Clinical Drug Resistance Linked to Interconvertible Phenotypic and

Clinical Drug Resistance Linked to Inter-convertible Phenotypic and Functional States of Tumour. Propagating cells in Multiple Myeloma Dr Aristeidis Chaidos MD Ph. D Centre for Haematology, Imperial College London Hammersmith Hospital, Imperial Healthcare NHS Trust
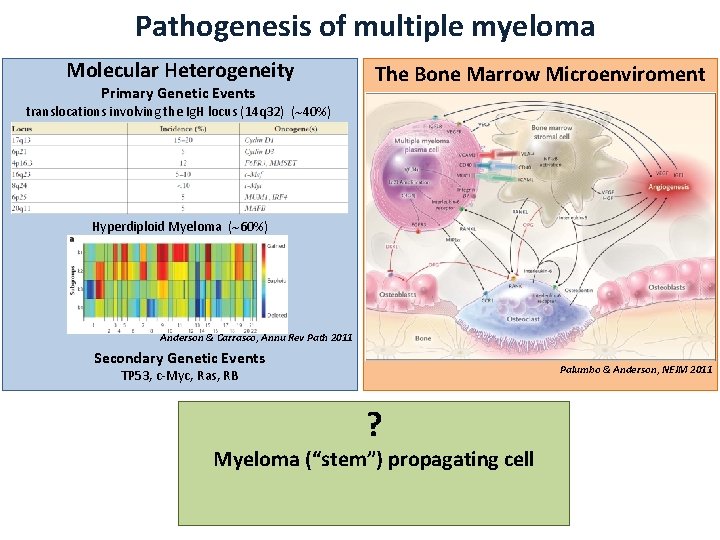
Pathogenesis of multiple myeloma Molecular Heterogeneity Primary Genetic Events The Bone Marrow Microenviroment translocations involving the Ig. H locus (14 q 32) ( 40%) Hyperdiploid Myeloma ( 60%) Anderson & Carrasco, Annu Rev Path 2011 Secondary Genetic Events Palumbo & Anderson, NEJM 2011 TP 53, c-Myc, Ras, RB ? Myeloma (“stem”) propagating cell
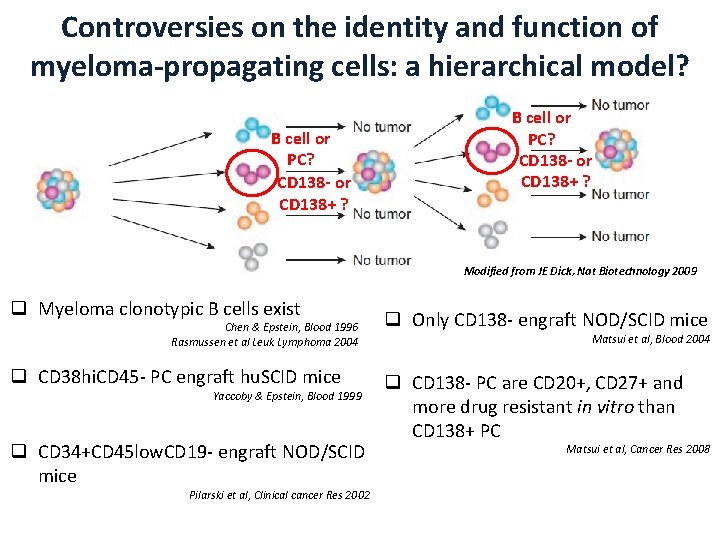
Controversies on the identity and function of myeloma-propagating cells: a hierarchical model? B cell or PC? CD 138 - or CD 138+ ? Modified from JE Dick, Nat Biotechnology 2009 q Myeloma clonotypic B cells exist Chen & Epstein, Blood 1996 Rasmussen et al Leuk Lymphoma 2004 q CD 38 hi. CD 45 - PC engraft hu. SCID mice Yaccoby & Epstein, Blood 1999 q CD 34+CD 45 low. CD 19 - engraft NOD/SCID mice Pilarski et al, Clinical cancer Res 2002 q Only CD 138 - engraft NOD/SCID mice Matsui et al, Blood 2004 q CD 138 - PC are CD 20+, CD 27+ and more drug resistant in vitro than CD 138+ PC Matsui et al, Cancer Res 2008
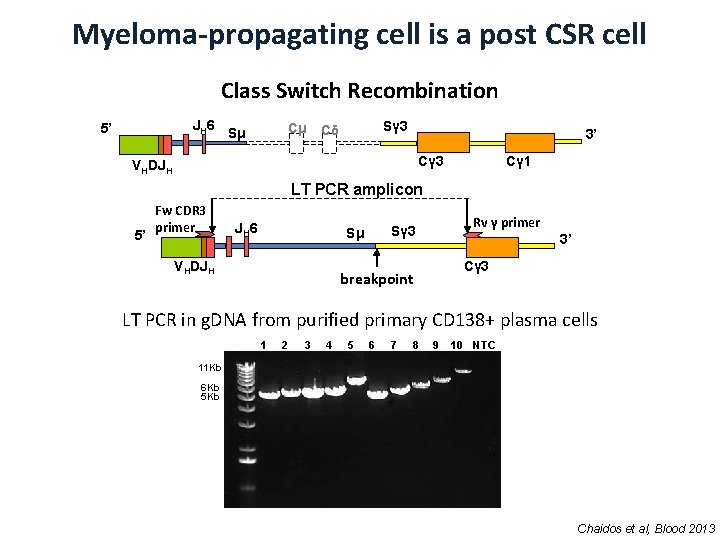
Myeloma-propagating cell is a post CSR cell Class Switch Recombination JH 6 5’ Cμ Sμ Sγ 3 Cδ 3’ Cγ 3 VHDJH Cγ 1 LT PCR amplicon 5’ Fw CDR 3 primer JH 6 VHDJH Rv γ primer Sγ 3 Sμ 3’ Cγ 3 breakpoint LT PCR in g. DNA from purified primary CD 138+ plasma cells 1 2 3 4 5 6 7 8 9 10 NTC 11 Kb 6 Kb 5 Kb Chaidos et al, Blood 2013
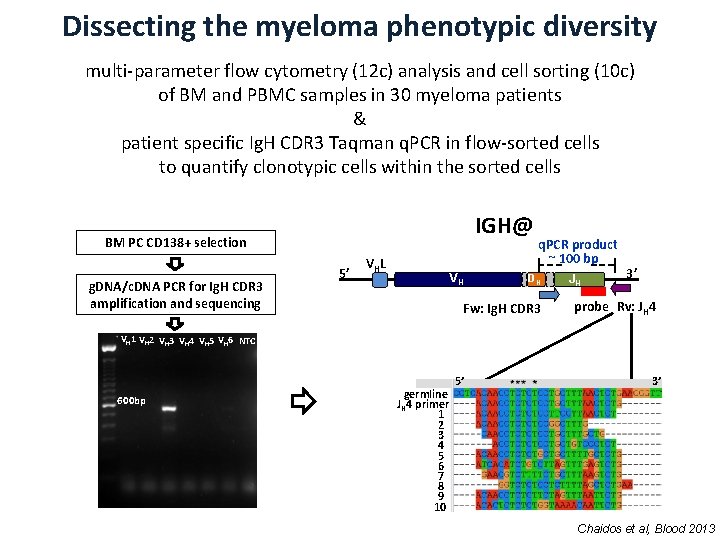
Dissecting the myeloma phenotypic diversity multi-parameter flow cytometry (12 c) analysis and cell sorting (10 c) of BM and PBMC samples in 30 myeloma patients & patient specific Ig. H CDR 3 Taqman q. PCR in flow-sorted cells to quantify clonotypic cells within the sorted cells IGH@ BM PC CD 138+ selection g. DNA/c. DNA PCR for Ig. H CDR 3 amplification and sequencing 5’ VH L VH VH q. PCR product ~ 100 bp DH Fw: Ig. H CDR 3 JH 3’ probe Rv: JH 4 VH 1 VH 2 VH 3 VH 4 VH 5 VH 6 NTC 600 bp germline JH 4 primer 1 2 3 4 5 6 7 8 9 10 5’ 3’ Chaidos et al, Blood 2013
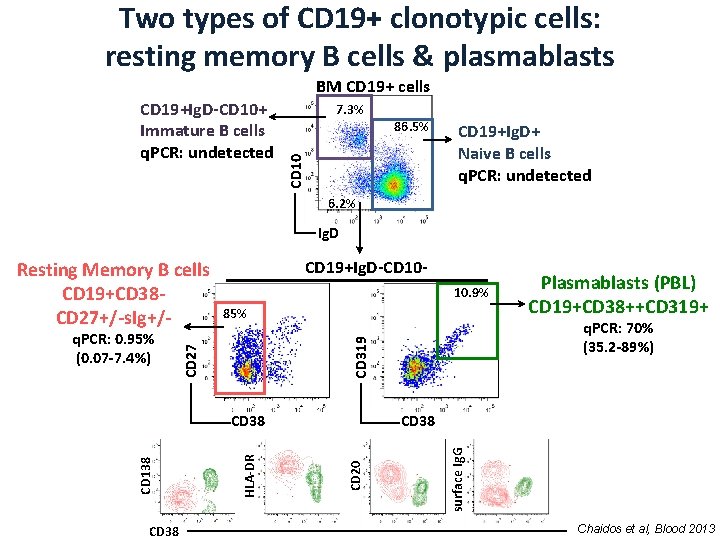
Two types of CD 19+ clonotypic cells: resting memory B cells & plasmablasts BM CD 19+ cells 7. 3% 86. 5% CD 10 CD 19+Ig. D-CD 10+ Immature B cells q. PCR: undetected CD 19+Ig. D+ Naive B cells q. PCR: undetected 6. 2% Ig. D 10. 9% 85% CD 20 HLA-DR CD 138 surface Ig. G CD 38 Plasmablasts (PBL) CD 19+CD 38++CD 319+ q. PCR: 70% (35. 2 -89%) CD 319 q. PCR: 0. 95% (0. 07 -7. 4%) CD 19+Ig. D-CD 10 - CD 27 Resting Memory B cells CD 19+CD 38 CD 27+/-s. Ig+/- Chaidos et al, Blood 2013
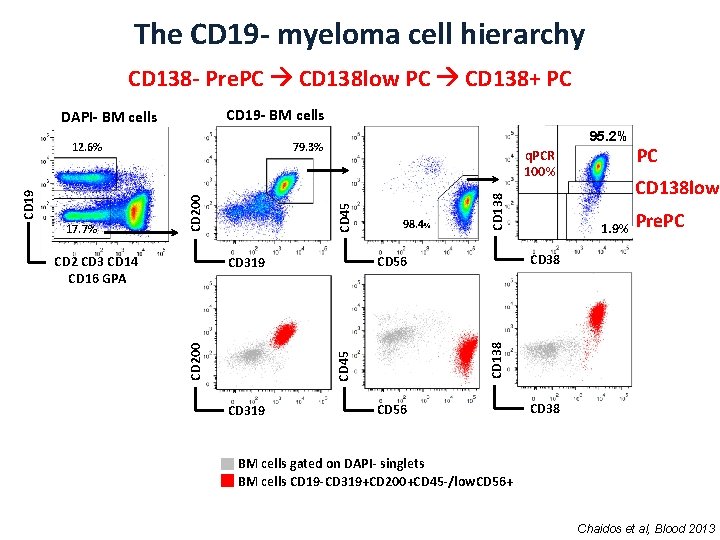
The CD 19 - myeloma cell hierarchy CD 138 - Pre. PC CD 138 low PC CD 138+ PC CD 19 - BM cells DAPI- BM cells CD 138 CD 45 CD 319 CD 138 low 1. 9% Pre. PC CD 38 CD 56 CD 319 CD 200 98. 4% PC CD 138 CD 2 CD 3 CD 14 CD 16 GPA q. PCR 100% CD 45 17. 7% 95. 2% 79. 3% CD 200 CD 19 12. 6% CD 56 CD 38 BM cells gated on DAPI- singlets BM cells CD 19 -CD 319+CD 200+CD 45 -/low. CD 56+ Chaidos et al, Blood 2013
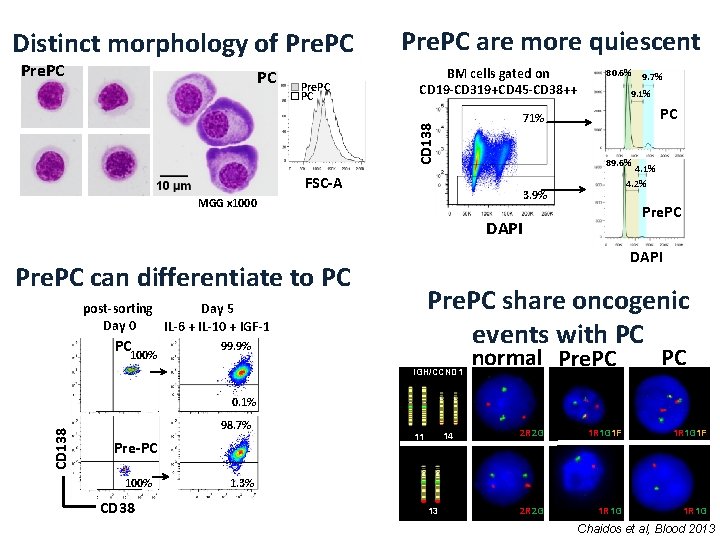
Distinct morphology of Pre. PC PC BM cells gated on CD 19 -CD 319+CD 45 -CD 38++ 80. 6% 9. 1% PC 89. 6% FSC-A 4. 1% 4. 2% 3. 9% MGG x 1000 Pre. PC DAPI Pre. PC can differentiate to PC Pre. PC share oncogenic events with PC post-sorting Day 5 Day 0 IL-6 + IL-10 + IGF-1 PC 100% 9. 7% 71% CD 138 PC Pre. PC are more quiescent 99. 9% IGH/CCND 1 normal Pre. PC PC CD 138 0. 1% 98. 7% Pre-PC 100% CD 38 14 11 2 R 2 G 1 R 1 G 1 F 2 R 2 G 1 R 1 G 1. 3% 13 Chaidos et al, Blood 2013
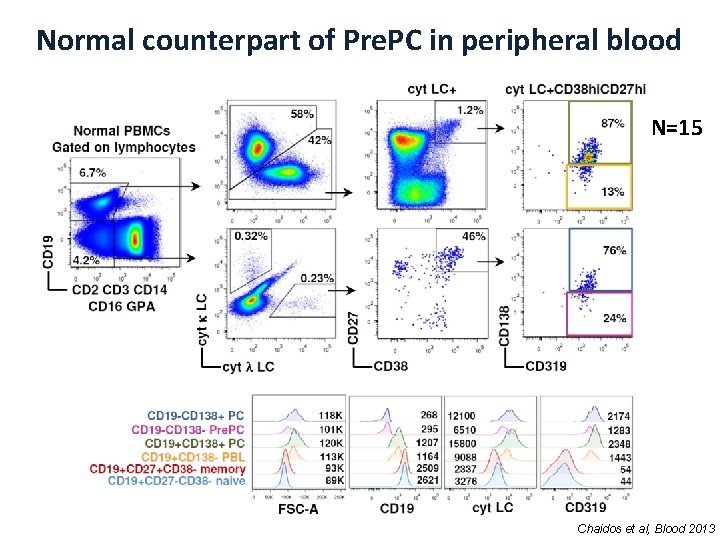
Normal counterpart of Pre. PC in peripheral blood N=15 Chaidos et al, Blood 2013
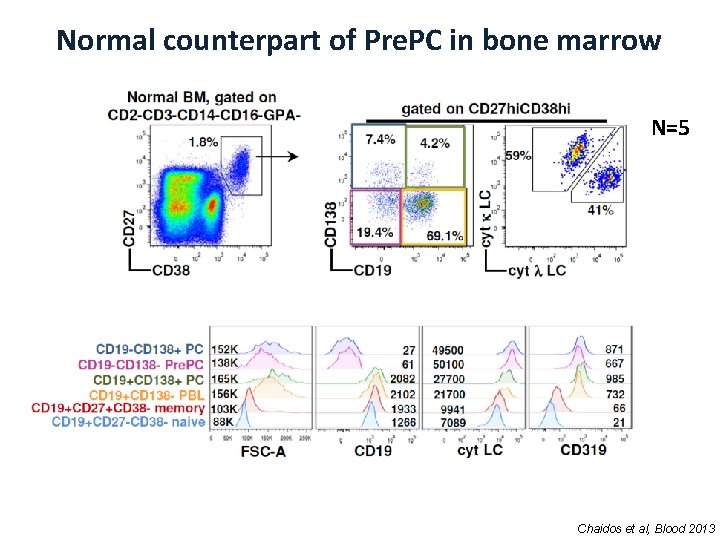
Normal counterpart of Pre. PC in bone marrow N=5 Chaidos et al, Blood 2013
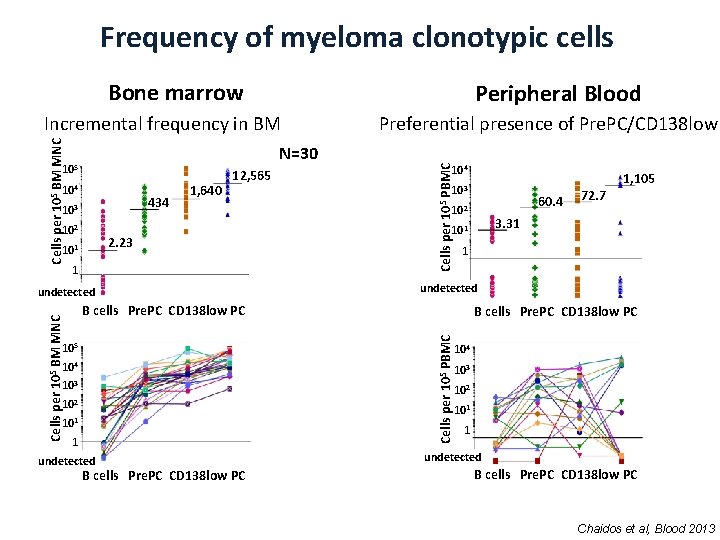
Frequency of myeloma clonotypic cells Bone marrow Peripheral Blood N=30 105 104 434 103 102 1, 640 12, 565 2. 23 101 1 104 103 102 101 1 undetected B cells Pre. PC CD 138 low PC 60. 4 102 72. 7 3. 31 101 1 undetected B cells Pre. PC CD 138 low PC 105 1, 105 103 B cells Pre. PC CD 138 low PC Cells per 105 PBMC Cells per 105 BM MNC undetected Preferential presence of Pre. PC/CD 138 low Cells per 105 PBMC Cells per 105 BM MNC Incremental frequency in BM 104 103 102 101 1 undetected B cells Pre. PC CD 138 low PC Chaidos et al, Blood 2013
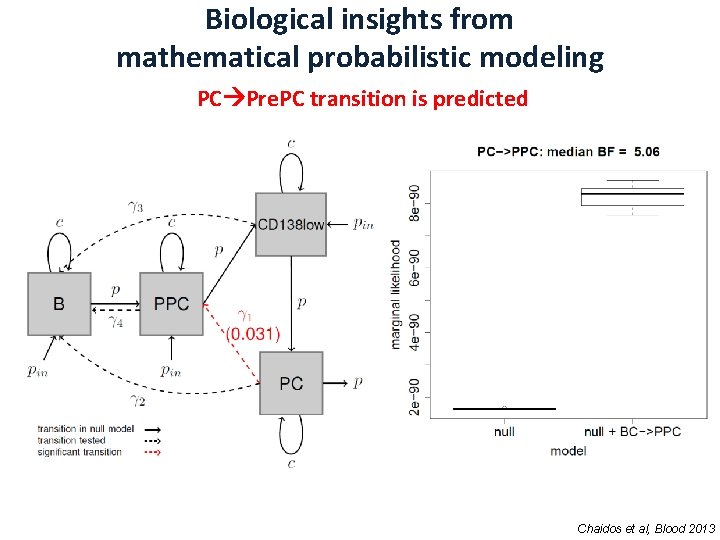
Biological insights from mathematical probabilistic modeling PC Pre. PC transition is predicted Chaidos et al, Blood 2013
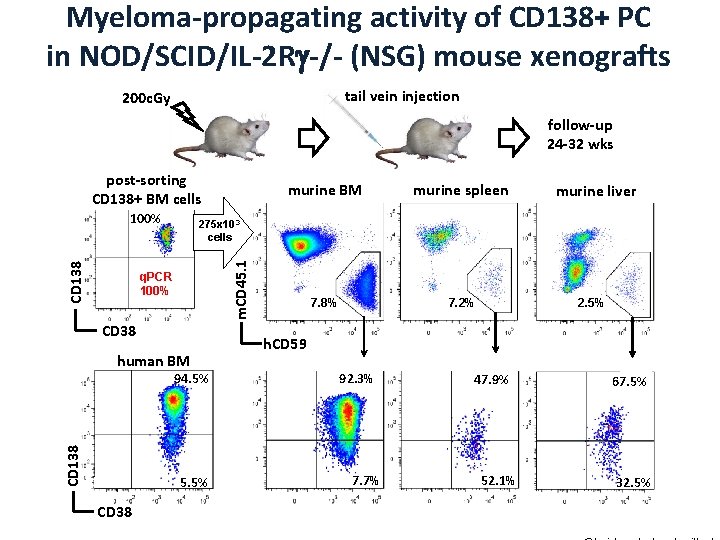
Myeloma-propagating activity of CD 138+ PC in NOD/SCID/IL-2 R -/- (NSG) mouse xenografts tail vein injection 200 c. Gy follow-up 24 -32 wks post-sorting CD 138+ BM cells CD 38 human BM 94. 5% CD 138 7. 8% 7. 2% murine liver 275 x 103 cells q. PCR 100% 5. 5% CD 38 murine spleen m. CD 45. 1 CD 138 100% murine BM 2. 5% h. CD 59 92. 3% 7. 7% 47. 9% 52. 1% 67. 5% 32. 5%
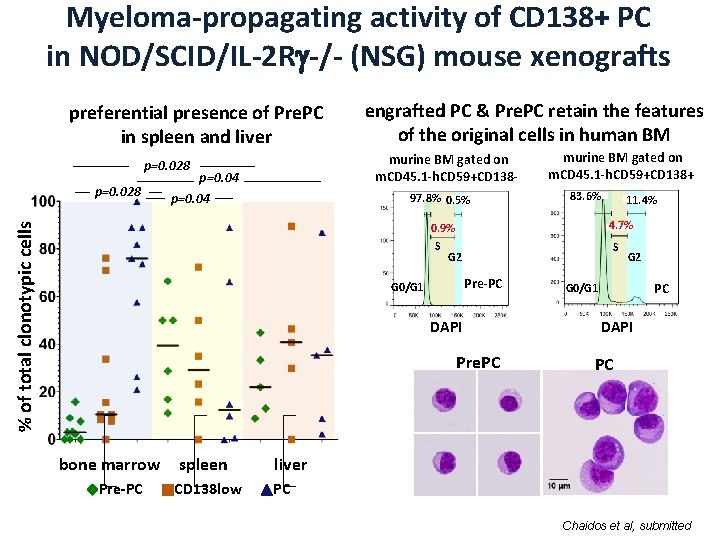
Myeloma-propagating activity of CD 138+ PC in NOD/SCID/IL-2 R -/- (NSG) mouse xenografts preferential presence of Pre. PC in spleen and liver murine BM gated on m. CD 45. 1 -h. CD 59+CD 138 - p=0. 028 engrafted PC & Pre. PC retain the features of the original cells in human BM p=0. 04 97. 8% 0. 5% murine BM gated on m. CD 45. 1 -h. CD 59+CD 138+ 83. 6% 4. 7% % of total clonotypic cells 0. 9% S S G 2 Pre-PC G 0/G 1 DAPI Pre. PC bone marrow Pre-PC spleen liver CD 138 low PC 11. 4% G 2 PC G 0/G 1 DAPI PC Chaidos et al, submitted
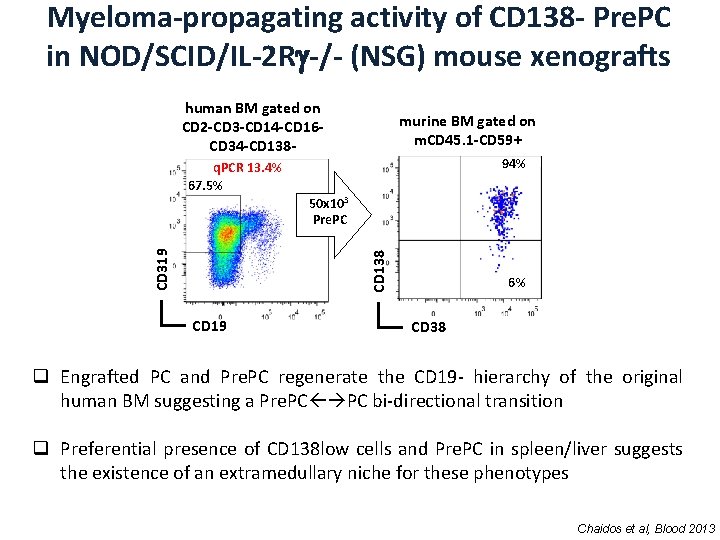
Myeloma-propagating activity of CD 138 - Pre. PC in NOD/SCID/IL-2 R -/- (NSG) mouse xenografts human BM gated on CD 2 -CD 3 -CD 14 -CD 16 CD 34 -CD 138 - murine BM gated on m. CD 45. 1 -CD 59+ 94% q. PCR 13. 4% 67. 5% CD 138 CD 319 50 x 103 Pre. PC CD 19 6% CD 38 q Engrafted PC and Pre. PC regenerate the CD 19 - hierarchy of the original human BM suggesting a Pre. PC PC bi-directional transition q Preferential presence of CD 138 low cells and Pre. PC in spleen/liver suggests the existence of an extramedullary niche for these phenotypes Chaidos et al, Blood 2013
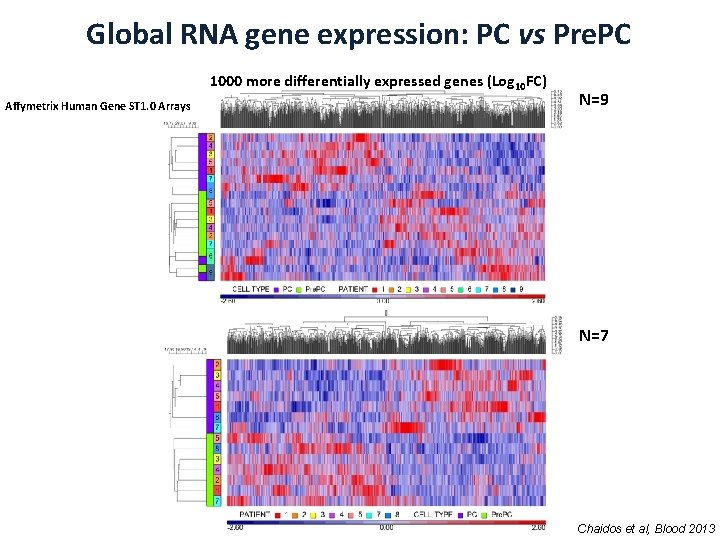
Global RNA gene expression: PC vs Pre. PC 1000 more differentially expressed genes (Log 10 FC) Affymetrix Human Gene ST 1. 0 Arrays N=9 N=7 Chaidos et al, Blood 2013

Global RNA gene expression: PC vs Pre. PC 1509 differentially expressed genes (p<0. 05) PCA p=4 x 10 -6 1059 genes Functional Annotation Clustering (DAVID) Top Annotation Cluster Enrichment Score: 12. 8 p SP_PIR_KEYWORDS chromatin regulator 3. 27 x 10 -15 GOTERM_BP_FAT chromatin modification 1. 69 x 10 -13 GOTERM_BP_FAT chromatin organization 2. 99 x 10 -13 GOTERM_BP_FAT chromosome organization 1. 97 x 10 -12 Chaidos et al, Blood 2013

Epigenetic basis of phenotypic transition Pre. PC are enriched in epigenetic regulators Chromatin remodeling q Several member of the SWI/SNF chromatin remodelers q Polycomb Repressive Complex 2 Polycomb repressive complex Histone methylation Histone acetylation q MLL complex q KDMs demethylases
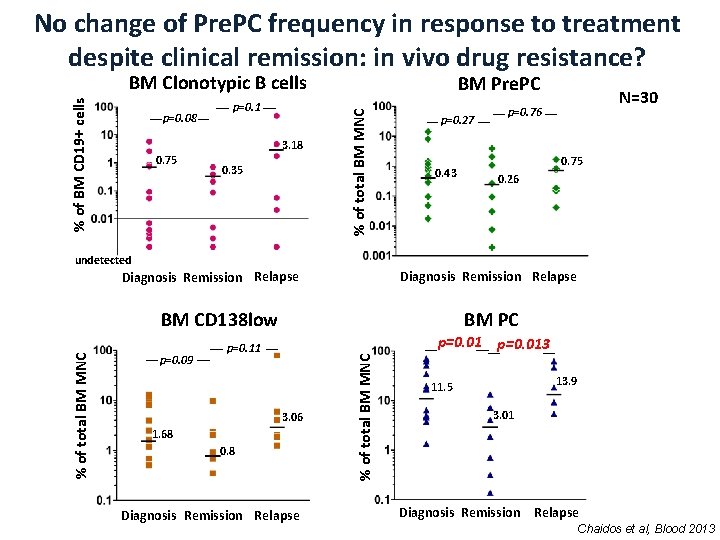
No change of Pre. PC frequency in response to treatment despite clinical remission: in vivo drug resistance? p=0. 08 p=0. 1 3. 18 0. 75 0. 35 BM Pre. PC % of total BM MNC % of BM CD 19+ cells BM Clonotypic B cells p=0. 27 0. 43 N=30 p=0. 76 0. 75 0. 26 undetected Diagnosis Remission Relapse BM PC p=0. 09 p=0. 013 p=0. 11 3. 06 1. 68 0. 8 Diagnosis Remission Relapse % of total BM MNC BM CD 138 low 13. 9 11. 5 3. 01 Diagnosis Remission Relapse Chaidos et al, Blood 2013
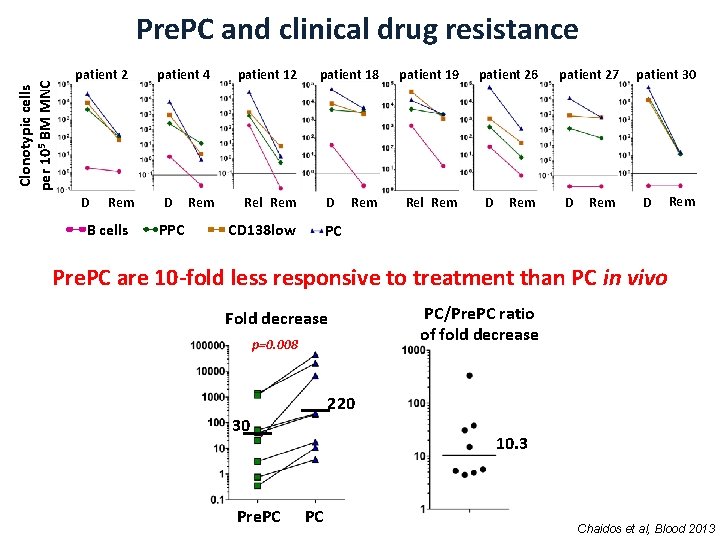
Clonotypic cells per 105 BM MNC Pre. PC and clinical drug resistance patient 2 D Rem B cells patient 4 D PPC Rem patient 12 patient 18 Rel Rem D CD 138 low Rem patient 19 Rel Rem patient 26 D Rem patient 27 D Rem patient 30 D Rem PC Pre. PC are 10 -fold less responsive to treatment than PC in vivo Fold decrease p=0. 008 PC/Pre. PC ratio of fold decrease 220 30 Pre. PC 10. 3 PC Chaidos et al, Blood 2013
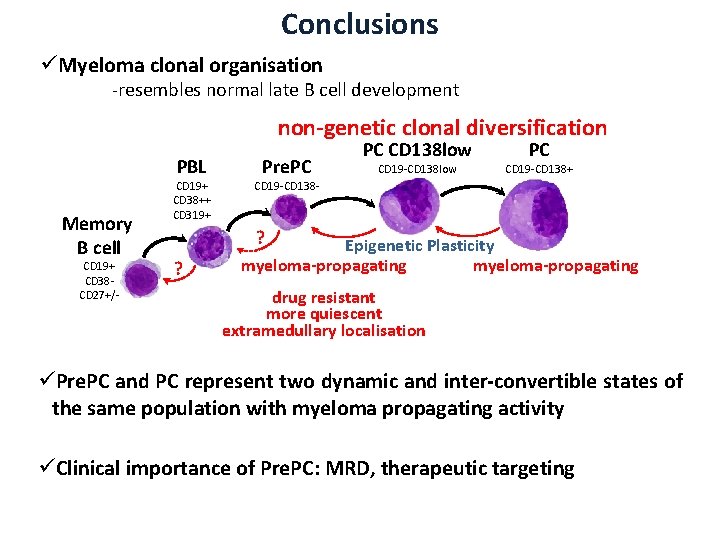
Conclusions üMyeloma clonal organisation -resembles normal late B cell development non-genetic clonal diversification PBL Memory B cell CD 19+ CD 38 CD 27+/- CD 19+ CD 38++ CD 319+ Pre. PC PC CD 138 low CD 19 -CD 138 low PC CD 19 -CD 138+ CD 19 -CD 138 - ? ? Epigenetic Plasticity myeloma-propagating drug resistant more quiescent extramedullary localisation üPre. PC and PC represent two dynamic and inter-convertible states of the same population with myeloma propagating activity üClinical importance of Pre. PC: MRD, therapeutic targeting

Acknowledgements Tassos Karadimitris Amin Rahemtulla Irene Roberts Mauritius Kleijnen Jane Apperley Saad Abdalla Helen Yarranton Evangelos Terpos Athens, Greece Maria Papaioannou Thessaloniki, Greece Evdoxia Hatjiharissi Thessaloniki, Greece Members of Prof Karadimitris & Prof Roberts group Department of Cell and Developmental Biology, University College London Helen Doolittle Gillian Cowan Georg Bohn Valentina Caputo Suhail Chaudry David O’Connor Deena Iskander Katerina Goudevenou Kalliopi Makarona Joana Simoes-Costa Andi Roy Chris Barnes Centre for Bioinformatics and Institute of Mathematical Sciences Heather Harrington Michael Stumpf Imperial Molecular Pathology Laboratory Letizia Foroni Gareth Gerrard Cytogenetics Alistair Reid Valeria Melo Philippa May Histopathology Kikkeri Naresh Weatheral Institute of Molecular Medicine, Oxford Paresh Vyas
- Slides: 22