ClingE coli Bacteria on target Harvard i GEM

Cling-E. coli : Bacteria on target Harvard i. GEM 2007 Ellenor Brown Stephanie Lo Alex Pickett Sammy Sambu Kevin Shee Perry Tsai Shaunak Vankudre George Xu

The motivation To develop a system for directing bacteria to a target of interest and effecting downstream activity • Bacterial targeting is necessary for spatiallyspecific activity in the body or in nature • Post-targeting activity and transmembrane signalling are the next step in engineering genetic circuits that interface extracellular and intracellular environments

The vision: Bacterial targeting via membrane display
The vision: Inter-cellular activation via Lux quorum-sensing

The vision: Intra-cellular activation via Fec signal transduction
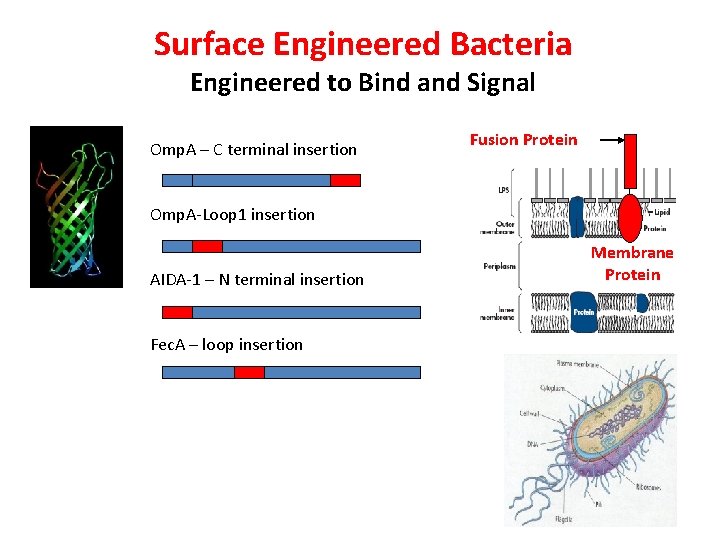
Surface Engineered Bacteria Engineered to Bind and Signal Omp. A – C terminal insertion Fusion Protein Omp. A-Loop 1 insertion AIDA-1 – N terminal insertion Fec. A – loop insertion Membrane Protein
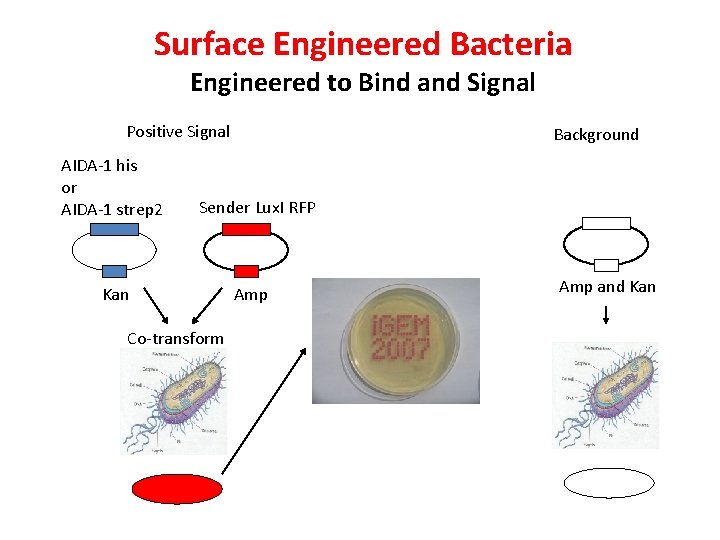
Surface Engineered Bacteria Engineered to Bind and Signal Positive Signal AIDA-1 his or AIDA-1 strep 2 Background Sender Lux. I RFP Kan Co-transform Amp and Kan
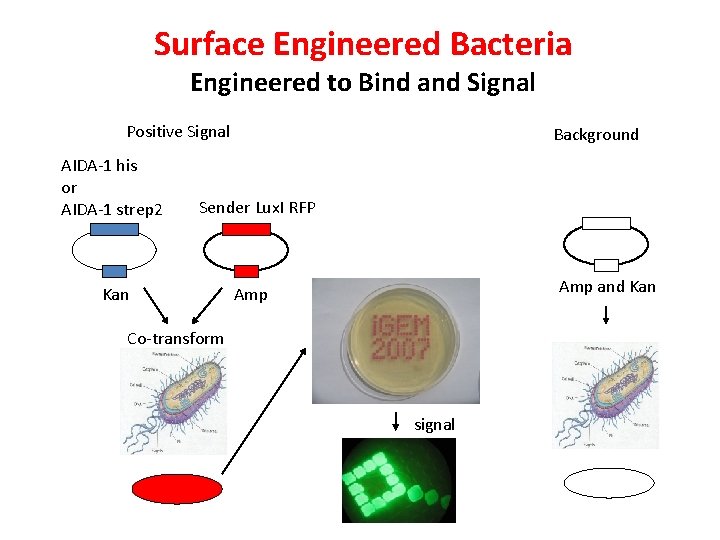
Surface Engineered Bacteria Engineered to Bind and Signal Positive Signal AIDA-1 his or AIDA-1 strep 2 Background Sender Lux. I RFP Kan Amp and Kan Amp Co-transform signal
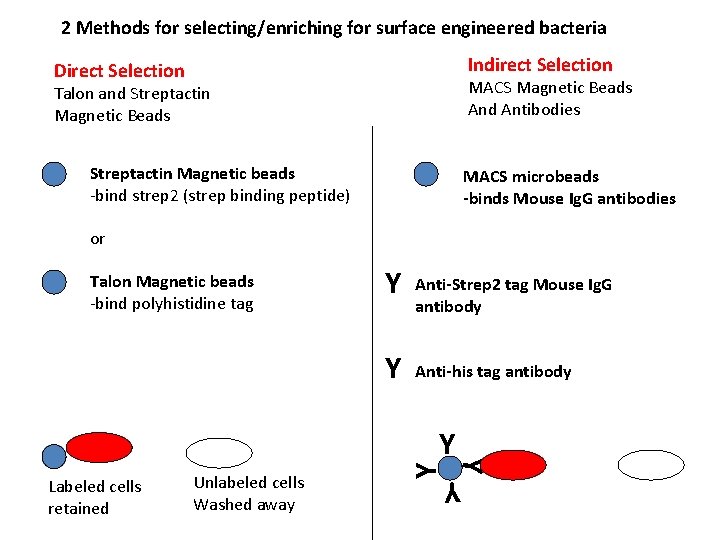
2 Methods for selecting/enriching for surface engineered bacteria Indirect Selection Direct Selection MACS Magnetic Beads And Antibodies Talon and Streptactin Magnetic Beads Streptactin Magnetic beads -bind strep 2 (strep binding peptide) MACS microbeads -binds Mouse Ig. G antibodies or Talon Magnetic beads -bind polyhistidine tag Y Y Anti-Strep 2 tag Mouse Ig. G antibody Anti-his tag antibody Y Unlabeled cells Washed away Y Labeled cells retained Y Y
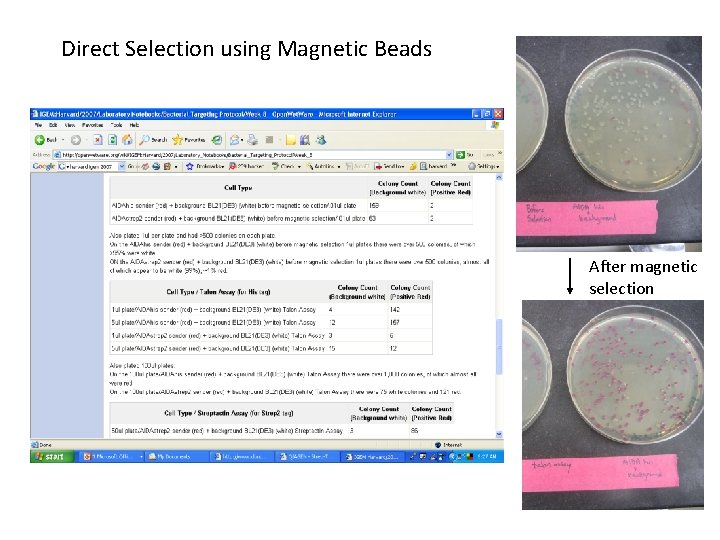
Direct Selection using Magnetic Beads After magnetic selection
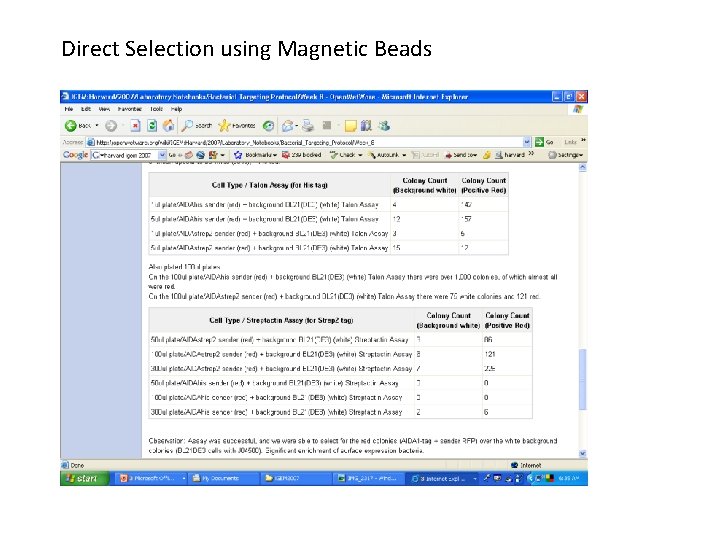
Direct Selection using Magnetic Beads
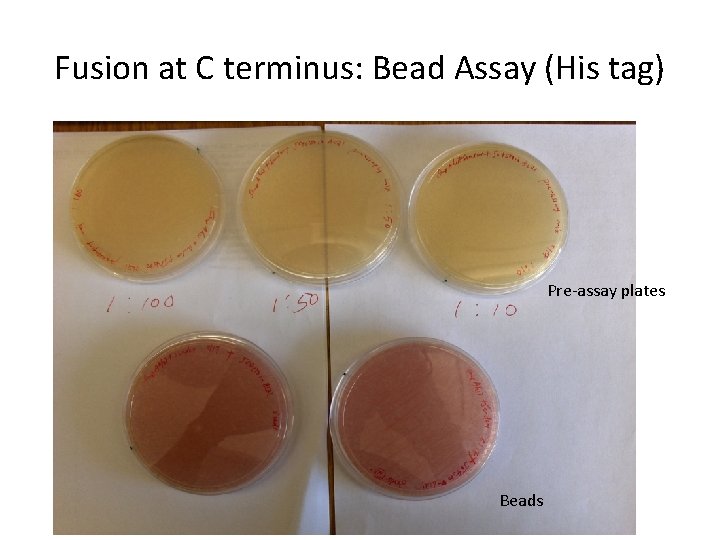
Fusion at C terminus: Bead Assay (His tag) Pre-assay plates Beads

Loop Insertion • PCR product digestion & ligation – Primer design – Digest-Ligate-Transform motif • Gene design – Insertion points created for inserting synthetic constructs
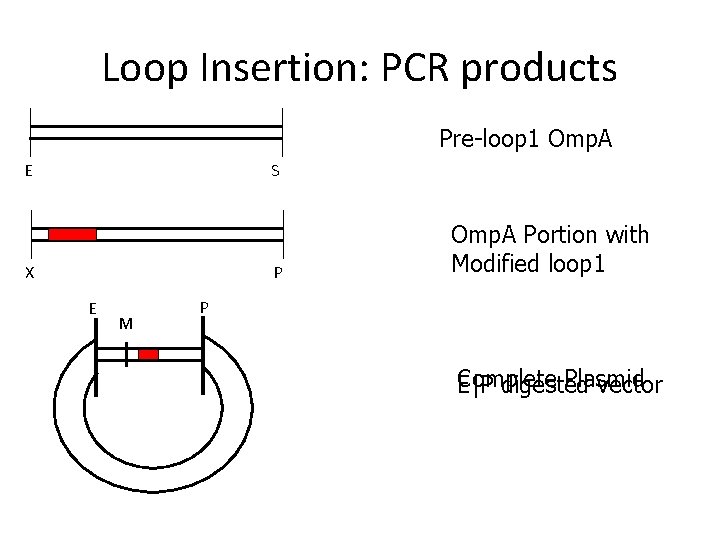
Loop Insertion: PCR products Pre-loop 1 Omp. A E S X P E M Omp. A Portion with Modified loop 1 P Complete Plasmid E|P digested vector
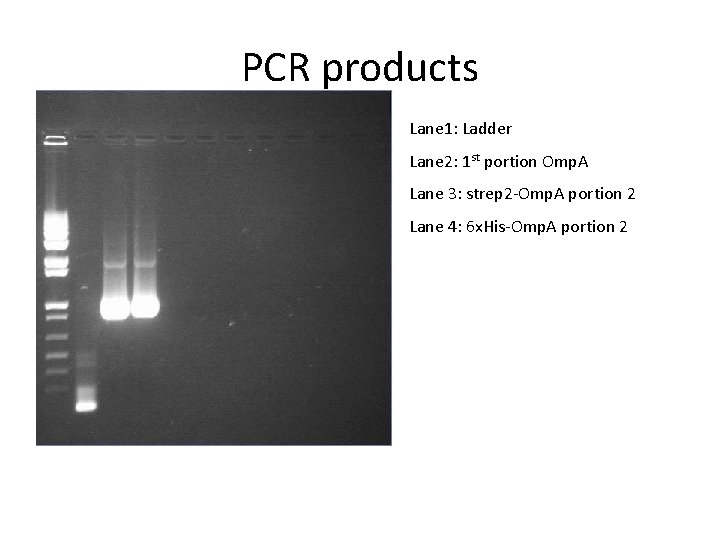
PCR products Lane 1: Ladder Lane 2: 1 st portion Omp. A Lane 3: strep 2 -Omp. A portion 2 Lane 4: 6 x. His-Omp. A portion 2
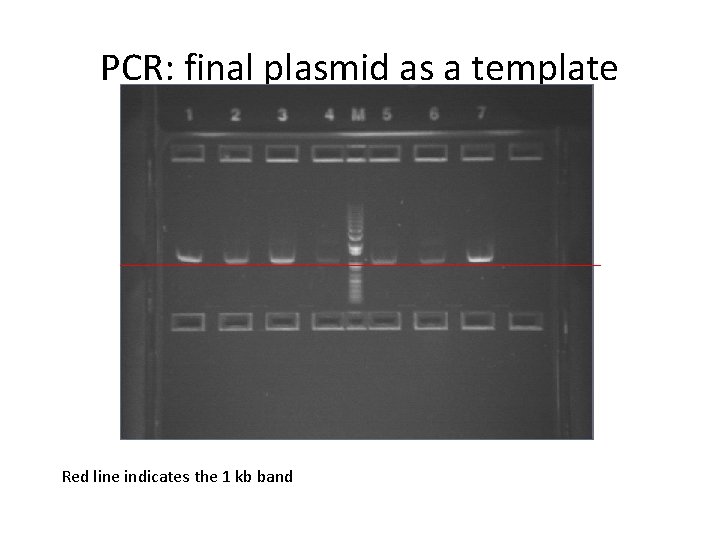
PCR: final plasmid as a template Red line indicates the 1 kb band
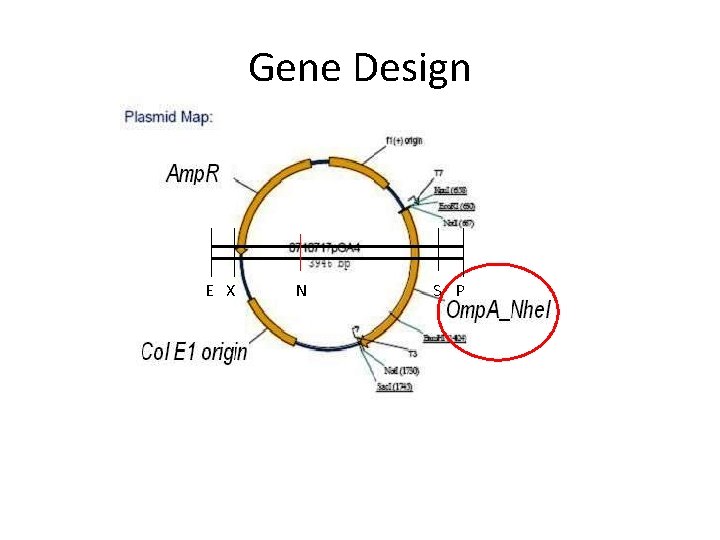
Gene Design E X N S P
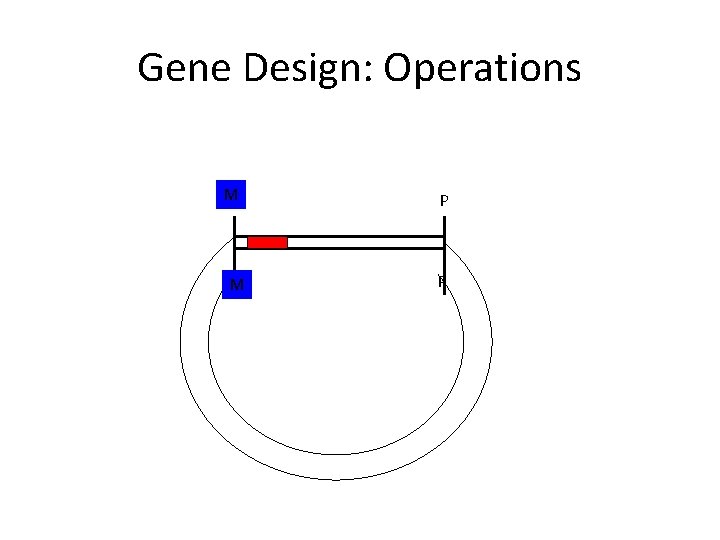
Gene Design: Operations N M X M P P
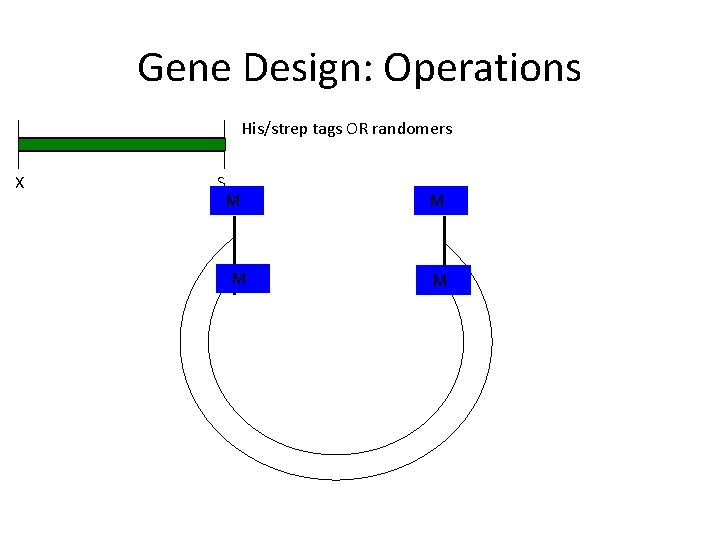
Gene Design: Operations His/strep tags OR randomers X S N M M MN M
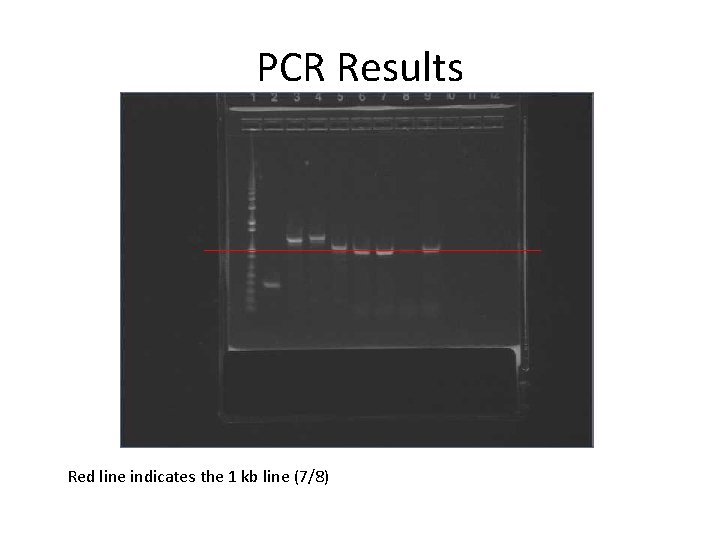
PCR Results Red line indicates the 1 kb line (7/8)
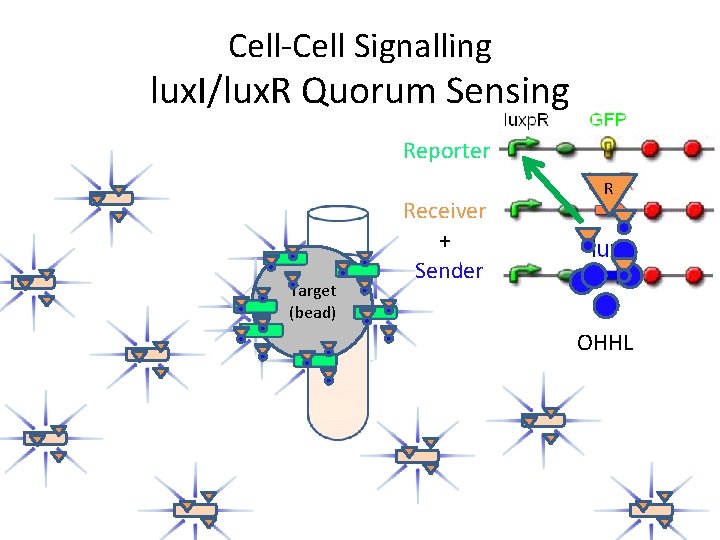
Cell-Cell Signalling lux. I/lux. R Quorum Sensing Reporter Target (bead) Receiver + Sender R OHHL
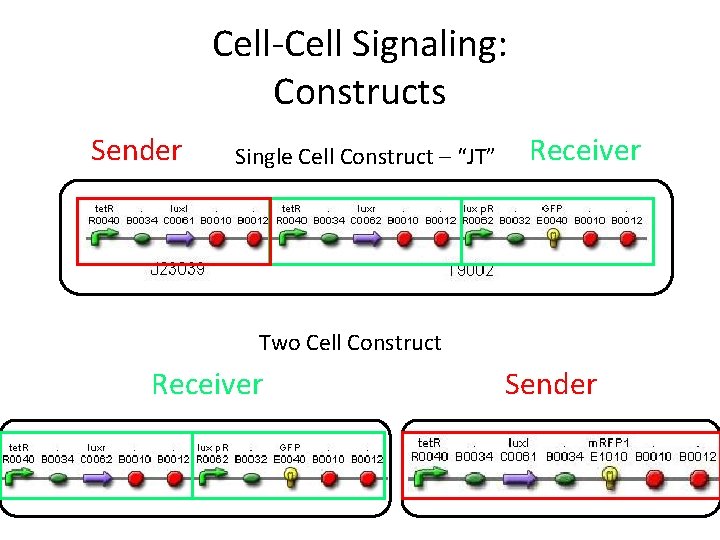
Cell-Cell Signaling: Constructs Sender Single Cell Construct – “JT” Receiver Two Cell Construct Receiver Sender
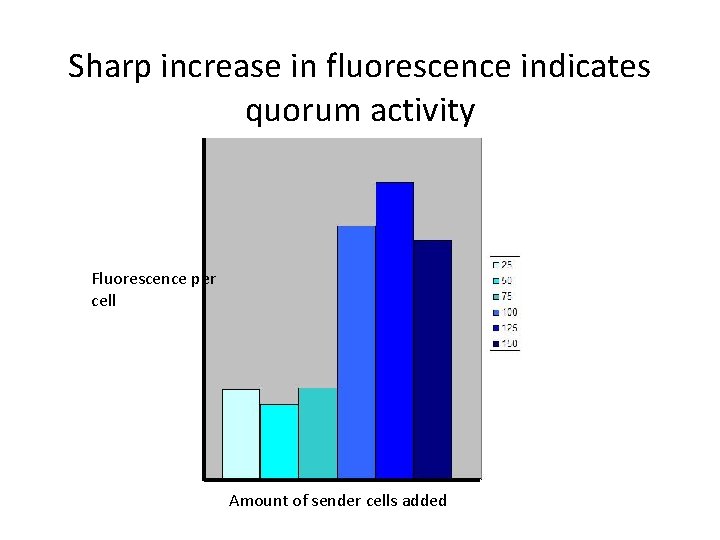
Sharp increase in fluorescence indicates quorum activity Fluorescence per cell Amount of sender cells added
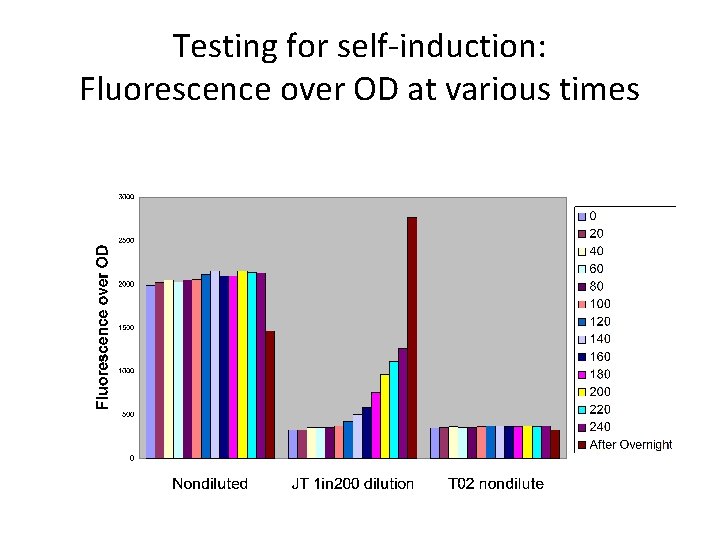
Testing for self-induction: Fluorescence over OD at various times

Direct Magnetic Beads: Good Enrichment
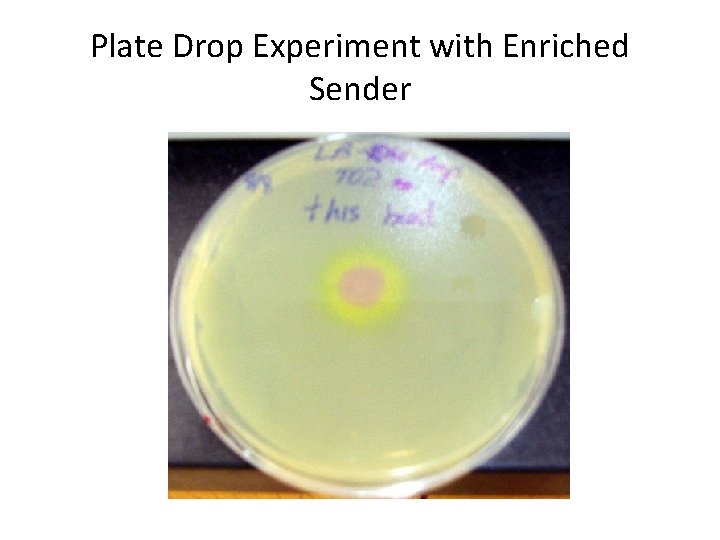
Plate Drop Experiment with Enriched Sender
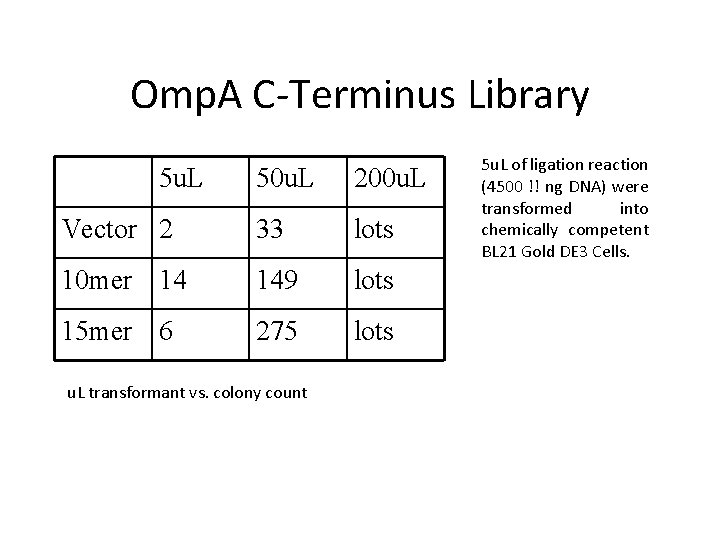
Omp. A C-Terminus Library 5 u. L 50 u. L 200 u. L Vector 2 33 lots 10 mer 14 149 lots 15 mer 6 275 lots u. L transformant vs. colony count 5 u. L of ligation reaction (4500 !! ng DNA) were transformed into chemically competent BL 21 Gold DE 3 Cells.
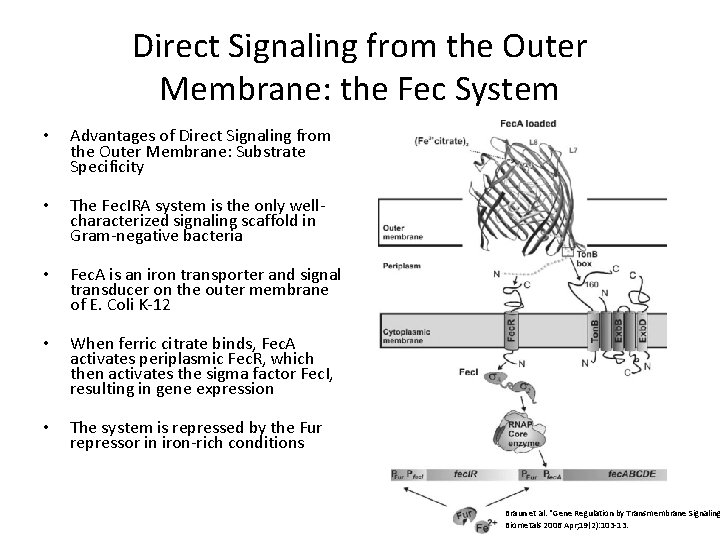
Direct Signaling from the Outer Membrane: the Fec System • Advantages of Direct Signaling from the Outer Membrane: Substrate Specificity • The Fec. IRA system is the only wellcharacterized signaling scaffold in Gram-negative bacteria • Fec. A is an iron transporter and signal transducer on the outer membrane of E. Coli K-12 • When ferric citrate binds, Fec. A activates periplasmic Fec. R, which then activates the sigma factor Fec. I, resulting in gene expression • The system is repressed by the Fur repressor in iron-rich conditions Braun et al. “Gene Regulation by Transmembrane Signaling Biometals 2006 Apr; 19(2): 103 -13.

Fec: Motivation and Methods • Structural information suggests possibility of maintaining signaling with changed binding. – L 7 moves up to 11Å, helix unwinds – L 8 moves up to 15Å • Select binding targets by inserting random library, controls known to bind nickel and streptavidin into loops 7 and 8. – Even if signaling cannot be maintained, binding of controls proves that Fec. A can be used as scaffold for surface expression of peptides • Computational approach in collaboration with the lab of Costas Maranas, Penn State Dept of Chemical Engineering. Ferguson AD et al. “Structural Basis of Gating by the Outer Membrane Transport Science 2002 Mar 1: 295(5560) 1715 -9
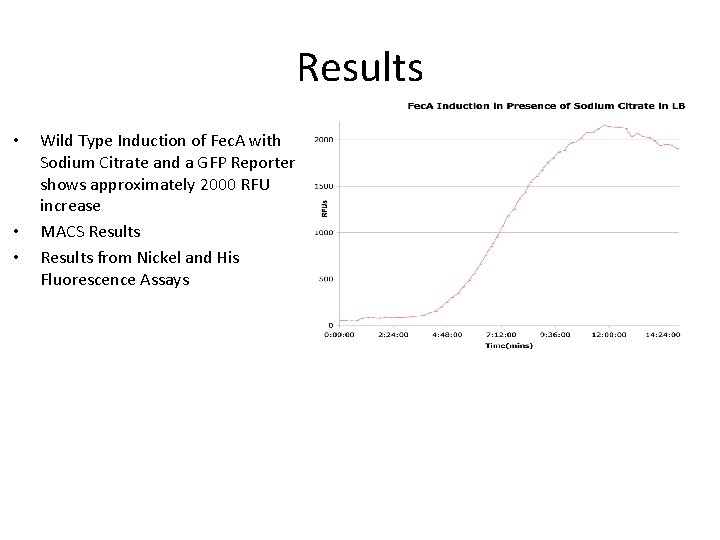
Results • • • Wild Type Induction of Fec. A with Sodium Citrate and a GFP Reporter shows approximately 2000 RFU increase MACS Results from Nickel and His Fluorescence Assays

Biobricking the Fec System Construct Features: • Swappable Fec. A - Fec. A is flanked by Nhe 1 and Afl. II sites to allow the easy mutagenesis and replacement of Fec. A. • Variable Promoters - each component will be on a separate constitutive promoter. • The optimization of GFP expression using promoters of different strengths is planned.

Biobricking the Fec System • Mutagenesis of Fec promoter to weaken gene expression, providing a range of sensitivity. • Mutagenesis of the Fec promoter to remove FUR repressor binding site, allowing easier assays.

CONCLUSION • To be added

ACKNOWLEDGEMENTS • Thank you.
- Slides: 34