Classification of Cell Membrane Proteins Seyed Koosha Golmohammadi

Classification of Cell Membrane Proteins Seyed Koosha Golmohammadi, Lukasz Kurgan, Brendan Crowley, and Marek Reformat University of Alberta Department of Electrical and Computer Engineering This presentation and other related information are available at http: //www. ualberta. ca/~golmoham/ Frontiers in the Convergence of Bioscience and Information Technologies 2007

Problem definition • Knowledge of cell membrane protein type is important – Critical for determining their function – Determining type of protein using traditional experimental methods is costly and time consuming • Large and widening gap between known proteins (over 3. 3 million) and annotated proteins Automated and accurate methods of classifying uncharacterized proteins are highly desirable Classification of Cell Membrane Proteins 1/10
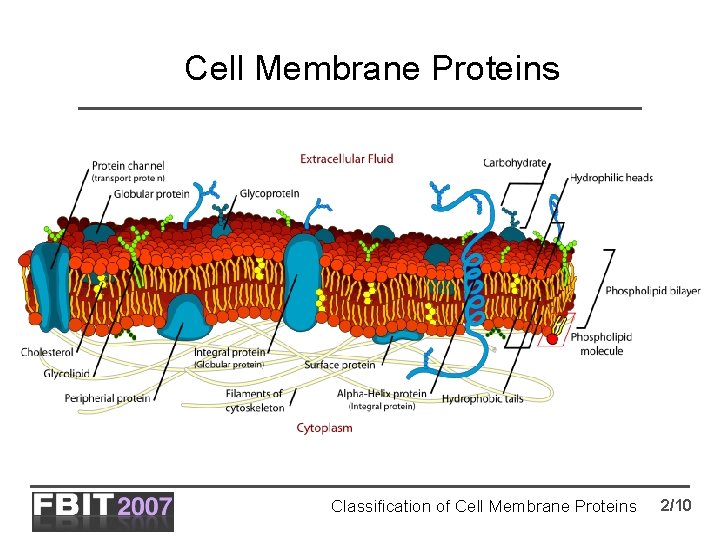
Cell Membrane Proteins Classification of Cell Membrane Proteins 2/10
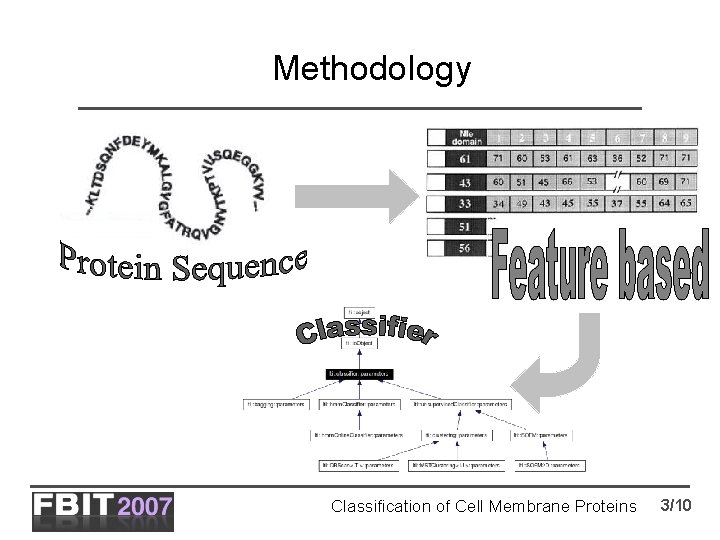
Methodology Classification of Cell Membrane Proteins 3/10

Datasets and test procedures • Two datasets were used to design and test our system. These standard benchmark datasets allow for a fair comparison with other methods – 2059 proteins were used to design the prediction system Chou, Prediction of protein cellular attributes using pseudo amino acid composition, Proteins (2001) 43: 246 -55 – 2625 proteins were used for an independent test Chou and Elrod, Prediction of membrane protein types and subcellular locations, Proteins (1999) 34: 137 -53 • Three test methods were used for evaluation of the performance of proposed prediction system – in-sample resubstitution (self-consistency) on the design dataset – out-of-sample jackknife (leave-one-out) on the design dataset – out-of-sample test on the independent dataset Classification of Cell Membrane Proteins 5/10
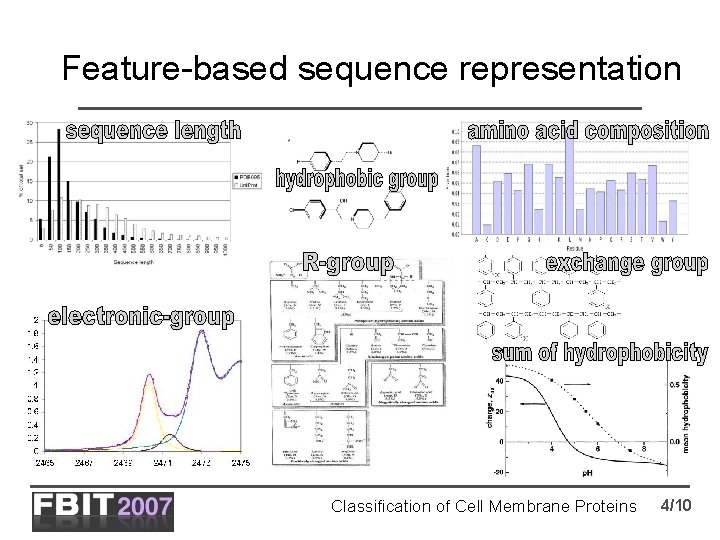
Feature-based sequence representation Classification of Cell Membrane Proteins 4/10
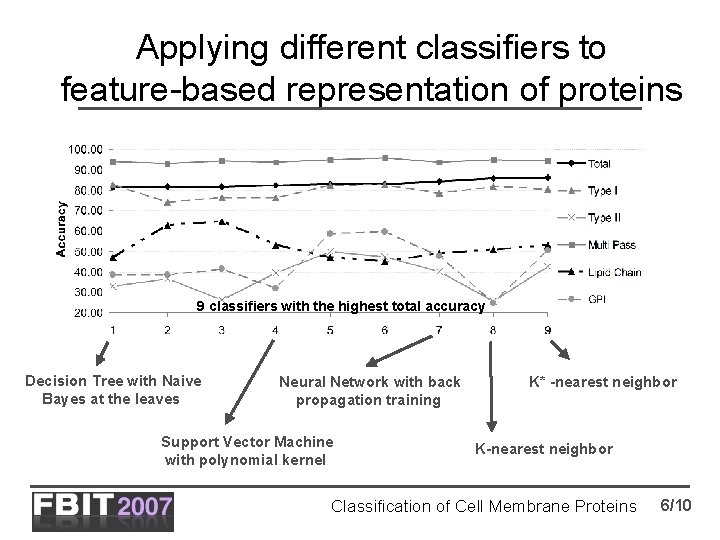
Applying different classifiers to feature-based representation of proteins 9 classifiers with the highest total accuracy Decision Tree with Naive Bayes at the leaves Neural Network with back propagation training Support Vector Machine with polynomial kernel K* -nearest neighbor K-nearest neighbor Classification of Cell Membrane Proteins 6/10
![Our method results in a glance Test method Accuracy [%] Specificity [%] Selfconsistency Jackknife Our method results in a glance Test method Accuracy [%] Specificity [%] Selfconsistency Jackknife](http://slidetodoc.com/presentation_image_h2/7fa084d48b8fb7210d172d20ee3a70ab/image-8.jpg)
Our method results in a glance Test method Accuracy [%] Specificity [%] Selfconsistency Jackknife Independent Overall 99. 9 86. 9 97. 1 Type I 100 83. 5 Type II 100 52. 6 96. 4 80. 6 Multipass 100 95. 8 Lipid 100 45. 1 99. 0 78. 6 GPI 99. 1 61. 5 96. 5 Type I 100 94. 7 Type II 100 98. 3 99. 2 99. 8 Multipass 99. 9 83. 4 Lipid 100 99. 9 93. 9 99. 9 GPI 100 98. 7 99. 8 Classification of Cell Membrane Proteins 7/10
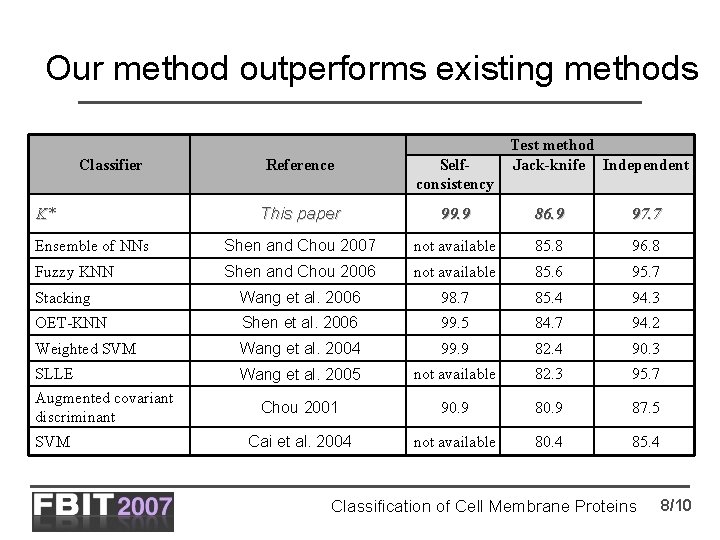
Our method outperforms existing methods Classifier Test method Jack-knife Independent Reference Selfconsistency This paper 99. 9 86. 9 97. 7 Ensemble of NNs Shen and Chou 2007 not available 85. 8 96. 8 Fuzzy KNN Shen and Chou 2006 not available 85. 6 95. 7 Stacking Wang et al. 2006 98. 7 85. 4 94. 3 OET-KNN Shen et al. 2006 99. 5 84. 7 94. 2 Weighted SVM Wang et al. 2004 99. 9 82. 4 90. 3 SLLE Augmented covariant discriminant Wang et al. 2005 not available 82. 3 95. 7 Chou 2001 90. 9 87. 5 Cai et al. 2004 not available 80. 4 85. 4 K* SVM Classification of Cell Membrane Proteins 8/10

Conclusions • The proposed method outperforms existing methods – higher accuracy in both jackknife and independent dataset tests • The improved prediction quality of our method is a result of applying a comprehensive feature-based sequence representation – existing methods use either composition or pseudo amino acid composition for protein representation. – in contrast, our method uses seven feature sets for the same task – there might be other features that are not tested in this study and could further improve the prediction accuracy Classification of Cell Membrane Proteins 9/10
- Slides: 10