Cloning and expression of HEAT repeat containing subunits

Cloning and expression of HEAT repeat containing subunits of condensin complexes in C. elegans Christina Borchers & Calvin Yoon ERCAN Lab NYU, 2012

Background Information • The large protein complexes, known as condensins, play a central role in chromosome assembly and segregation in Eukaryotic cells. • Many Eukaryotic cells have 2 condensin complexes: Condensin I and Condensin II • Caenorhabditis elegans have both Condensin I and Condensin II, but also have a third complex closely related to condensin 1 that is involved in dosage compensation (DCC) 2

Background Information • C. elegans hermaphrodites (XX), partially repress both X chromosomes in order to balance the gene expression between hermaphrodites and males • The study is directed to pursue the hypothesis that the differential recruitment of the condensins to chromosomes is achieved because of the role of the histone modifications in the interaction. • These histone modifications act to mediate the affinity of the condensins to the chromosomes. • Because there are many different types of these condensins that bind to various regions on the chromosomes, they embody different functions in chromosome structure and transcription. 3

Goals • Long-term: to determine how the HEAT repeat containing condensin subunits interact with histone modifications. • Short-term (summer): clone, bacterially express and purify soluble recombinant HEAT repeat containing condensin subunits in C. elegans 4

Overall procedure Obtain RNA from C. elegans embryos Synthesize first-strand of c. DNA PCR amplify DNA Digest DNA to form sticky ends (or perform T 4 DNA Polymerase treatment) • Ligate products into bacteria plasmids • Undergo basic transformation • •

Overall procedure • Prepare miniprep cultures • Make minipreps of DNA using Qiagen set • Digest at Pst. I +Eco. RI) site to check for proper transformation • Sequence minipreps • Obtain recombinant protein and run protein gel to check for solubility • Purify protein using GST tag and run SDS-PAGE

Preparation of RNA samples Obtained RNA from frozen C. elegans embryo Used Trizol extraction Qiagen Rneasy kit Measured concentration of RNA using Nano Drop spectrophotometer • 5/31/11: 142 ng/µl 6/1/11: 0. 9 • •

Synthesis of first-strand c. DNA from RNA • Oligo(d. T) added to hybridize onto poly-A tail of m. RNA • Added d. NTPs & SS III (along with Mg. Cl 2, DTT, RNase. OUT) to RNA • Treat with RNase. H

PCR Step 1: Initiation- Sample is heated to 95°C, melting the Hydrogen bonds, and creating single-stranded DNA Step 2: Annealing- Sample is cooled to 55°C, primers case-pair with the template strands Step 3: Elongation- Sample heated to 72°C. The DNA polymerase elongates strands of DNA that are complementary to each template strand 9 9

PCR CAPG-1 275 -377, 451 -550, 704 -822 CAPG-2 109 -377 DPY-28 200 -550, 664 -1455 HCP-6 500 -800, 801 -1357, 621 -1357 KLE-2 & AIR-2 10
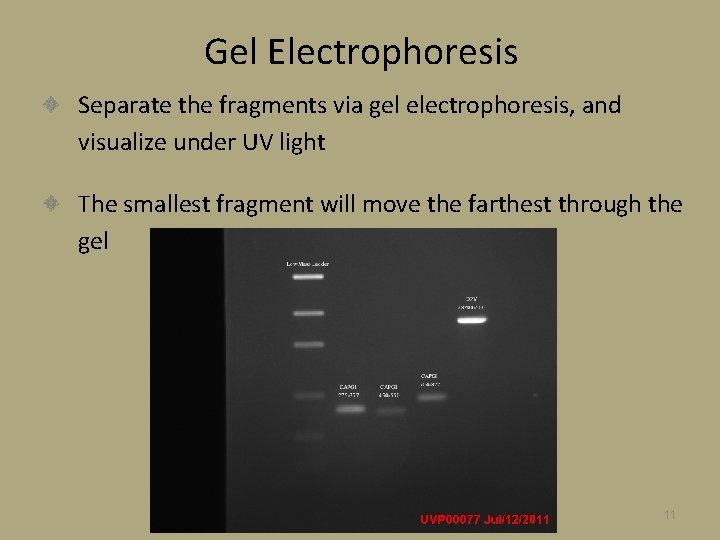
Gel Electrophoresis Separate the fragments via gel electrophoresis, and visualize under UV light The smallest fragment will move the farthest through the gel 11

PCR Column Purification The product is purified before sequenced because components like competing enzymes or buffer components can cause issues The kit is used to purify PCR fragments from an agarose gel Works well for DNA in the 70 bp-10 Kbp range. Fragments larger should be extracted w/ the Gel Extraction Kit 12
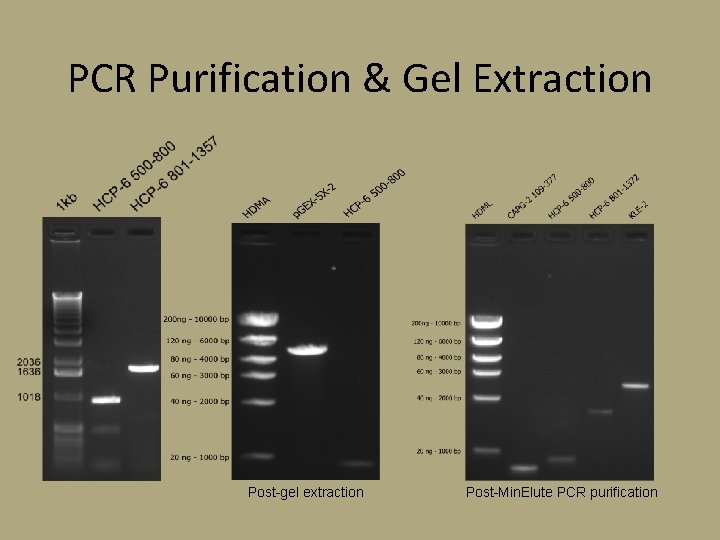
PCR Purification & Gel Extraction Post-gel extraction Post-Min. Elute PCR purification

Gel Extraction • Weigh samples • Gel slices dissolve in a buffer containing a p. H indicator for optimal DNA binding • add to spin columns, nucleic acids absorb the silica membrane due to the buffer • impurities are washed away and pure DNA remains 14
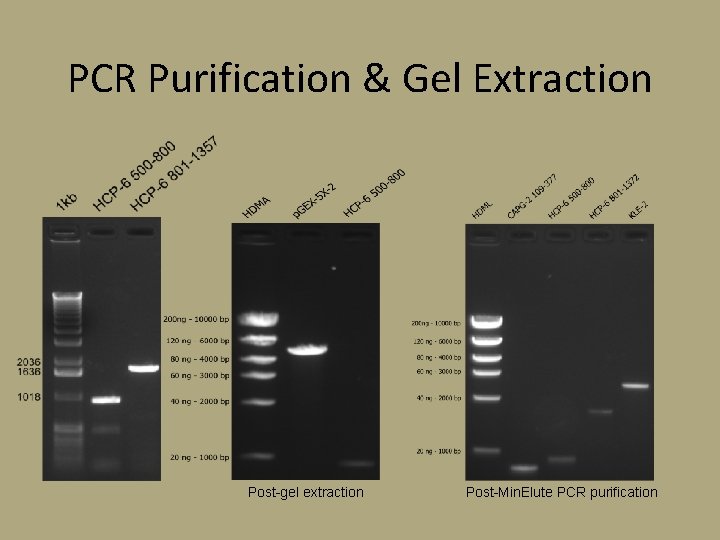
PCR Purification & Gel Extraction Post-gel extraction Post-Min. Elute PCR purification

Digestion of DNA To form the free ends of the DNA molecule that were complementary, known as sticky Ends. They are able to pair with other DNA molecules w/ similar sequences. The restriction enzyme sites in plasmid DNA allows our desired sequence w/ the complementary sticky ends to be inserted into the plasmid. The plasmid then can be reintroduced into the bacteria cell Bam. HI and Not. I restrictive enzymes used 10 X Buffer 3, 1000 X BSA, PCR products, REs 16

17
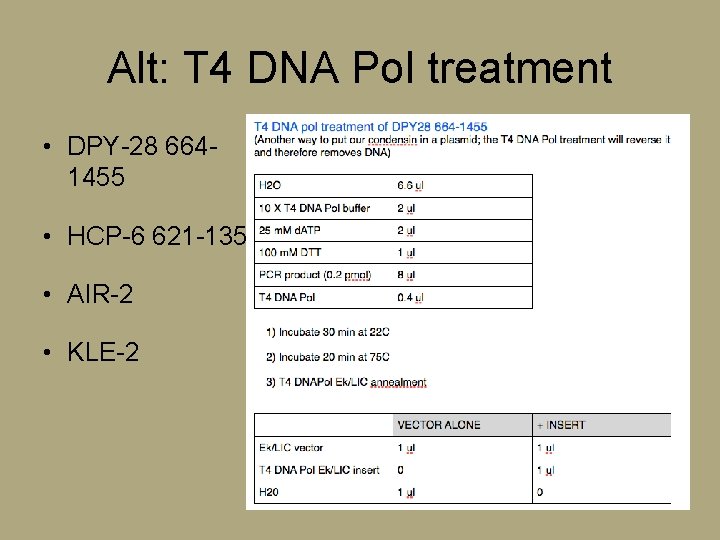
Alt: T 4 DNA Pol treatment • DPY-28 6641455 • HCP-6 621 -1357 • AIR-2 • KLE-2
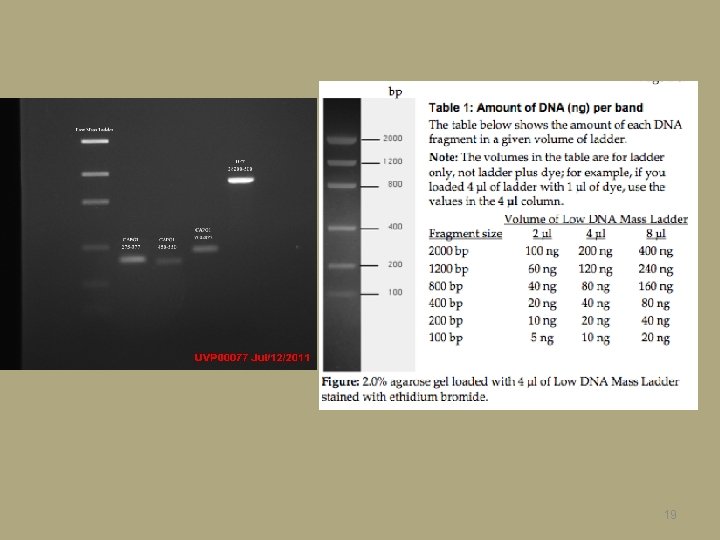
19

Ligation • To ligate a DNA fragment (the insert) into a plasmid (p. GEX-5 X-2) they must have compatible ends after digestion Alt: T 4 DNAPol Ek/LIC Annealment Vector alone Ek/LIC vector 1 T 4 DNA Pol Ek/LIC insert 0 H 2 O 2 +insert 1 2 0

Restriction Enzyme Reaction 21

Basic Transformation • Introduce a plasmid into a bacteria that will amplify the plasmid to make large quantities of it. • Based on Plasmid function of transferring genetic information necessary to the survival of bacteria • The Plasmid: used to amplify a gene of interest. Used as a vector if containing resistance to an antibiotic (ampicillin) • Gene of interest is inserted into the vector plasmid and then put into E. Coli, sensitive to ampicillin. • 5 µl of 5 -10 ng of DNA to One Shot cells • Started off by plating 10 -1, 10 -2 & 10 -3 dilutions with BL-21 • Redid transformations with TOP 10 with undiluted, 10 -1 & 10 -2 dilutions 22

Preparation of miniprep cultures • 3 ml LB with antibiotics (50µl amp in 50 ml LB) 1000 • Used pipette to obtain culture (10 per protein) • Obtained DNA using Qiagen kit
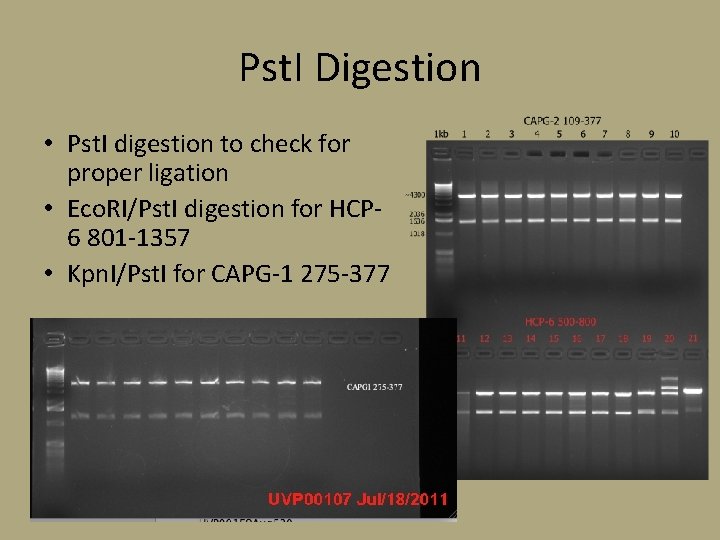
Pst. I Digestion • Pst. I digestion to check for proper ligation • Eco. RI/Pst. I digestion for HCP 6 801 -1357 • Kpn. I/Pst. I for CAPG-1 275 -377
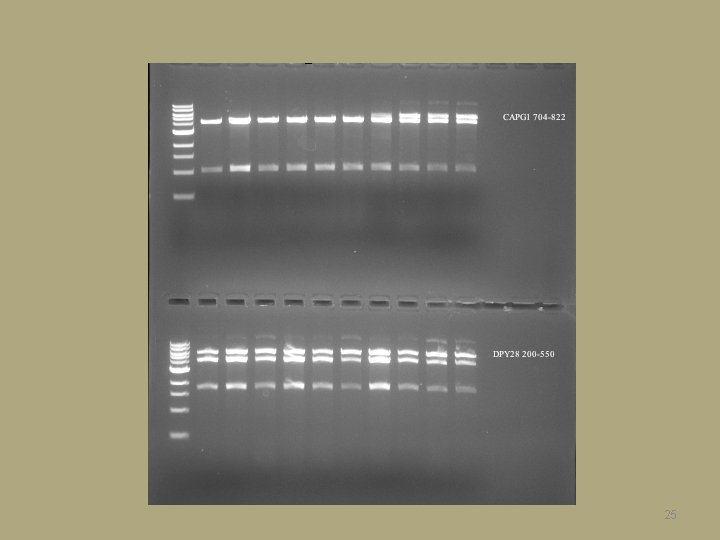
25

26
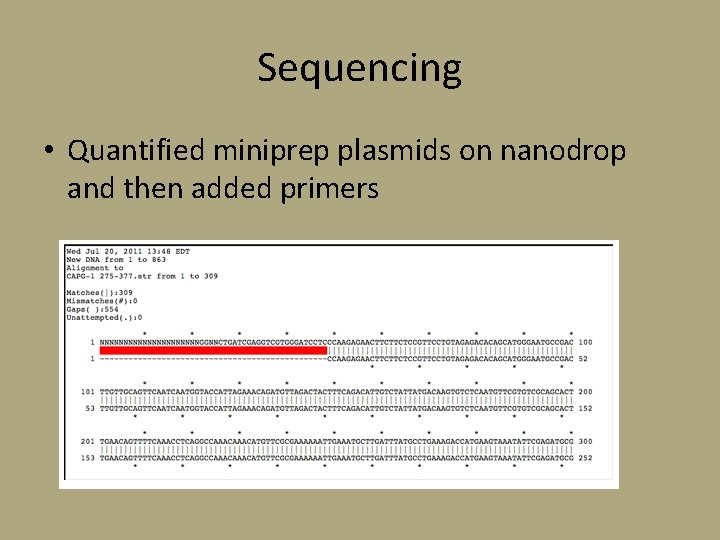
Sequencing • Quantified miniprep plasmids on nanodrop and then added primers

Optical Density • Measured OD of culture tubes (10 µl sample in 3 ml LB/amp) • Calculated for 0. 1 OD in 5 ml LB/amp & shook • At 0. 4 -0. 6 OD, added 5 µl of 1 M IPTG (initially 1 m. M) to induce expression • Froze using liquid nitrogen
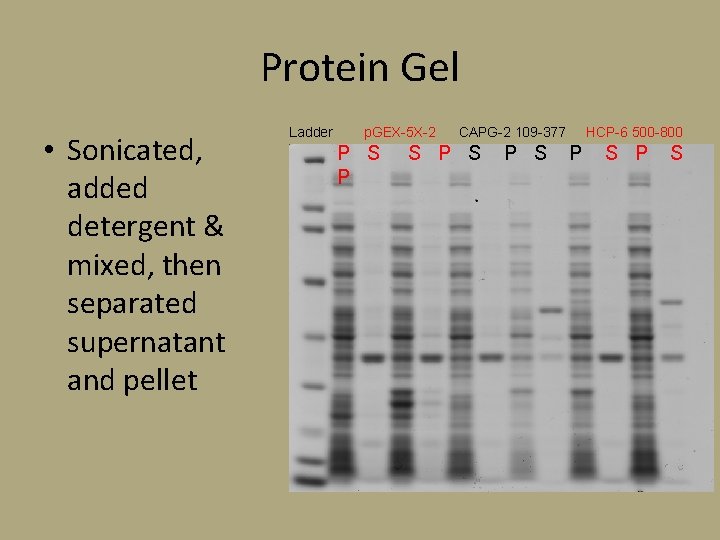
Protein Gel • Sonicated, added detergent & mixed, then separated supernatant and pellet Ladder p. GEX-5 X-2 P S P CAPG-2 109 -377 S P S HCP-6 500 -800 P S

GSH Pulldown
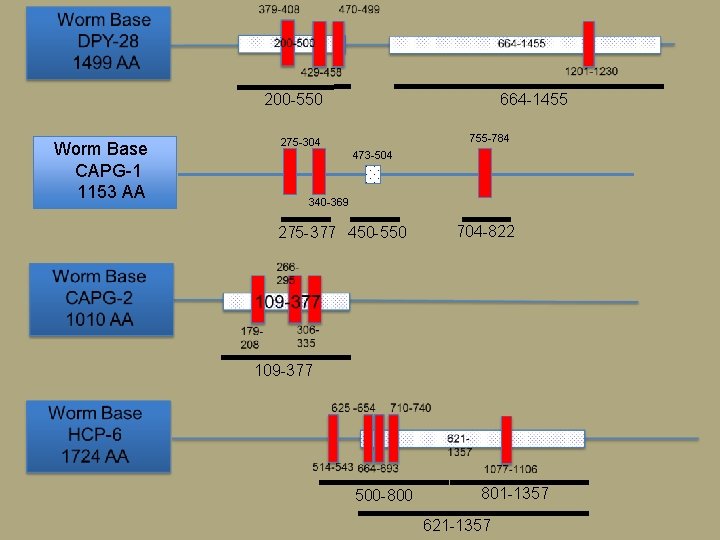
200 -550 Worm Base CAPG-1 1153 AA 664 -1455 755 -784 275 -304 473 -504 340 -369 275 -377 450 -550 704 -822 109 -377 500 -800 801 -1357 621 -1357
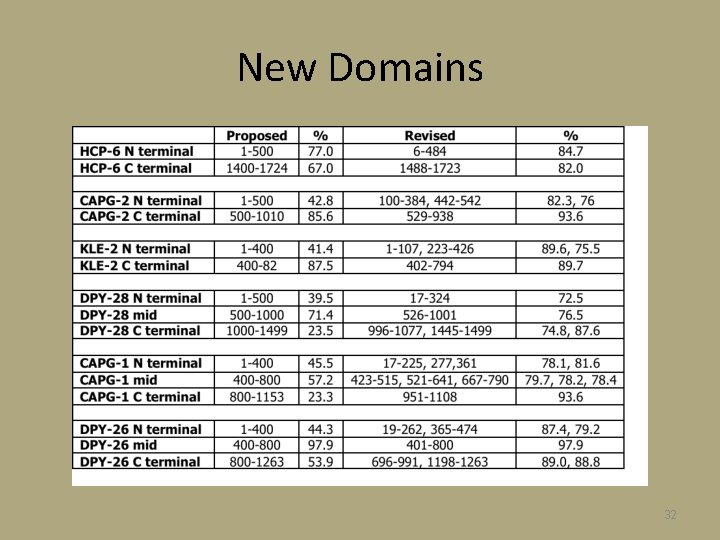
New Domains 32
- Slides: 32