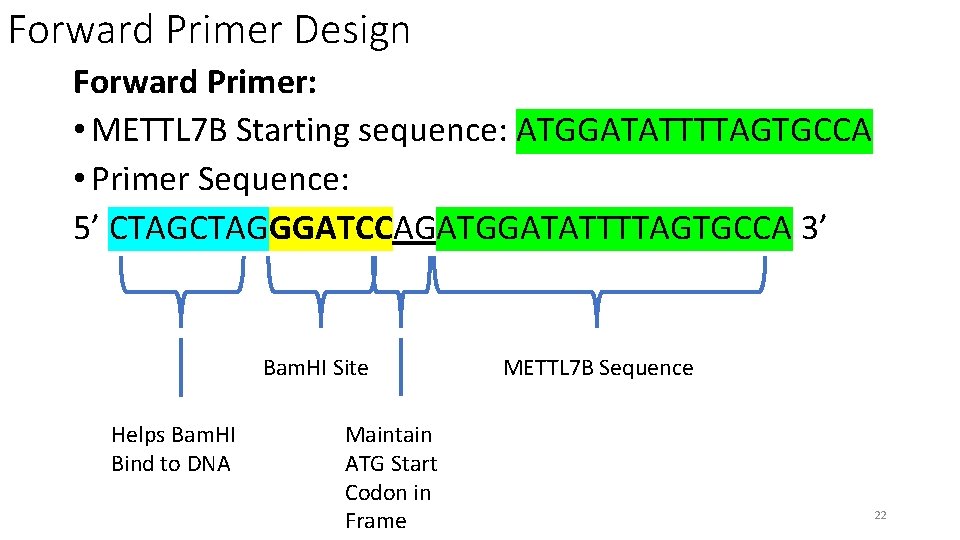
Forward Primer Design Forward Primer: • METTL 7 B Starting sequence: ATGGATATTTTAGTGCCA • Primer Sequence: 5’ CTAGGGATCCAGATGGATATTTTAGTGCCA 3’ Bam. HI Site Helps Bam. HI Bind to DNA Maintain ATG Start Codon in Frame METTL 7 B Sequence 22
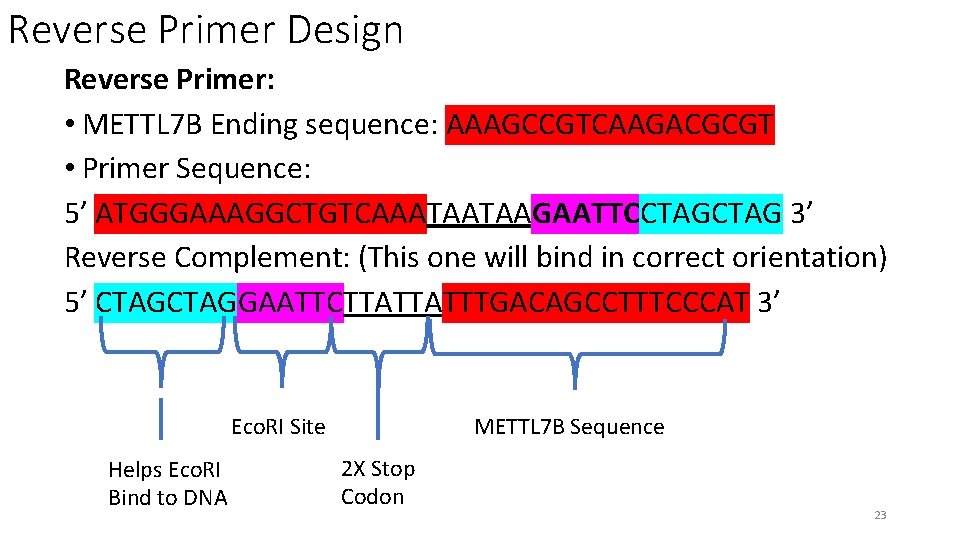
Reverse Primer Design Reverse Primer: • METTL 7 B Ending sequence: AAAGCCGTCAAGACGCGT • Primer Sequence: 5’ ATGGGAAAGGCTGTCAAATAATAAGAATTCCTAG 3’ Reverse Complement: (This one will bind in correct orientation) 5’ CTAGGAATTCTTATTATTTGACAGCCTTTCCCAT 3’ Eco. RI Site Helps Eco. RI Bind to DNA METTL 7 B Sequence 2 X Stop Codon 23
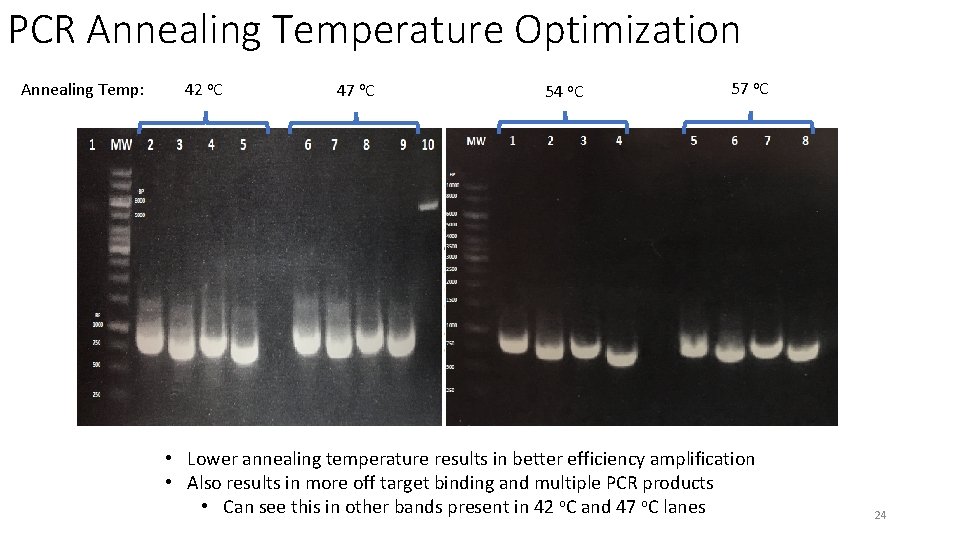
PCR Annealing Temperature Optimization Annealing Temp: 42 o. C 47 o. C 54 o. C 57 o. C • Lower annealing temperature results in better efficiency amplification • Also results in more off target binding and multiple PCR products • Can see this in other bands present in 42 o. C and 47 o. C lanes 24
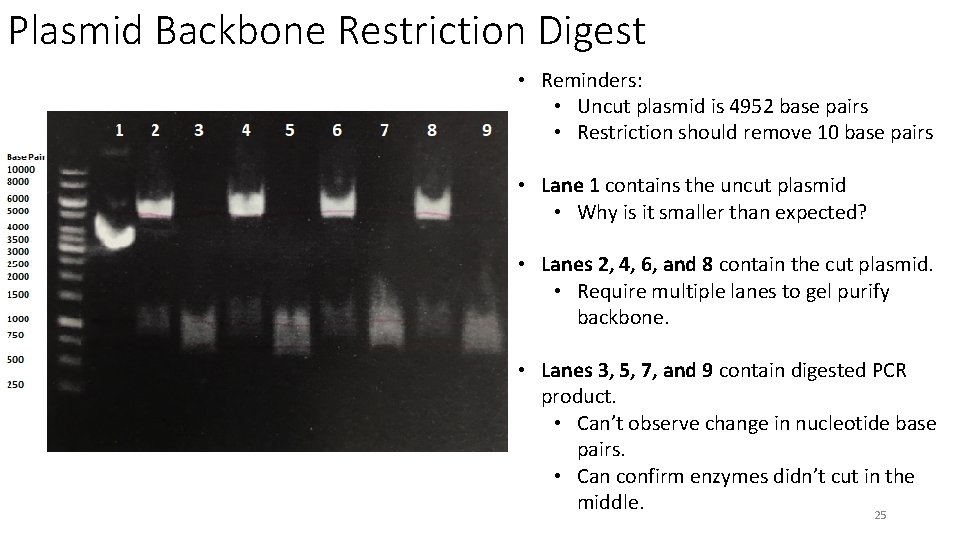
Plasmid Backbone Restriction Digest • Reminders: • Uncut plasmid is 4952 base pairs • Restriction should remove 10 base pairs • Lane 1 contains the uncut plasmid • Why is it smaller than expected? • Lanes 2, 4, 6, and 8 contain the cut plasmid. • Require multiple lanes to gel purify backbone. • Lanes 3, 5, 7, and 9 contain digested PCR product. • Can’t observe change in nucleotide base pairs. • Can confirm enzymes didn’t cut in the middle. 25
Finch TV • Go to example 26

Clustal Omega • Multiple sequence alignment • Useful to examine sequencing data for presence of SNPs or frameshift mutations. • Not all PCR is perfect • Risk of mutation increases with length of PCR amplification • Use of high fidelity enzymes mitigates the risk • Might be better/easier tools in other sequencing viewer software. This is just how I do it. 27

Conclusions • Lots of online tools and guides for workflow and design aspects for cloning • Kits for every possible cloning thing you want to do • Not all companies make the same quality kits • Personal favorite is Qiagen but New England Biolabs kits have worked before • UW Biochem Stores in J-wing basement sells all the kits you need • Plan out all the design before synthesizing primers • Really annoying to have to re-submit a synthesis order because you messed up • Ask for help! Multiple people looking at primer design or digestion sites might find problems that were overlooked. 28
- Slides: 28